Plant CRISPRi Gene Circuits Kit
(Kit #
1000000276
)
Depositing Lab: Ryan Lister
This kit provides a modular CRISPRi-based plasmid collection enabling programmable and reversible control of gene expression in plants. It includes engineered repressible synthetic promoters (SynPro variants) driving luciferase reporters, compatible sgRNA modules, and plant-expression constructs for dCas9 and dCas9-ZAT10 expression. Together, these components enable rapid quantitative characterization of synthetic gene circuits in transient assays and facilitate the construction of logic gates (e.g. NOT/NOR) that can be layered to create more complex circuits and interface with host regulatory sequences.
This kit will be sent as bacterial glycerol stocks in 96-well plate format.
Original Publication
CRISPRi-based circuits to control gene expression in plants. Khan MA, Herring G, Zhu JY, Oliva M, Fourie E, Johnston B, Zhang Z, Potter J, Pineda L, Pflueger J, Swain T, Pflueger C, Lloyd JPB, Secco D, Small I, Kidd BN, Lister R. Nat Biotechnol. 2025 Mar;43(3):416-430. doi: 10.1038/s41587-024-02236-w. Epub 2024 May 20. PubMed (Link opens in a new window) Article (Link opens in a new window)
Description
Engineering synthetic gene circuits in plants has been limited by the availability of orthogonal, modular genetic parts and compact designs that can be rapidly prototyped in relevant plant contexts. This plasmid collection consists of our CRISPRi-based gene circuit platform (Figure 1), which is programmable, reversible, and highly compact, and can be layered to create more complex circuits. The utility of this system has been demonstrated in Arabidopsis thaliana and shown to function across diverse plant protoplast systems, including Physcomitrium patens, Triticum aestivum, and Brassica napus.
This collection includes two libraries of engineered repressible promoters based on the CaMV 35S and AtTCTP promoters with varying strengths. These engineered promoters, termed “integrators,” contain unique sgRNA target sites positioned just upstream and downstream of the TATA box to enable dCas9-mediated repression. These SynPros are supplied both as luciferase reporter constructs enabling ratiometric, quantitative measurements (Renilla luciferase as the primary output reporter and firefly luciferase as a transfection/normalization control), and as Golden Gate Level 0 parts to accelerate reassembly into new circuit designs. Furthermore, these SynPros can be used to drive expression of genes of interest.
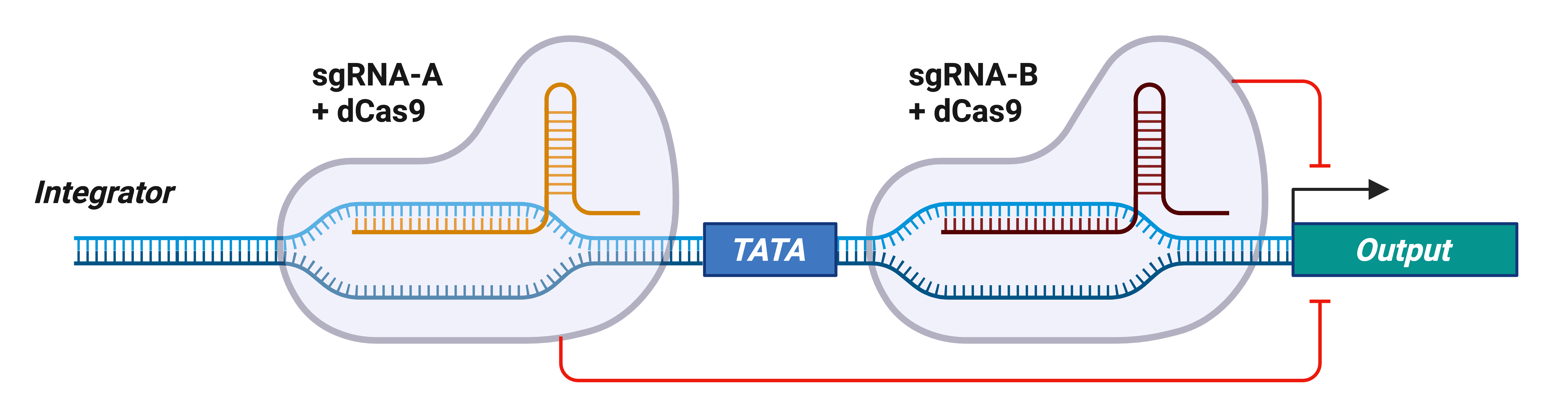
Figure 1: CRISPRi-based synthetic gene circuit system. In the CRISPRi-based gene circuit system, the integrator is an engineered promoter that contains two unique sgRNA target sites positioned just upstream and downstream of the TATA box, enabling dCas9-based repression. The integrator is used for driving expression of an output gene, which can be a reporter or any gene of interest. In the absence of input sgRNAs, the output is expressed. However, in the presence of input sgRNA-A, sgRNA-B, or sgRNA-A + sgRNA-B, dCas9 is recruited to the integrator and the output is repressed, implementing a NOR-gate logic function. These sgRNAs can be expressed from cell-type-specific or inducible promoters to control when and where the output gene is expressed.
