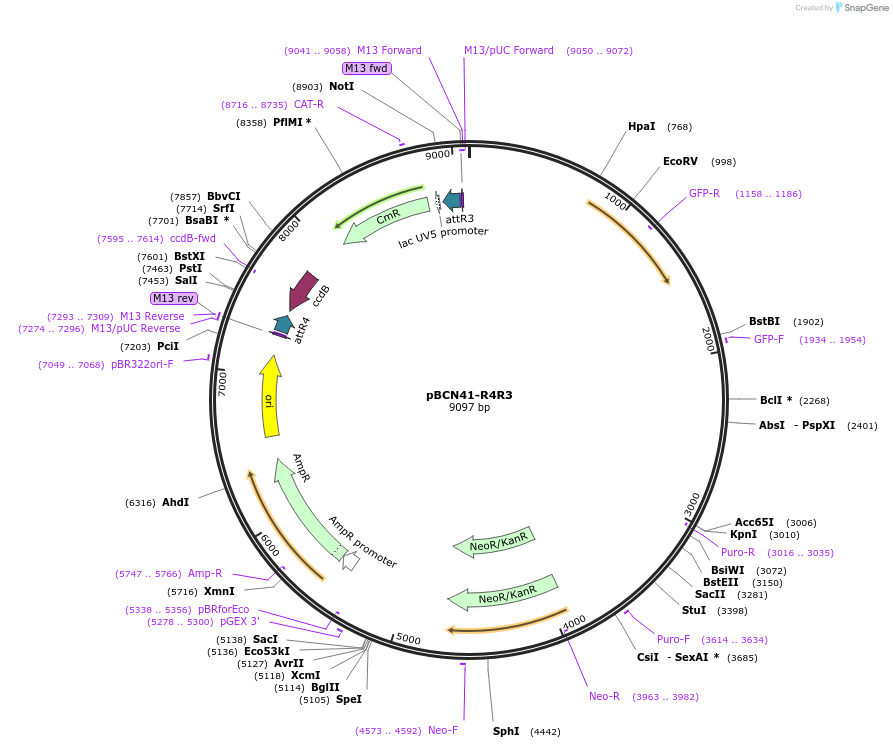
pBCN41-R4R3
(Plasmid
#34916)
-
Purpose(Empty Backbone)
-
Depositing Lab
-
Sequence Information
Ordering
| Item | Catalog # | Description | Quantity | Price (USD) | |
|---|---|---|---|---|---|
| Plasmid | 34916 | Standard format: Plasmid sent in bacteria as agar stab | 1 | $85 | |
Backbone
-
Vector backbonepBCN21-R4R3
-
Modifications to backboneContains the dual antibiotic resistance operon Prpl-28::PuroR::rpl-16_outron::NeoR::let-858_3'UTR instead of the puromycin resistance gene. Spe I, Xcm I, Bgl II, Avr I and Sac I restriction sites introduced into backbone for linearisation before bombardment. The ccdB gene was replaced with a version from pDONR221 (Invitrogen) that is compatible with commercially available ccdB Survival E. coli, no longer requiring DB3.1 cells. Contains a new version of Pmyo-2::EGFP::myo-2_3'UTR transgene in the backbone that expresses well in various Caenorhabditis species.
-
Vector typeWorm Expression
-
Selectable markersNeomycin (select with G418), Puromycin
Growth in Bacteria
-
Bacterial Resistance(s)Chloramphenicol and Ampicillin, 25 & 100 μg/mL
-
Growth Temperature37°C
-
Growth Strain(s)ccdB Survival
-
Growth instructionsAs a Gateway destination vector the empty vector must be grown in ccdB resistant cells such as DB3.1 or ccdB Survival. Use both Ampicillin and Chloramphenicol selection to avoid loss of the Gateway cassette. Once the Gateway cassette has been replaced with the gene of interest select with Ampicillin in standard E. coli strain such as Top10 or DH5alpha.
-
Copy numberUnknown
Resource Information
-
Supplemental Documents
-
A portion of this plasmid was derived from a plasmid made byPuromycin resistance gene from pBabePuro (Addgene). rpl-28 promoter and let-858 3'UTR sequences from pPD129.57 vector (Addgene). Neomycin resistance gene from pEGFP-C1 (Clontech). EGFP gene from pPD129.57 (Addgene).
Terms and Licenses
-
Academic/Nonprofit Terms
-
Industry Terms
- Not Available to Industry
Trademarks:
- Zeocin® is an InvivoGen trademark.
Depositor Comments
Dual resistance (PuroR-NeoR) vector for drug selection following biolistic bombardment in worms.
This vector contains a Pmyo-2::EGFP::myo-2_3'UTR pharyngeal marker on the backbone. Worm strains generated with vector can be crossed with strains generated with other visual markers such as in pBCN40-R4R3 which has a red fluorescent pharyngeal marker.
This is a Gateway 3-fragment compatible destination vector. It contains AttR4 and AttR3 sites (not AttR1 and AttR2 sites as identified automatically by the Addgene algorithm).
These plasmids were created by your colleagues. Please acknowledge the Principal Investigator, cite the article in which the plasmids were described, and include Addgene in the Materials and Methods of your future publications.
-
For your Materials & Methods section:
pBCN41-R4R3 was a gift from Ben Lehner (Addgene plasmid # 34916 ; http://n2t.net/addgene:34916 ; RRID:Addgene_34916) -
For your References section:
Generating transgenic nematodes by bombardment and antibiotic selection. Semple JI, Biondini L, Lehner B. Nat Methods. 2012 Jan 30;9(2):118-9. doi: 10.1038/nmeth.1864. 10.1038/nmeth.1864 PubMed 22290182