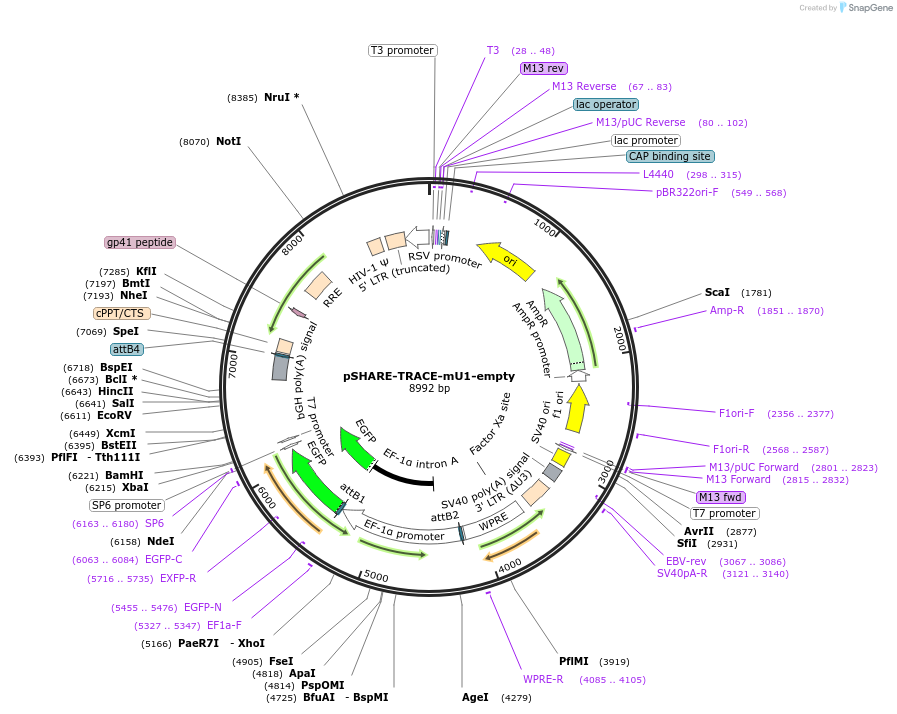
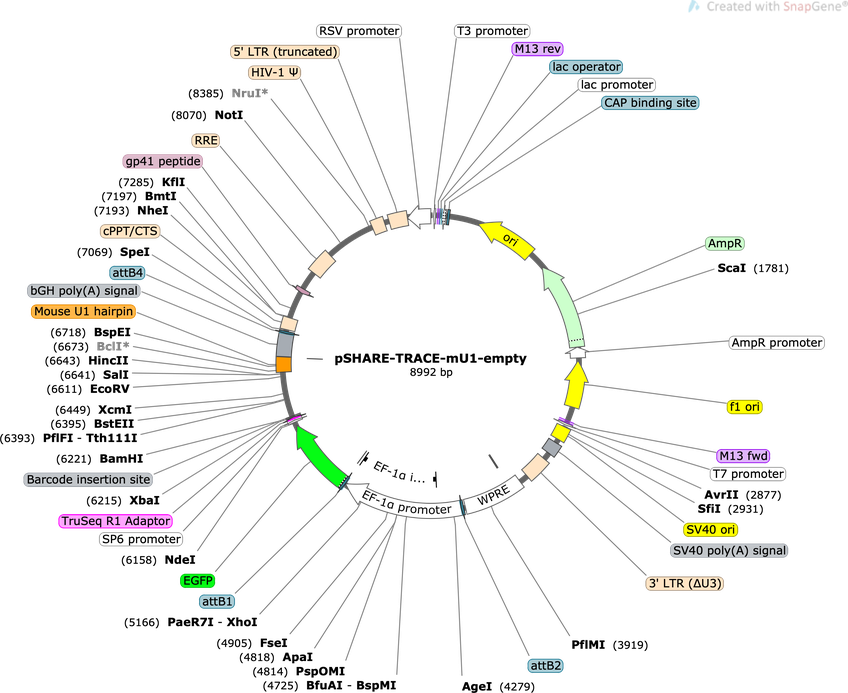
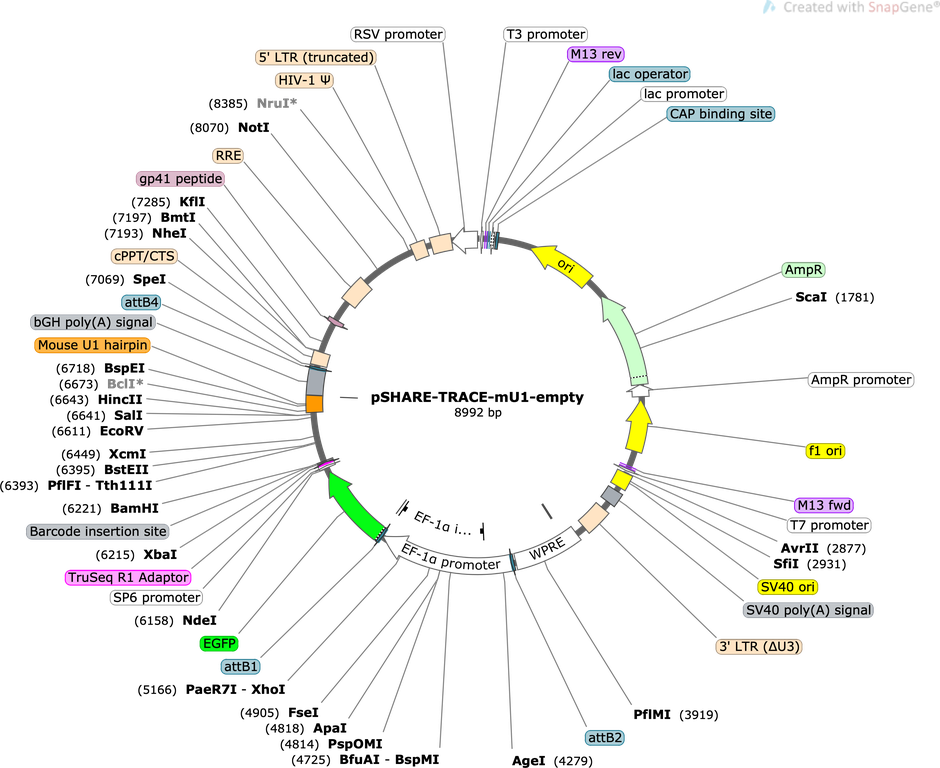
pSHARE-TRACE-mU1-empty
(Plasmid
#253332)
-
Purpose(Empty Backbone) Lineage tracing vector compatible with SHARE-seq, modified from Weinreb at al 2020. Expresses GFP from EF1a promoter with a mouse U1 inserted into 3' UTR to improve nuclear localization.
-
Depositing Lab
-
Sequence Information
Ordering
| Item | Catalog # | Description | Quantity | Price (USD) | |
|---|---|---|---|---|---|
| Plasmid | 253332 | Standard format: Plasmid sent in bacteria as agar stab | 1 | $89 | |
Backbone
-
Vector backbonepSHARE-TRACE-mU1-empty
-
Backbone manufacturerThis publication
- Backbone size (bp) 8991
-
Modifications to backboneThe pLARRY empty vector (Addgene #140025) was first modified to insert a TruSeq sequencing adaptor (ACACTCTTTCCCTACACGACGCTCTTCCGATCT) upstream of the barcode insertion site to allow for direct amplification. Additional sequence, including a mouse U1 hairpin, was introduced downstream of the barcode site to promote nuclear translocation of RNA transcripts and more efficient SHARE-seq capture.
-
Vector typeMammalian Expression, Lentiviral
- Promoter EF1a
-
Selectable markersGFP
