-
Depositing Lab
-
Publication
-
Sequence Information
Ordering
| Item | Catalog # | Description | Quantity | Price (USD) | |
|---|---|---|---|---|---|
| Plasmid | 24349 | Standard format: Plasmid sent in bacteria as agar stab | 1 | $85 | |
Backbone
-
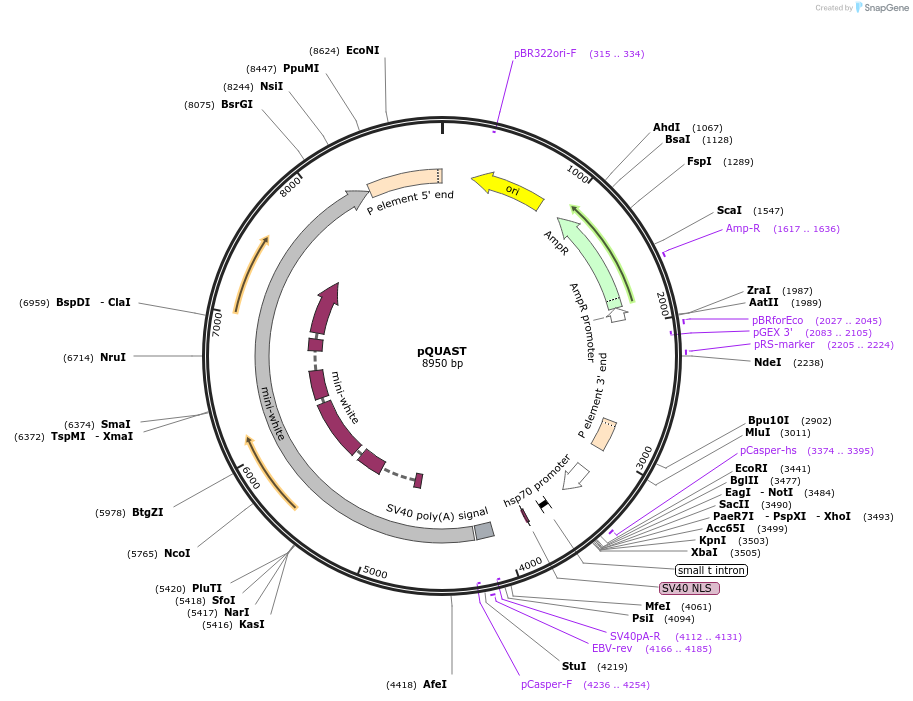
Vector backbonepUAST
-
Backbone manufacturerBrand and Perrimon, 1993
- Backbone size w/o insert (bp) 8950
-
Vector typeInsect Expression
Growth in Bacteria
-
Bacterial Resistance(s)Ampicillin, 100 μg/mL
-
Growth Temperature37°C
-
Growth Strain(s)DH5alpha
-
Copy numberHigh Copy
Gene/Insert
-
Gene/Insert name5xQUAS-Pmin promoter
-
SpeciesD. melanogaster (fly)
Cloning Information
- Cloning method Restriction Enzyme
- 5′ cloning site BamHI (destroyed during cloning)
- 3′ cloning site EcoRI (destroyed during cloning)
- 5′ sequencing primer pCasper-hs
- 3′ sequencing primer SV40pA-R (Common Sequencing Primers)
Resource Information
-
Reference
-
Articles Citing this Plasmid
Terms and Licenses
-
Academic/Nonprofit Terms
-
Industry Terms
- Not Available to Industry
Trademarks:
- Zeocin® is an InvivoGen trademark.
Depositor Comments
This vector was designed to mimic the multi-cloning site of the pUAST vector (Brand and
Perrimon, 1993) thereby allowing easy exchange of inserts between pUAST and pQUAST. pUAST
was digest by SphI and EcoRI and blunted with Klenow to remove the 5xUAS and hsp70 minimal
promoter (minP). pQUAS-GG was digested with BamHI and EcoRI to excise the 5xQUAS and Pmin
and then blunted. The 5xQUAS-Pmin promoter was then ligated into the modified pUAST vector to
generate pQUAST. Any gene X can be subcloned from pUAST-geneX into pQUAST using the
same restriction sites that were originally utilized for pUAST-geneX construction. If the pUAST-
geneX plasmid is not available, genomic DNA from flies containing the UAS-geneX transgene can
be used. In this case, pQUAST-geneX can be constructed as follows:
1) PCR amplify the UAS insert from UAS-geneX genomic DNA by using the primer pairs
genUASFOR (GCTTCGTCTACGGAGCGACAATTCAATTCAAAC) and genUASREVsv40
(GCAGTAGCCTCATCATCACTAGATGGCATTTCTTC). These primers will amplify the insert for
any UAS construct, including the restriction sites used for cloning of that insert into the UAS vector.
2) If the restriction sites used for cloning of the UAS insert are unknown, sequence the PCR
fragment using UASFOR-SEQ (TCAAACAAGCAAAGTGAACACG) and SV40REV-SEQ
(CCATTCATCAGTTCCATAGGTTGG) primers.
3) Digest the PCR product with appropriate enzymes for cloning into pQUAST.
These plasmids were created by your colleagues. Please acknowledge the Principal Investigator, cite the article in which the plasmids were described, and include Addgene in the Materials and Methods of your future publications.
-
For your Materials & Methods section:
pQUAST was a gift from Liqun Luo (Addgene plasmid # 24349 ; http://n2t.net/addgene:24349 ; RRID:Addgene_24349) -
For your References section:
The Q System: A Repressible Binary System for Transgene Expression, Lineage Tracing, and Mosaic Analysis. Potter CJ, Tasic B, Russler EV, Liang L, Luo L.. Volume 141, Issue 3, 536-548, 30 April 2010 10.1016/j.cell.2010.02.025 PubMed 20434990