-
Purpose(Empty Backbone)
-
Depositing Lab
-
Publication
-
Sequence Information
Ordering
| Item | Catalog # | Description | Quantity | Price (USD) | |
|---|---|---|---|---|---|
| Plasmid | 21873 | Standard format: Plasmid sent in bacteria as agar stab | 1 | $85 | |
This material is available to academics and nonprofits only.
Backbone
-
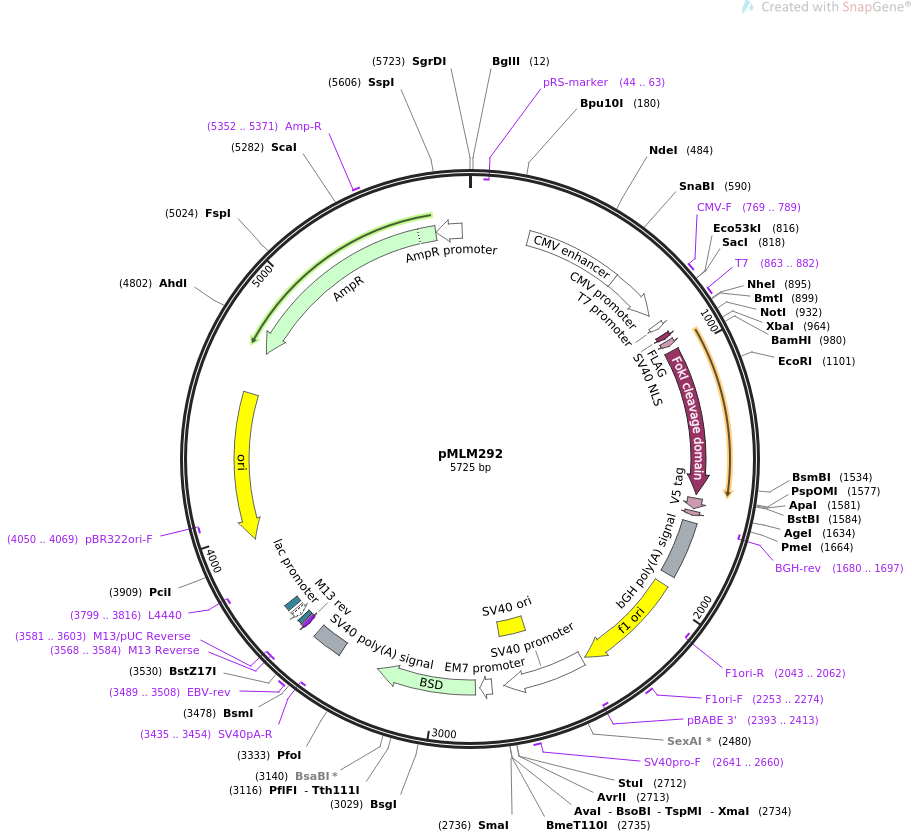
Vector backbonepMLM292
- Backbone size (bp) 5725
-
Vector typeMammalian Expression ; zebrafish expression
Growth in Bacteria
-
Bacterial Resistance(s)Ampicillin, 100 μg/mL
-
Growth Temperature37°C
-
Growth Strain(s)DH5alpha
-
Copy numberHigh Copy
Gene/Insert
-
Gene/Insert nameNone
-
Tag
/ Fusion Protein
- Flag (N terminal on backbone)
Cloning Information
- Cloning method Restriction Enzyme
- 5′ cloning site XbaI (not destroyed)
- 3′ cloning site BamHI (not destroyed)
- 5′ sequencing primer CMV-F (Common Sequencing Primers)
Resource Information
-
Articles Citing this Plasmid
Terms and Licenses
-
Academic/Nonprofit Terms
-
Industry Terms
- Not Available to Industry
Trademarks:
- Zeocin® is an InvivoGen trademark.
Depositor Comments
Zinc fingers are cloned in between the unique XbaI and BamHI sites. This results in an expression vector which encodes a ZFN with the OPEN zinc finger array fused to the mutant FokI nuclease domain.
pMLM292 is identical to pST1374 except that it harbors two mutations in the FokI nuclease domain (Q486E, I499L; aka the “-“ mutation see Miller et al., Nat. Biotech 2007, PMID 17603475) which confers heterodimeric behavior on these domains
235- 822: CMV promoter / 771-775: CAAT box / 804-808: TATA box / 819: 3' end of hCMV / 904-906: ZFN start codon / 910-930: SV40 nuclease localization signal (NLS) / 940- 963: FLAG tag / 964- 969: XbaI site / 980- 985: BamHI site / 986-1573: FokI nuclease domain / 1292&1294: Q486E heterodimer FokI mutation / 1331: I499L heterodimer FokI mutation / 1574-1576: ZFN stop codon / 1686-1910: BGH polyadenylation sequence / 1956-2384: f1 origin / 2411-2720: SV40 early promoter and origin / 2775-2841: EM-7 Promoter / 2842-3240: Blasticidin resistance gene / 3398-3528: SV40 late polyA signal / 3955-4574: pUC origin / 4729-5589: ampicillin (bla) resistance gene (complementary strand) / 5590-5688: bla promoter (complementary strand)
These plasmids were created by your colleagues. Please acknowledge the Principal Investigator, cite the article in which the plasmids were described, and include Addgene in the Materials and Methods of your future publications.
-
For your Materials & Methods section:
pMLM292 was a gift from Keith Joung (Addgene plasmid # 21873 ; http://n2t.net/addgene:21873 ; RRID:Addgene_21873) -
For your References section:
Targeted mutagenesis in zebrafish using customized zinc-finger nucleases. Foley JE, Maeder ML, Pearlberg J, Joung JK, Peterson RT, Yeh JR. Nat Protoc. 2009. 4(12):1855-67. 10.1038/nprot.2009.209 PubMed 20010934