To interface external inputs (for example, temperature) with these integrators, this kit provides sgRNA expression modules as well as Csy4 and its cognate site to enable expression of sgRNAs from RNA polymerase II promoters (including cell-type-specific or inducible promoters) for circuit programmability. This collection also includes CRISPRi effectors (dCas9 and dCas9–ZAT10) for programmable control of gene expression in plants.
Kit Documentation
Our step-by-step protocol for designing and testing CRISPRi-based gene circuits in plants (Khan et al. 2026 Nat Protoc. (Link opens in a new window)) includes:
- Design principles
- Protoplast isolation across multiple tissues/species
- 96-well plate transfection
- Dual-luciferase measurement for ratiometric quantification
How to Cite this Kit
These plasmids were created by your colleagues. Please acknowledge the Principal Investigator, cite the article in which they were created, and include Addgene in the Materials and Methods of your future publications.
For your Materials and Methods section:
"The Plant CRISPRi Gene Circuits Kit was a gift from Ryan Lister (Addgene kit #1000000276)."
For your Reference section:
CRISPRi-based circuits to control gene expression in plants. Khan MA, Herring G, Zhu JY, Oliva M, Fourie E, Johnston B, Zhang Z, Potter J, Pineda L, Pflueger J, Swain T, Pflueger C, Lloyd JPB, Secco D, Small I, Kidd BN, Lister R. Nat Biotechnol. 2025 Mar;43(3):416-430. doi: 10.1038/s41587-024-02236-w. Epub 2024 May 20. PubMed (Link opens in a new window) Article (Link opens in a new window)
Plant CRISPRi Gene Circuits Kit - #1000000276
- Resistance Color Key
Each circle corresponds to a specific antibiotic resistance in the kit plate map wells.
- Inventory
Searchable and sortable table of all plasmids in kit. The Well column lists the plasmid well location in its plate. The Plasmid column links to a plasmid's individual web page.
- Kit Plate Map
96-well plate map for plasmid layout. Hovering over a well reveals the plasmid name, while clicking on a well opens the plasmid page.
Resistance Color Key
| Ampicillin | |
| Kanamycin | |
| Spectinomycin |
Inventory
| Well | Plasmid | Resistance |
|---|---|---|
| A / 1 | LLP947 |
|
| A / 2 | LLP948 |
|
| A / 3 | LLP949 |
|
| A / 4 | LLP950 |
|
| A / 5 | LLP951 |
|
| A / 6 | LLP952 |
|
| A / 7 | LLP953 |
|
| A / 8 | LLP954 |
|
| A / 9 | LLP955 |
|
| A / 10 | LLP956 |
|
| A / 11 | LLP957 |
|
| A / 12 | LLP958 |
|
| B / 1 | LLP959 |
|
| B / 2 | LLP960 |
|
| B / 3 | LLP961 |
|
| B / 4 | LLP962 |
|
| B / 5 | LLP963 |
|
| B / 6 | LLP964 |
|
| B / 7 | LLP965 |
|
| B / 8 | LLP966 |
|
| B / 9 | LLP967 |
|
| B / 10 | LLP968 |
|
| B / 11 | LLP969 |
|
| B / 12 | LLP970 |
|
| C / 1 | LLP971 |
|
| C / 2 | LLP972 |
|
| C / 3 | LLP973 |
|
| C / 4 | LLP974 |
|
| C / 5 | LLP975 |
|
| C / 6 | LLP976 |
|
| C / 7 | LLP977 |
|
| C / 8 | LLP978 |
|
| C / 9 | LLP979 |
|
| C / 10 | LLP980 |
|
| C / 11 | LLP1015 |
|
| C / 12 | LLP1016 |
|
| D / 1 | LLP1017 |
|
| D / 2 | LLP1023 |
|
| D / 3 | LLP1024 |
|
| D / 4 | LLP1014 |
|
| D / 5 | LLP1020 |
|
| D / 6 | LLP981 |
|
| D / 7 | LLP982 |
|
| D / 8 | LLP983 |
|
| D / 9 | LLP984 |
|
| D / 10 | LLP985 |
|
| D / 11 | LLP986 |
|
| D / 12 | LLP987 |
|
| E / 1 | LLP988 |
|
| E / 2 | LLP989 |
|
| E / 3 | LLP990 |
|
| E / 4 | LLP991 |
|
| E / 5 | LLP992 |
|
| E / 6 | LLP993 |
|
| E / 7 | LLP994 |
|
| E / 8 | LLP995 |
|
| E / 9 | LLP996 |
|
| E / 10 | LLP997 |
|
| E / 11 | LLP998 |
|
| E / 12 | LLP999 |
|
| F / 1 | LLP1000 |
|
| F / 2 | LLP1001 |
|
| F / 3 | LLP1002 |
|
| F / 4 | LLP1003 |
|
| F / 5 | LLP1004 |
|
| F / 6 | LLP1005 |
|
| F / 7 | LLP1006 |
|
| F / 8 | LLP1007 |
|
| F / 9 | LLP1008 |
|
| F / 10 | LLP1009 |
|
| F / 11 | LLP1010 |
|
| F / 12 | LLP1011 |
|
| G / 1 | LLP1012 |
|
| G / 2 | LLP1013 |
|
| G / 3 | LLP1018 |
|
| G / 4 | LLP1019 |
|
| G / 5 | LLP1021 |
|
| G / 6 | LLP1022 |
|


