-
Depositing Lab
-
Publication
-
Sequence Information
Ordering
| Item | Catalog # | Description | Quantity | Price (USD) | |
|---|---|---|---|---|---|
| Plasmid | 14785 | Standard format: Plasmid sent in bacteria as agar stab | 1 | $85 | |
Backbone
-
Vector backbonepIS1
-
Backbone manufacturerAvailable from Addgene (#12179)
- Backbone size w/o insert (bp) 4000
-
Vector typeMammalian Expression, Luciferase
Growth in Bacteria
-
Bacterial Resistance(s)Ampicillin, 100 μg/mL
-
Growth Temperature37°C
-
Growth Strain(s)DH5alpha
-
Copy numberHigh Copy
Gene/Insert
-
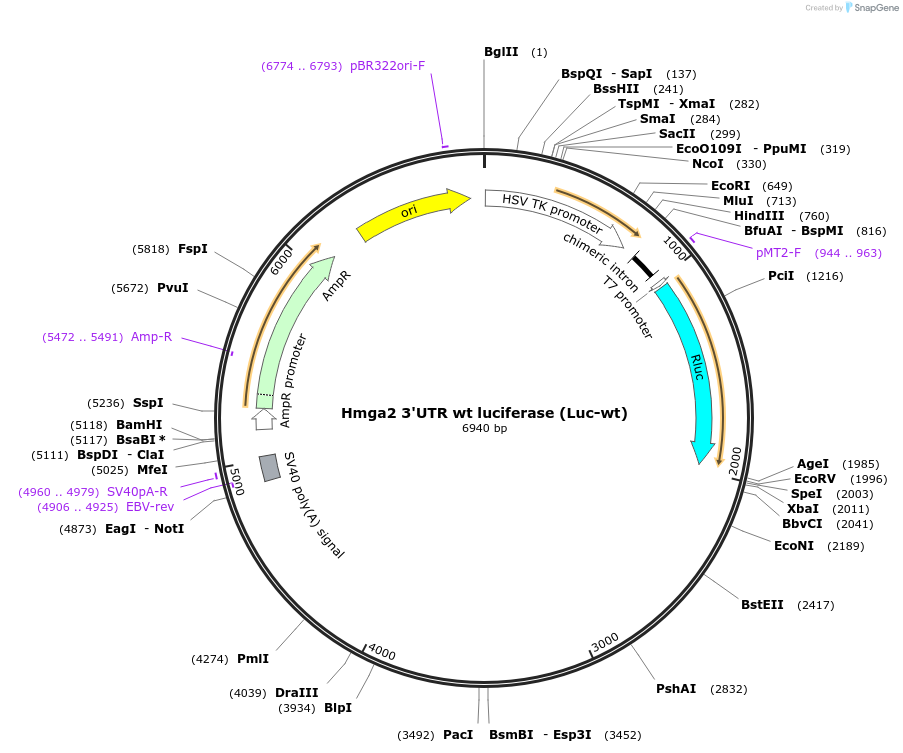
Gene/Insert nameLuc-wt
-
Alt nameHMGA2
-
SpeciesM. musculus (mouse)
-
Insert Size (bp)3000
-
GenBank IDBC052158
-
Entrez GeneHmga2 (a.k.a. 9430083A20Rik, HMGI-C, Hmgic, pg, pygmy)
-
Tag
/ Fusion Protein
- luciferase (N terminal on backbone)
Cloning Information
- Cloning method Restriction Enzyme
- 5′ cloning site XbaI (not destroyed)
- 3′ cloning site NotI (not destroyed)
- 5′ sequencing primer N/A
- 3′ sequencing primer EBV rev primer (Common Sequencing Primers)
Resource Information
-
Articles Citing this Plasmid
Terms and Licenses
-
Academic/Nonprofit Terms
-
Industry Terms
- Not Available to Industry
Trademarks:
- Zeocin® is an InvivoGen trademark.
Depositor Comments
The 3' UTR of the mouse Hmga2 cDNA (BC052158) was subcloned into pCR2.1-TOPO (Invitrogen) for site-directed mutagenesis. To disrupt each let-7 complementary site, the nucleotides that paired to nucleotides 3 and 5 of the miRNA were substituted (see Author's map), using the Quikchange site-directed mutagenesis kit (Stratagene). To construct the Renilla luciferase reporters, wild-type and mutant Hmga2 3' UTRs were amplified (PCR primers, 5'-GCGTCTCGAGGGGCGCCGACATTC and 5'GGCGCGGCCGCAGTCAGAGGGCACAC) and cloned into the XbaI and NotI sites of pIS1.
These plasmids were created by your colleagues. Please acknowledge the Principal Investigator, cite the article in which the plasmids were described, and include Addgene in the Materials and Methods of your future publications.
-
For your Materials & Methods section:
Hmga2 3'UTR wt luciferase (Luc-wt) was a gift from David Bartel (Addgene plasmid # 14785 ; http://n2t.net/addgene:14785 ; RRID:Addgene_14785) -
For your References section:
Disrupting the pairing between let-7 and Hmga2 enhances oncogenic transformation. Mayr C, Hemann MT, Bartel DP. Science. 2007 Mar 16. 315(5818):1576-9. 10.1126/science.1137999 PubMed 17322030