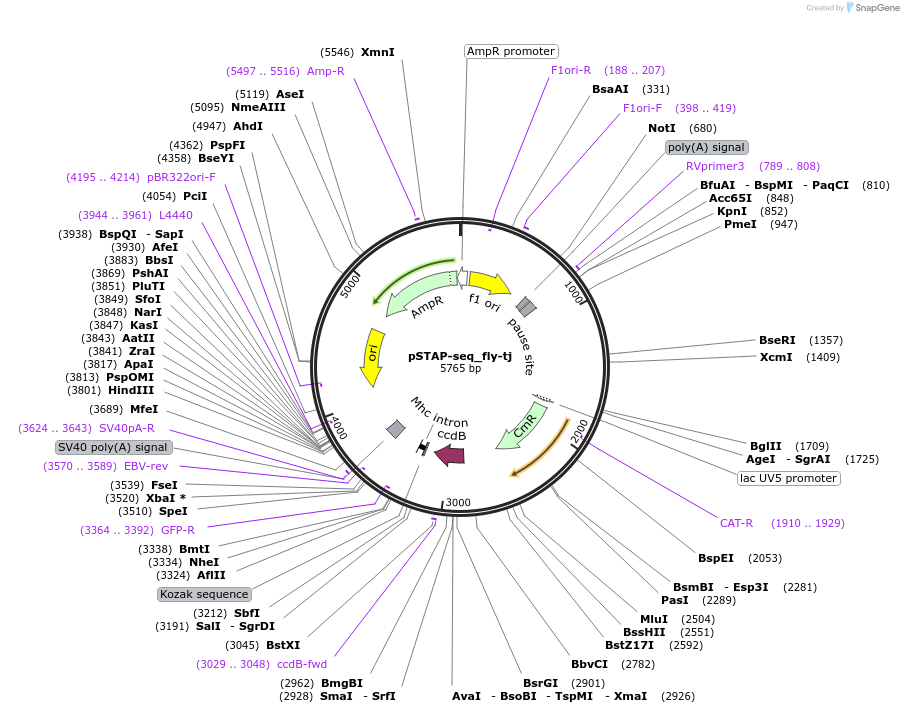
pSTAP-seq_fly-tj
(Plasmid
#86386)
-
PurposeVector to measure enhancer responsiveness of candidates from a genomic library (cloned instead of the core promoter) by determining the abundance of transcripts originating from each candidate.
-
Depositing Lab
-
Sequence Information
Ordering
| Item | Catalog # | Description | Quantity | Price (USD) | |
|---|---|---|---|---|---|
| Plasmid | 86386 | Standard format: Plasmid sent in bacteria as agar stab | 1 | $94 | |
Backbone
-
Vector backbonepGL3
-
Backbone manufacturerPromega
- Backbone size w/o insert (bp) 3426
-
Vector typeSTAP-seq screening vector
Growth in Bacteria
-
Bacterial Resistance(s)Chloramphenicol and Ampicillin, 25 & 100 μg/mL
-
Growth Temperature37°C
-
Growth Strain(s)ccdB Survival
-
Copy numberHigh Copy
Gene/Insert
-
Gene/Insert nameD. melanogaster tj enhancer
-
SpeciesD. melanogaster (fly)
-
Insert Size (bp)856
Cloning Information
- Cloning method Restriction Enzyme
- 5′ cloning site unknown (unknown if destroyed)
- 3′ cloning site unknown (unknown if destroyed)
- 5′ sequencing primer RVprimer3
- (Common Sequencing Primers)
Resource Information
Terms and Licenses
-
Academic/Nonprofit Terms
-
Industry Terms
- Not Available to Industry
Trademarks:
- Zeocin® is an InvivoGen trademark.
Depositor Comments
to consruct the STAP-seq screening vector from pGL3 Promoter we replaced the sequence between the BglII and FseI sites with the following sequence, containing a ccdB suicide gene next to the Chloramphenicole resistance gene flanked by homology arms (used for cloning the candidates during library generation), an intron (mhc16), an ORF (truncated sgGFP, Qbiogene, Inc)
These plasmids were created by your colleagues. Please acknowledge the Principal Investigator, cite the article in which the plasmids were described, and include Addgene in the Materials and Methods of your future publications.
-
For your Materials & Methods section:
pSTAP-seq_fly-tj was a gift from Alexander Stark (Addgene plasmid # 86386 ; http://n2t.net/addgene:86386 ; RRID:Addgene_86386) -
For your References section:
Genome-wide assessment of sequence-intrinsic enhancer responsiveness at single-base-pair resolution. Arnold CD, Zabidi MA, Pagani M, Rath M, Schernhuber K, Kazmar T, Stark A. Nat Biotechnol. 2016 Dec 26. doi: 10.1038/nbt.3739. 10.1038/nbt.3739 PubMed 28024147