-
PurposeEGFP-tagged constructs to monitor the aggregation state of the sensors and the ability of cells to solubilize or degrade the aggregated proteins
-
Depositing Lab
-
Sequence Information
Ordering
| Item | Catalog # | Description | Quantity | Price (USD) | |
|---|---|---|---|---|---|
| Plasmid | 90170 | Standard format: Plasmid sent in bacteria as agar stab | 1 | $94 | |
Backbone
-
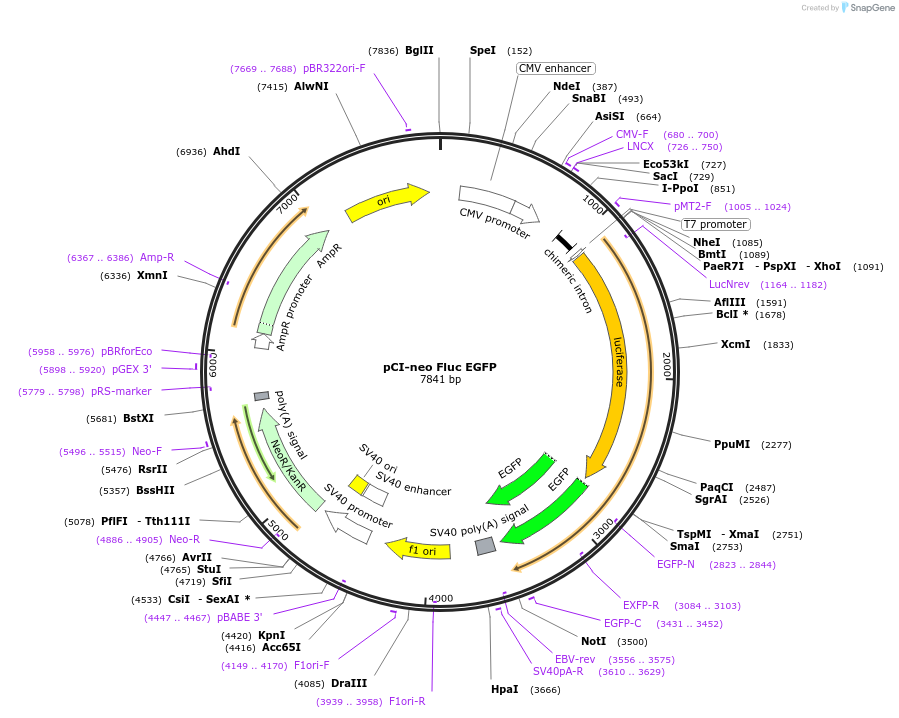
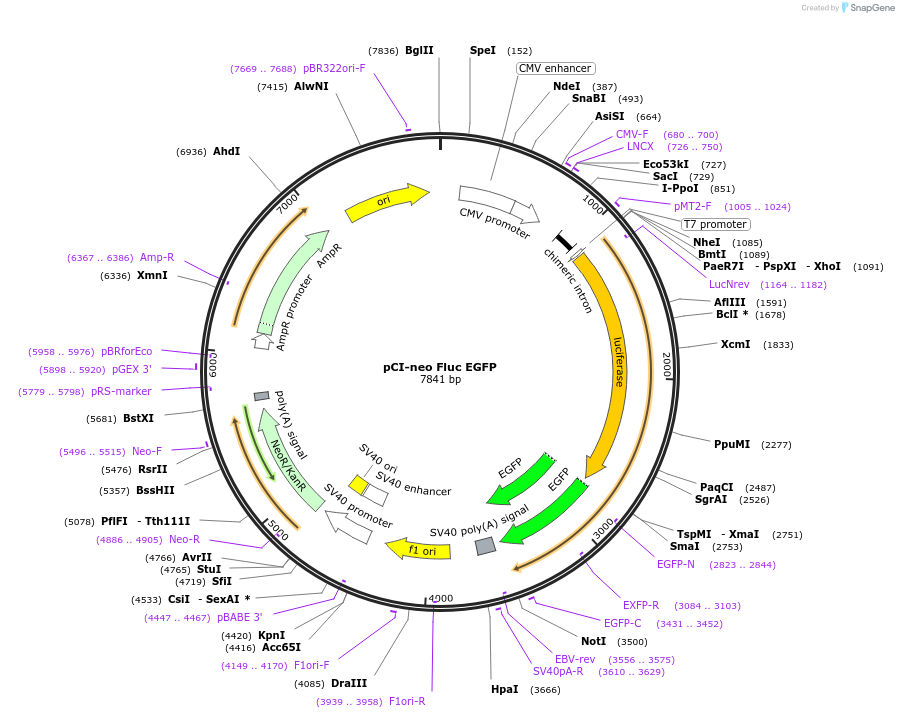
Vector backbonepCIneo
-
Backbone manufacturerpromega
- Backbone size w/o insert (bp) 5472
- Total vector size (bp) 7860
-
Vector typeMammalian Expression
-
Selectable markersNeomycin (select with G418)
Growth in Bacteria
-
Bacterial Resistance(s)Ampicillin, 100 μg/mL
-
Growth Temperature37°C
-
Growth Strain(s)DH5alpha
-
Copy numberUnknown
Gene/Insert
-
Gene/Insert nameFLuc EGFP
-
Speciesfirefly
-
Insert Size (bp)2373
Cloning Information
- Cloning method Restriction Enzyme
- 5′ cloning site XmaI (unknown if destroyed)
- 3′ cloning site NotI (unknown if destroyed)
- 5′ sequencing primer T7
- 3′ sequencing primer T3
- (Common Sequencing Primers)
Resource Information
-
Articles Citing this Plasmid
Terms and Licenses
-
Academic/Nonprofit Terms
-
Industry Terms
- Not Available to Industry
Trademarks:
- Zeocin® is an InvivoGen trademark.
These plasmids were created by your colleagues. Please acknowledge the Principal Investigator, cite the article in which the plasmids were described, and include Addgene in the Materials and Methods of your future publications.
-
For your Materials & Methods section:
pCI-neo Fluc EGFP was a gift from Franz-Ulrich Hartl (Addgene plasmid # 90170 ; http://n2t.net/addgene:90170 ; RRID:Addgene_90170) -
For your References section:
Firefly luciferase mutants as sensors of proteome stress. Gupta R, Kasturi P, Bracher A, Loew C, Zheng M, Villella A, Garza D, Hartl FU, Raychaudhuri S. Nat Methods. 2011 Sep 4;8(10):879-84. doi: 10.1038/nmeth.1697. 10.1038/nmeth.1697 PubMed 21892152