Growth in Bacteria
-
Bacterial Resistance(s)Ampicillin, 100 μg/mL
-
Growth Temperature37°C
-
Growth Strain(s)NEB Stable
-
Copy numberUnknown
Cloning Information
- Cloning method Gibson Cloning
Resource Information
Terms and Licenses
-
Academic/Nonprofit Terms
-
Industry Terms
- Not Available to Industry
Trademarks:
- Zeocin® is an InvivoGen trademark.
Depositor Comments
Please visit https://doi.org/10.1101/2025.02.13.638099 for bioRxiv preprint.
These plasmids were created by your colleagues. Please acknowledge the Principal Investigator, cite the article in which the plasmids were described, and include Addgene in the Materials and Methods of your future publications.
-
For your Materials & Methods section:
pSHARE-TRACE-mU1-empty was a gift from Jason Buenrostro (Addgene plasmid # 253332 ; http://n2t.net/addgene:253332 ; RRID:Addgene_253332) -
For your References section:
Clonal memory of colitis accumulates and promotes tumor growth. Nagaraja S, Ojeda-Miron L, Zhang R, Oreskovic E, Hu Y, Zeve D, Sharma K, Hyman RR, Zhang Q, Castillo A, Breault DT, Yilmaz OH, Buenrostro JD. bioRxiv [Preprint]. 2025 Feb 17:2025.02.13.638099. doi: 10.1101/2025.02.13.638099. 10.1101/2025.02.13.638099 PubMed 40027722