-
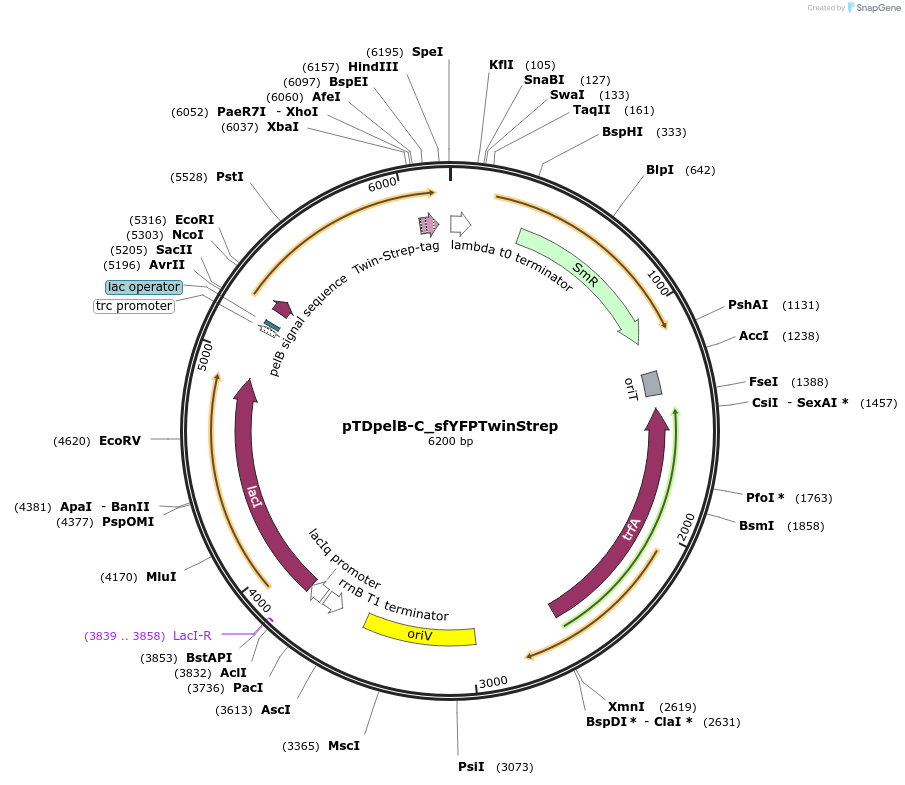
Purposeexpresses P.putida codon optimized PelB-sfYFP-Twin-Strep fusion protein for periplasmic translocation via PelB signal sequence
-
Depositing Lab
-
Publication
-
Sequence Information
Ordering
| Item | Catalog # | Description | Quantity | Price (USD) | |
|---|---|---|---|---|---|
| Plasmid | 45944 | Standard format: Plasmid sent in bacteria as agar stab | 1 | $94 | |
Backbone
-
Vector backbonepTDpelB-CTwinStrep
-
Backbone manufacturerDammeyer et al. 2013
-
Modifications to backbonecloning of synthetic sfYFP insert into EcoRI and XbaI sites of pTDpelB-CTwinStrep for N-termina fusion to PelB export signal and C-terminal fusion to TwinStrep-tag
-
Vector typeBacterial Expression
Growth in Bacteria
-
Bacterial Resistance(s)Streptomycin, 50 μg/mL
-
Growth Temperature37°C
-
Growth Strain(s)DH5alpha
-
Growth instructionsFor growth in P.putida KT2440 (30C); on plates use spectinomycin in addition.
-
Copy numberLow Copy
Gene/Insert
-
Gene/Insert namesynthetic sfYFP (codon usage adapted to P.putida KT2440)
-
SpeciesAequorea victoria
- Promoter lacIq/Ptrc
-
Tags
/ Fusion Proteins
- Twin-Strep-tag (C terminal on backbone)
- PelB export sequence (N terminal on backbone)
Cloning Information
- Cloning method Restriction Enzyme
- 5′ cloning site EcoRI (not destroyed)
- 3′ cloning site XbaI (not destroyed)
- 5′ sequencing primer TGTGTGGAATTGTGAGCGG
- 3′ sequencing primer ACTTTGTTTTAGGGCGACTG
- (Common Sequencing Primers)
Terms and Licenses
-
Academic/Nonprofit Terms
-
Industry Terms
- Not Available to Industry
Trademarks:
- Zeocin® is an InvivoGen trademark.
Depositor Comments
This plasmid was tested in the Gram-negative soil bacterium Pseudomonas putida KT2440 and Escherichia coli K12 and is especially suited for protein production, affinity purification, protein complex copurification with SPINE (Strep Protein Interaction Experiments) or (co-)localization studies. Due to the broad host range of the RK2 origin of replication, the plasmid facilitates experimental verification of hypothetical proteins and protein production yield assessment in different expression hosts possibly including new isolates.
The Supplementary Table S1 in the following publication lists approximately 30 strains in which the RK2 origin of replication should be functional. Silva-Rocha et al., The Standard European Vector Architecture (SEVA): a coherent platform for the analysis and deployment of complex prokaryotic phenotypes. Nucleic Acids Research 2013, 41:D666-675. http://nar.oxfordjournals.org/content/41/D1/D666.long
These plasmids were created by your colleagues. Please acknowledge the Principal Investigator, cite the article in which the plasmids were described, and include Addgene in the Materials and Methods of your future publications.
-
For your Materials & Methods section:
pTDpelB-C_sfYFPTwinStrep was a gift from Thorben Dammeyer (Addgene plasmid # 45944 ; http://n2t.net/addgene:45944 ; RRID:Addgene_45944) -
For your References section:
Broad host range vectors for expression of proteins with (Twin-) Strep-tag, His-tag and engineered, export optimized yellow fluorescent protein. Dammeyer T, Timmis KN, Tinnefeld P. Microb Cell Fact. 2013 May 20;12(1):49. 10.1186/1475-2859-12-49 PubMed 23687945