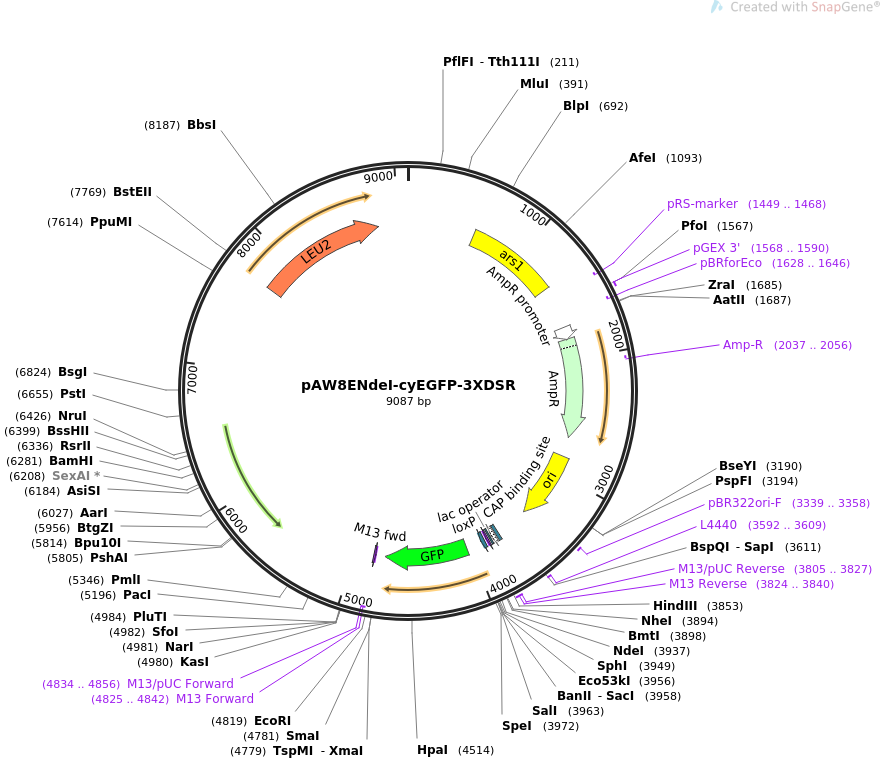
pAW8ENdeI-cyEGFP-3XDSR
(Plasmid
#49863)
-
Purpose(Empty Backbone) for Cre-lox cassette exchange of C-terminal yEGFP tagged gene sequences together with 3 copies of the core DSR motif into the S. pombe urg1 locus
-
Depositing Lab
-
Sequence Information
Ordering
| Item | Catalog # | Description | Quantity | Price (USD) | |
|---|---|---|---|---|---|
| Plasmid | 49863 | Standard format: Plasmid sent in bacteria as agar stab | 1 | $85 | |
This material is available to academics and nonprofits only.
Backbone
-
Vector backbonepUC
-
Vector typeCre/Lox
-
Selectable markersLEU2
-
Tag
/ Fusion Protein
- yEGFP (C terminal on backbone)
Growth in Bacteria
-
Bacterial Resistance(s)Ampicillin, 100 μg/mL
-
Growth Temperature37°C
-
Growth Strain(s)DH5alpha
-
Copy numberHigh Copy
Cloning Information
- Cloning method Restriction Enzyme
Terms and Licenses
-
Academic/Nonprofit Terms
-
Industry Terms
- Not Available to Industry
Trademarks:
- Zeocin® is an InvivoGen trademark.
Depositor Comments
This plasmid contains a wild-type loxP site and a mutant loxP site termed loxM3. Addgene's mapping software can not differentiate between the two sites and identifies the loxM3 site as loxP. The locations of the loxP and loxM3 sites are:
loxP = 3859-3892
loxM3 = 4785-4818
These plasmids were created by your colleagues. Please acknowledge the Principal Investigator, cite the article in which the plasmids were described, and include Addgene in the Materials and Methods of your future publications.
-
For your Materials & Methods section:
pAW8ENdeI-cyEGFP-3XDSR was a gift from Tony Carr (Addgene plasmid # 49863 ; http://n2t.net/addgene:49863 ; RRID:Addgene_49863) -
For your References section:
Optimisation of the Schizosaccharomyces pombe urg1 Expression System. Watson AT, Daigaku Y, Mohebi S, Etheridge TJ, Chahwan C, Murray JM, Carr AM. PLoS One. 2013 Dec 20;8(12):e83800. doi: 10.1371/journal.pone.0083800. PubMed 24376751