-
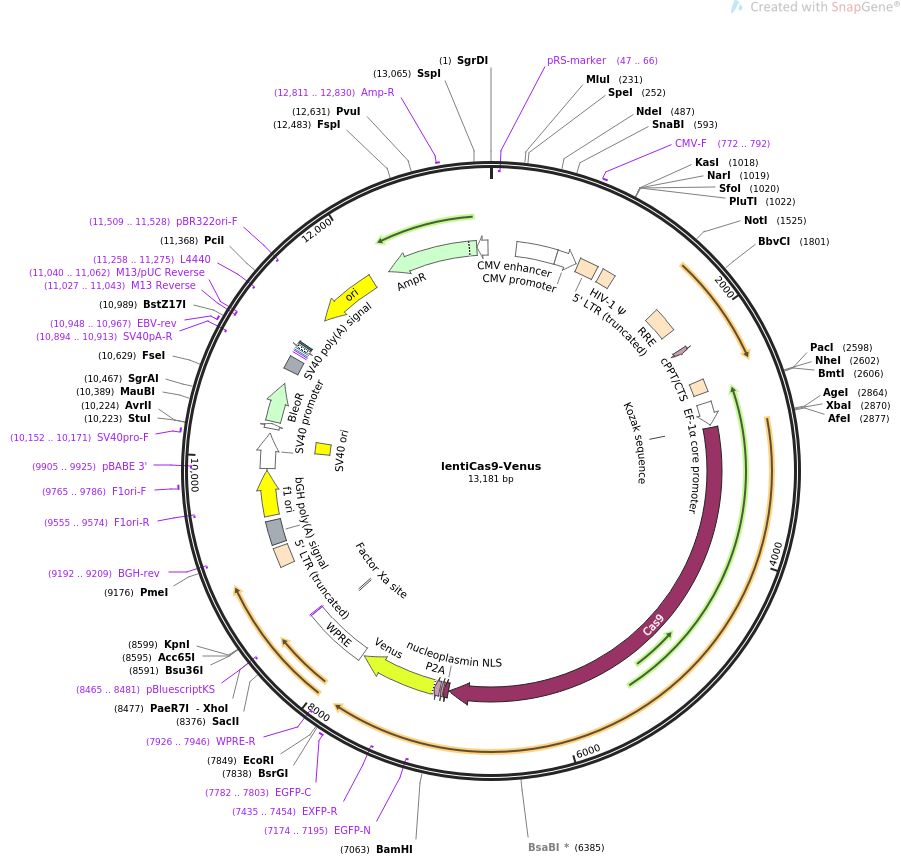
PurposeExpresses human codon-optimized S. pyogenes Cas9 protein and Venus reporter from EFS promoter. Lentiviral backbone.
-
Depositing Lab
-
Sequence Information
Ordering
| Item | Catalog # | Description | Quantity | Price (USD) | |
|---|---|---|---|---|---|
| Plasmid | 70267 | Standard format: Plasmid sent in bacteria as agar stab | 1 | $85 | |
This material is available to academics and nonprofits only.
Backbone
-
Vector backbonepFUGW
- Backbone size w/o insert (bp) 9080
- Total vector size (bp) 13181
-
Vector typeMammalian Expression, Lentiviral, CRISPR
-
Selectable markersVenus
Growth in Bacteria
-
Bacterial Resistance(s)Ampicillin, 100 μg/mL
-
Growth Temperature37°C
-
Growth Strain(s)NEB Stable
-
Growth instructionsOnly amplify in RecA- bacteria (eg. Stbl3).
-
Copy numberHigh Copy
Gene/Insert 1
-
Gene/Insert nameCas9
-
Alt nameS. pyogenes CRISPR-Cas9
-
SpeciesSynthetic
-
Insert Size (bp)4101
- Promoter EFS-NS
-
Tag
/ Fusion Protein
- FLAG (C terminal on insert)
Cloning Information for Gene/Insert 1
- Cloning method Restriction Enzyme
- 5′ cloning site AgeI, XbaI, AfeI (not destroyed)
- 3′ cloning site BamHI (not destroyed)
- 5′ sequencing primer TTCTGCAGACAAATGGCAGT
- 3′ sequencing primer GCTGAACTTGTGGCCGTTTA (Common Sequencing Primers)
Gene/Insert 2
-
Gene/Insert nameVenus Reporter
-
SpeciesSynthetic
-
Insert Size (bp)720
- Promoter P2A
Cloning Information for Gene/Insert 2
- Cloning method Restriction Enzyme
- 5′ cloning site BamHI (not destroyed)
- 3′ cloning site EcoRI (not destroyed)
- 5′ sequencing primer TACGAGACACGGATCGACCT
- 3′ sequencing primer AAGCCATACGGGAAGCAATA (Common Sequencing Primers)
Terms and Licenses
-
Academic/Nonprofit Terms
-
Industry Terms
- Not Available to Industry
Trademarks:
- Zeocin® is an InvivoGen trademark.
Depositor Comments
Subcloned Venus in place of Blast in lentiCas9-Blast.
Improved vectors and genome-wide libraries for CRISPR screening. Sanjana NE, Shalem O, Zhang F. Nat Methods. 2014 Aug;11(8):783-4. doi: 10.1038/nmeth.3047. 10.1038/nmeth.3047 PubMed 25075903
Canver MC, Smith EC, Sher F, Pinello L, Sanjana NE, Shalem O, Chen DD, Schupp PG, Vinjamur DS, Garcia SP, Luc S, Kurita R, Nakamura Y, Fujiwara Y, Maeda T, Yuan G, Zhang F, Orkin SH, Bauer DE. BCL11A enhancer dissection by Cas9-mediated in situ saturating mutagenesis. Nature. 2015 September 16; in press; doi:10.1038/nature15521.
These plasmids were created by your colleagues. Please acknowledge the Principal Investigator, cite the article in which the plasmids were described, and include Addgene in the Materials and Methods of your future publications.
-
For your Materials & Methods section:
lentiCas9-Venus was a gift from Daniel Bauer (Addgene plasmid # 70267 ; http://n2t.net/addgene:70267 ; RRID:Addgene_70267) -
For your References section:
BCL11A enhancer dissection by Cas9-mediated in situ saturating mutagenesis. Canver MC, Smith EC, Sher F, Pinello L, Sanjana NE, Shalem O, Chen DD, Schupp PG, Vinjamur DS, Garcia SP, Luc S, Kurita R, Nakamura Y, Fujiwara Y, Maeda T, Yuan GC, Zhang F, Orkin SH, Bauer DE. Nature. 2015 Sep 16. doi: 10.1038/nature15521. 10.1038/nature15521 PubMed 26375006