-
PurposeRFP with robust performance in protein fusions and useful as FRET acceptor for Clover in a FRET pair that offers bright fluorescence, dynamic range, and photostability while limiting emissions overlap
-
Depositing Lab
-
Sequence Information
Ordering
| Item | Catalog # | Description | Quantity | Price (USD) | |
|---|---|---|---|---|---|
| Plasmid | 40260 | Standard format: Plasmid sent in bacteria as agar stab | 1 | $85 | |
| Cloning Grade DNA | 40260-DNA.cg | 2 µg of cloning grade DNA in Tris buffer | 1 | $105 | |
Backbone
-
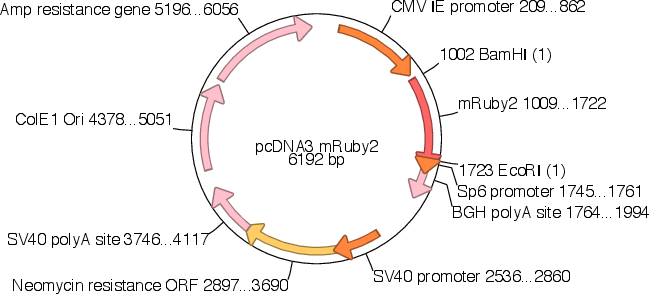
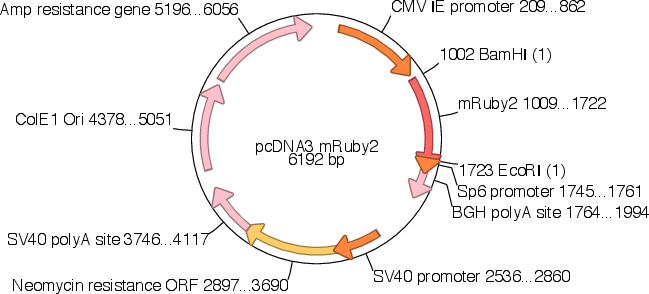
Vector backbonepcDNA3
-
Backbone manufacturerInvitrogen
- Backbone size w/o insert (bp) 5478
- Total vector size (bp) 6192
-
Modifications to backboneHis6 and Xpress tags at N-terminus of insert.
-
Vector typeMammalian Expression
-
Selectable markersNeomycin (select with G418)
Growth in Bacteria
-
Bacterial Resistance(s)Ampicillin, 100 μg/mL
-
Growth Temperature37°C
-
Growth Strain(s)DH5alpha
-
Copy numberHigh Copy
Gene/Insert
-
Gene/Insert namemRuby2
-
SpeciesSynthetic; Entacmaea quadricolor
-
Insert Size (bp)714
- Promoter CMV IE
-
Tags
/ Fusion Proteins
- His6 (N terminal on backbone)
- Xpress (N terminal on backbone)
Cloning Information
- Cloning method Restriction Enzyme
- 5′ cloning site BamHI (not destroyed)
- 3′ cloning site EcoRI (not destroyed)
- 5′ sequencing primer T7
- 3′ sequencing primer BGH Rev (Common Sequencing Primers)
Resource Information
-
Articles Citing this Plasmid
Terms and Licenses
-
Academic/Nonprofit Terms
-
Industry Terms
- Not Available to Industry
Trademarks:
- Zeocin® is an InvivoGen trademark.
Depositor Comments
mRuby2 is the brightest red fluorescent protein (RFP) characterized to date, with robust performance in protein fusions. Also, it is useful as FRET acceptor for Clover GFP in a FRET pair that offers bright fluorescence, dynamic range, and photostability while limiting emissions overlap.
Information for Cloning Grade DNA (Catalog # 40260-DNA.cg) ( Back to top)
Purpose
Cloning grade DNA is suitable for use in PCR, cloning reactions, or transformation into E. coli. The purity and amount is not suitable for direct transfections.
Delivery
- Amount 2 µg
- Guaranteed Concentration 100 ng/µl +/- 5 ng/µl
- Pricing $105 USD
- Storage DNA can be stored at 4℃ (short term) or -20℃ (long term).
Terms and Licenses
-
Academic/Nonprofit Terms
-
Industry Terms
- Not Available to Industry
Quality Control
Addgene has verified this plasmid using Next Generation Sequencing. Results are available here
These plasmids were created by your colleagues. Please acknowledge the Principal Investigator, cite the article in which the plasmids were described, and include Addgene in the Materials and Methods of your future publications.
-
For your Materials & Methods section:
pcDNA3-mRuby2 was a gift from Michael Lin (Addgene plasmid # 40260 ; http://n2t.net/addgene:40260 ; RRID:Addgene_40260) -
For your References section:
Improving FRET dynamic range with bright green and red fluorescent proteins. Lam AJ, St-Pierre F, Gong Y, Marshall JD, Cranfill PJ, Baird MA, McKeown MR, Wiedenmann J, Davidson MW, Schnitzer MJ, Tsien RY, Lin MZ. Nat Methods. 2012 Sep 9. doi: 10.1038/nmeth.2171. 10.1038/nmeth.2171 PubMed 22961245