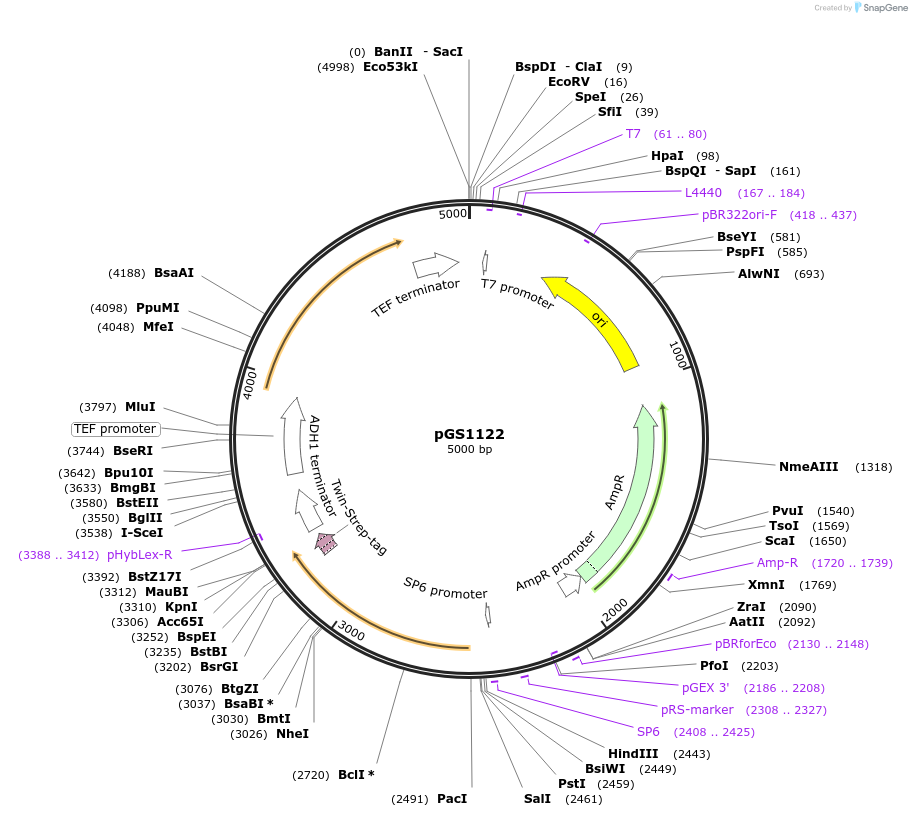
pGS1122
(Plasmid
#63790)
-
PurposeVector to create a cassette for homologous recombination in S. cerevisiae; adds a C-terminal eSapphire Twin-Strep tag followed by URA3 selection marker
-
Depositing Labs
-
Publication
-
Sequence Information
Ordering
| Item | Catalog # | Description | Quantity | Price (USD) | |
|---|---|---|---|---|---|
| Plasmid | 63790 | Standard format: Plasmid sent in bacteria as agar stab | 1 | $94 | |
Backbone
-
Vector backbonepGS1222
- Backbone size w/o insert (bp) 5000
- Total vector size (bp) 5000
-
Vector typeE. coli cloning vector
-
Selectable markersURA3
Growth in Bacteria
-
Bacterial Resistance(s)Ampicillin, 100 μg/mL
-
Growth Temperature37°C
-
Growth Strain(s)DH5alpha
-
Copy numberHigh Copy
Gene/Insert
-
Gene/Insert nameeSapphire-Twin-Strep tag followed by URA3
-
SpeciesSynthetic
- Promoter none
-
Tag
/ Fusion Protein
- eSapphire-Twin-Strep tag (C terminal on backbone)
Cloning Information
- Cloning method Restriction Enzyme
- 5′ cloning site SalI (not destroyed)
- 3′ cloning site BsrGI/Bsp1407I (not destroyed)
- 5′ sequencing primer CTGGCTTAACTATGCGGCATCAG
- 3′ sequencing primer gcaacctgacctacaggaaagag
- (Common Sequencing Primers)
Resource Information
-
A portion of this plasmid was derived from a plasmid made byMark A. Sheff and Kurt S. Thorn
Terms and Licenses
-
Academic/Nonprofit Terms
-
Industry Terms
- Not Available to Industry
Trademarks:
- Zeocin® is an InvivoGen trademark.
Depositor Comments
The sequence of the eSapphire gene has been optimized for S. cerevisiae and generated by gene synthesis. The Twin-Strep tag sequence is adapted from the sequence of commercial Twin-Strep tag plasmids offered by IBA, Goettingen, Germany.
These plasmids were created by your colleagues. Please acknowledge the Principal Investigator, cite the article in which the plasmids were described, and include Addgene in the Materials and Methods of your future publications.
-
For your Materials & Methods section:
pGS1122 was a gift from Monika Golas & Bjoern Sander (Addgene plasmid # 63790 ; http://n2t.net/addgene:63790 ; RRID:Addgene_63790) -
For your References section:
Strep-tag II and Twin-Strep Based Cassettes for Protein Tagging by Homologous Recombination and Characterization of Endogenous Macromolecular Assemblies in Saccharomyces cerevisiae. Rai J, Pemmasani JK, Voronovsky A, Jensen IS, Manavalan A, Nyengaard JR, Golas MM, Sander B. Mol Biotechnol. 2014 Jun 27. 10.1007/s12033-014-9778-5 PubMed 24969434