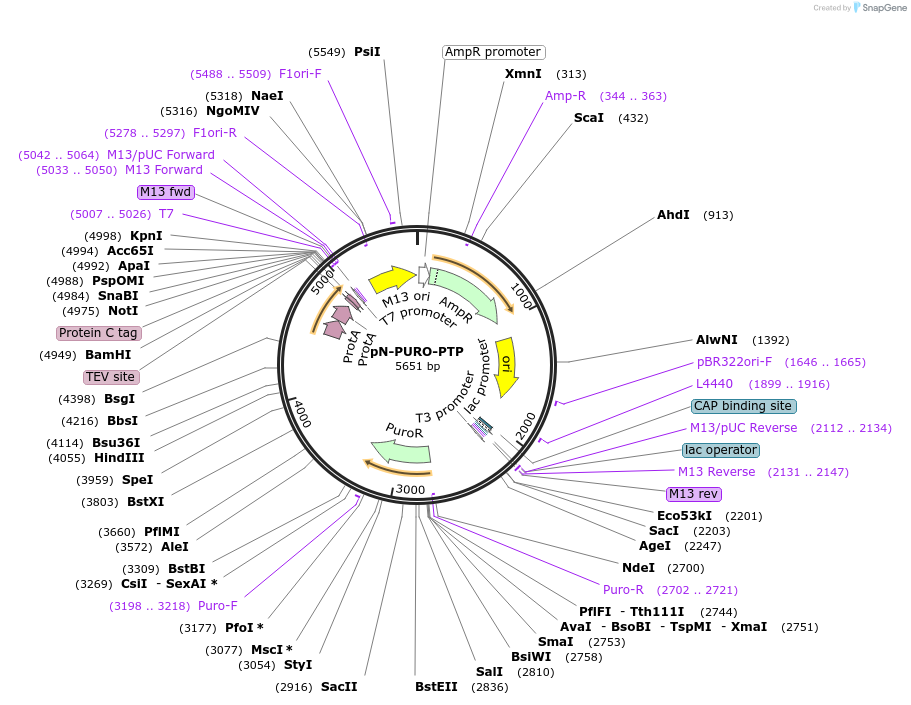
pN-PURO-PTP
(Plasmid
#51711)
-
Purpose(Empty Backbone) contains an N-terminal tandem affinity purification tag (relevant to any system) within a construct designed for genome integration into trypanosomes.
-
Depositing Lab
-
Publication
-
Sequence Information
Ordering
| Item | Catalog # | Description | Quantity | Price (USD) | |
|---|---|---|---|---|---|
| Plasmid | 51711 | Standard format: Plasmid sent in bacteria as agar stab | 1 | $94 | |
Backbone
-
Vector backbonepC-PTP-NEO (Addgene plasmid #51710)
-
Modifications to backboneThe resistance marker cassette was modified by replacing NEO-R by the coding region of the puromycin N-acetyltransferase gene (PURO-R).
-
Vector typeT. brucei genome integration vectors
-
Selectable markersPuromycin
-
Tag
/ Fusion Protein
- PTP (ProtC-TEV-ProtA) (N terminal on backbone)
Growth in Bacteria
-
Bacterial Resistance(s)Ampicillin, 100 μg/mL
-
Growth Temperature37°C
-
Growth Strain(s)DH5alpha
-
Copy numberUnknown
Cloning Information
- Cloning method Restriction Enzyme
- 5′ sequencing primer M13-F20
- 3′ sequencing primer M13R
- (Common Sequencing Primers)
Terms and Licenses
-
Academic/Nonprofit Terms
-
Industry Terms
- Not Available to Industry
Trademarks:
- Zeocin® is an InvivoGen trademark.
Depositor Comments
The PTP tag is a derivative of the original TAP tag and has CBP replaced by ProtC.
The PTP cassette is composed of 434 bp of 5′ flank from the TbRPA2 gene, encoding the second-largest subunit of RNA polymerase I, a translation initiation codon, the PTP tag coding sequence, and unique restriction sites for fusing target sequences to the tag.
These plasmids were created by your colleagues. Please acknowledge the Principal Investigator, cite the article in which the plasmids were described, and include Addgene in the Materials and Methods of your future publications.
-
For your Materials & Methods section:
pN-PURO-PTP was a gift from Arthur Gunzl (Addgene plasmid # 51711 ; http://n2t.net/addgene:51711 ; RRID:Addgene_51711) -
For your References section:
Highly efficient tandem affinity purification of trypanosome protein complexes based on a novel epitope combination. Schimanski B, Nguyen TN, Gunzl A. Eukaryot Cell. 2005 Nov;4(11):1942-50. 10.1128/EC.4.11.1942-1950.2005 PubMed 16278461