-
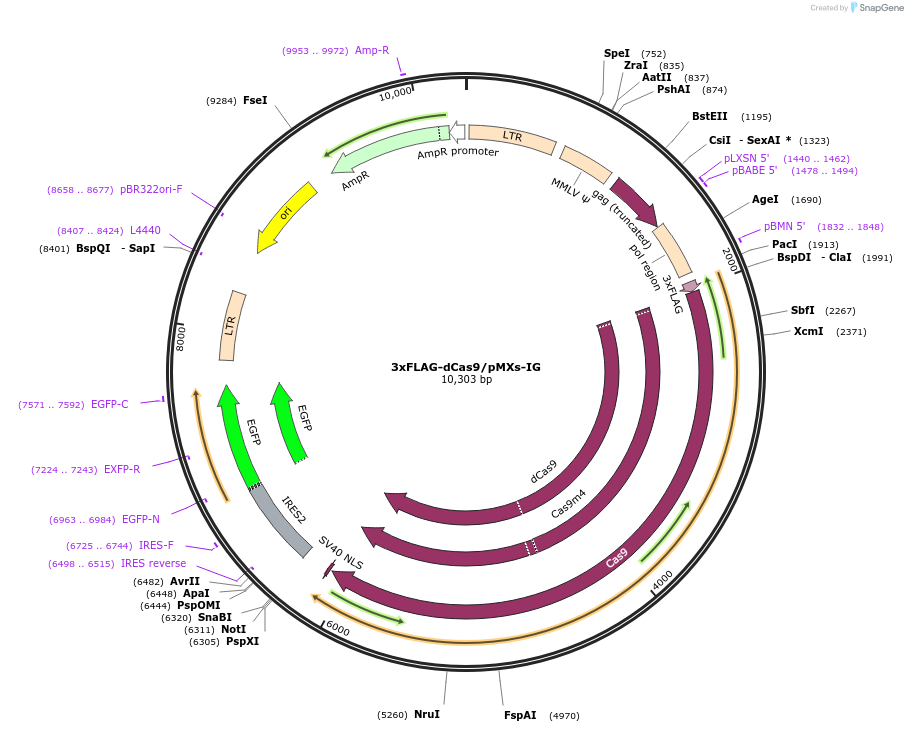
PurposeExpresses 3xFLAG-dCas9 in mammalian cells for enChIP analysis to purify specific genomic regions of interest.
-
Depositing Lab
-
Sequence Information
Ordering
| Item | Catalog # | Description | Quantity | Price (USD) | |
|---|---|---|---|---|---|
| Plasmid | 51258 | Standard format: Plasmid sent in bacteria as agar stab | 1 | $94 | |
Backbone
-
Vector backbonepMXs-IG
-
Backbone manufacturerToshio Kitamura
- Backbone size w/o insert (bp) 6091
- Total vector size (bp) 10303
-
Vector typeMammalian Expression, Retroviral, CRISPR
-
Selectable markersenhanced GFP
Growth in Bacteria
-
Bacterial Resistance(s)Ampicillin, 50 μg/mL
-
Growth Temperature30°C
-
Growth Strain(s)NEB Stable
-
Growth instructionsslow growing, culture for 18hrs.
-
Copy numberLow Copy
Gene/Insert
-
Gene/Insert name3xFLAG-dCas9
-
SpeciesSynthetic
-
Insert Size (bp)4212
-
Mutationhuman codon-optimized, D10A + H840A
- Promoter LTR
-
Tags
/ Fusion Proteins
- 3xFLAG tag (N terminal on insert)
- NLS (nuclear localization signal) (C terminal on insert)
Cloning Information
- Cloning method Restriction Enzyme
- 5′ cloning site Pac I (not destroyed)
- 3′ cloning site Not I (not destroyed)
- 5′ sequencing primer ggtggaccatcctctagact
- 3′ sequencing primer AAACGCACACCGGCCTTATT
- (Common Sequencing Primers)
Resource Information
-
Addgene Notes
Terms and Licenses
-
Academic/Nonprofit Terms
-
Industry Terms
- Not Available to Industry
Trademarks:
- Zeocin® is an InvivoGen trademark.
Depositor Comments
This plasmid is prone to recombination. It is recommended that recipient scientists screen multiple colonies by diagnostic digest prior to use.
The coding sequence of 3xFLAG-dCas9 can be cleaved with Pac I and Not I.
Construction strategy of gRNA retroviral vectors
1. Cleave gBlock from a gRNA vector constructed using gRNA cloning vector (Addgene #41824) with appropriate restriction enzymes (eg. [Xho I + Hind III], EcoR I).
2. Insert the cleaved gBlock into pSIR-based self-inactivating retroviral vectors.
Vectors & Sites of insertion
pSIR-neo (Addgene #51128): eg. [Xho I + Hind III]
pSIR-GFP (Addgene #51134): eg. [Xho I + Hind III], EcoR I
pSIR-DsRed-Express2 (Addgene #51135): eg. [Xho I + Hind III], EcoR I
pSIR-hCD2 (Addgene #51143): eg. EcoR I
These plasmids were created by your colleagues. Please acknowledge the Principal Investigator, cite the article in which the plasmids were described, and include Addgene in the Materials and Methods of your future publications.
-
For your Materials & Methods section:
3xFLAG-dCas9/pMXs-IG was a gift from Hodaka Fujii (Addgene plasmid # 51258 ; http://n2t.net/addgene:51258 ; RRID:Addgene_51258) -
For your References section:
Identification of Proteins Associated with an IFNgamma-Responsive Promoter by a Retroviral Expression System for enChIP Using CRISPR. Fujita T, Fujii H. PLoS One. 2014 Jul 22;9(7):e103084. doi: 10.1371/journal.pone.0103084. eCollection 2014. 10.1371/journal.pone.0103084 PubMed 25051498