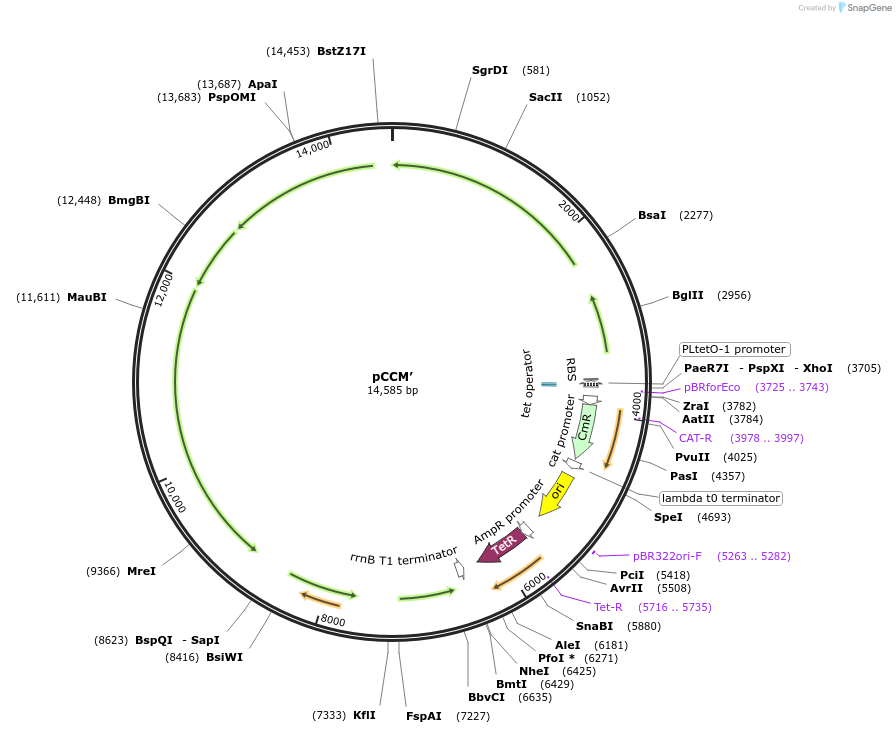
pCCM’
(Plasmid
#162709)
-
PurposepCCM reconstructed after selection for CCMB1 growth on glycerol under ambient air
-
Depositing Lab
-
Sequence Information
Ordering
| Item | Catalog # | Description | Quantity | Price (USD) | |
|---|---|---|---|---|---|
| Plasmid | 162709 | Standard format: Plasmid sent in bacteria as agar stab | 1 | $94 | |
Backbone
-
Vector backbonepFA31 (derived from pZA31 expressing under PLtet0-1 and constitutive tetR)
- Backbone size w/o insert (bp) 2980
- Total vector size (bp) 14585
-
Modifications to backboneThe p15A origin was replaced with high-copy colE1 origin.
-
Vector typeBacterial Expression
Growth in Bacteria
-
Bacterial Resistance(s)Chloramphenicol, 25 μg/mL
-
Growth Temperature37°C
-
Growth Strain(s)DH5alpha
-
Copy numberHigh Copy
Gene/Insert
-
Gene/Insert nameSecond CCM operon cloned from H. neapolitanus
-
SpeciesHalothiobacillus neapolitanus
-
Insert Size (bp)11605
-
Mutationp15A origin replaced with high-copy colE1 origin
- Promoter PLtet0-1 promoter
Cloning Information
- Cloning method Gibson Cloning
- 5′ sequencing primer ggaacctcttacgtgccgatc
- 3′ sequencing primer GCGTTCACCGACAAACAACA
- (Common Sequencing Primers)
Resource Information
Terms and Licenses
-
Academic/Nonprofit Terms
-
Industry Terms
- Not Available to Industry
Trademarks:
- Zeocin® is an InvivoGen trademark.
Depositor Comments
pCCM reconstructed after selection for CCMB1 growth on glycerol under ambient air. Please note that the I145V mutation in TetR does not impact plasmid function.
These plasmids were created by your colleagues. Please acknowledge the Principal Investigator, cite the article in which the plasmids were described, and include Addgene in the Materials and Methods of your future publications.
-
For your Materials & Methods section:
pCCM’ was a gift from David Savage (Addgene plasmid # 162709 ; http://n2t.net/addgene:162709 ; RRID:Addgene_162709) -
For your References section:
Functional reconstitution of a bacterial CO2 concentrating mechanism in E. coli. Flamholz AI, Dugan E, Blikstad C, Gleizer S, Ben-Nissan R, Amram S, Antonovsky N, Ravishankar S, Noor E, Bar-Even A, Milo R, Savage D. Elife. 2020 Oct 21;9. pii: 59882. doi: 10.7554/eLife.59882. 10.7554/eLife.59882 PubMed 33084575