-
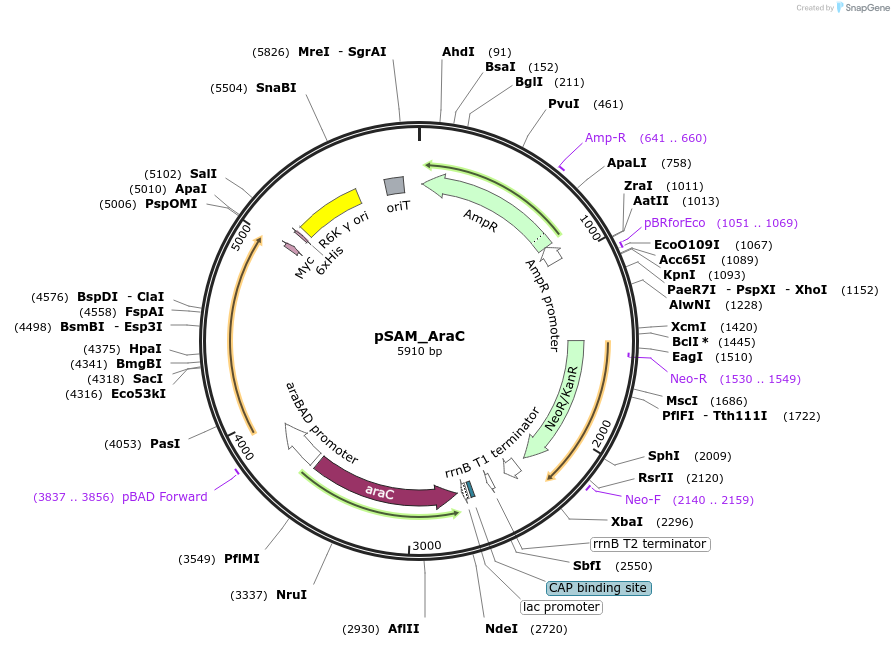
PurposeMariner transposon mutagenesis & sequencing vector - Transposon contains KanR and Illumina sequencing adapters. Arabinose promoter expresses transposase. Ampicillin resistance on backbone.
-
Depositing Lab
-
Sequence Information
Ordering
| Item | Catalog # | Description | Quantity | Price (USD) | |
|---|---|---|---|---|---|
| Plasmid | 91569 | Standard format: Plasmid sent in bacteria as agar stab | 1 | $94 | |
Backbone
-
Vector backbonepSAM_Ec
-
Backbone manufacturerN/A
- Backbone size w/o insert (bp) 4559
- Total vector size (bp) 5906
-
Modifications to backbonePut arabinose inducible promoter before transposase
-
Vector typeBacterial Expression
Growth in Bacteria
-
Bacterial Resistance(s)Ampicillin and Kanamycin, 100 & 50 μg/mL
-
Growth Temperature37°C
-
Growth Strain(s)S17-1λpir
-
Growth instructionsAdd 10 mM glucose to repress transposase. S17-1λpir may be EndA+ so extra purification steps may be necessary during miniprep.
-
Copy numberLow Copy
Gene/Insert
-
Gene/Insert nameAraC/PBAD
-
SpeciesE. coli
-
Insert Size (bp)1200
Cloning Information
- Cloning method Restriction Enzyme
- 5′ cloning site NdeI (unknown if destroyed)
- 3′ cloning site NcoI (unknown if destroyed)
- 5′ sequencing primer TTATGACAACTTGACGGCTA
- 3′ sequencing primer ATGCCATAGCATTTTTATCC
- (Common Sequencing Primers)
Resource Information
-
Articles Citing this Plasmid
Terms and Licenses
-
Academic/Nonprofit Terms
-
Industry Terms
- Not Available to Industry
Trademarks:
- Zeocin® is an InvivoGen trademark.
Depositor Comments
Mariner himar1C9 transposase driven by an arabinose promoter on an R6K gamma suicide plasmid. The transposon allows identification of fitness factors by Illumina sequencing. This transposon will randomly insert a kanamycin resistance cassette into genomic TA sites. Ampicillin resistance is contained on the plasmid backbone.
For further background, see:
Goodman, Andrew L., Meng Wu, and Jeffrey I. Gordon. “Identifying Microbial Fitness Determinants by Insertion Sequencing (INSeq) Using Genome-Wide Transposon Mutant Libraries.” Nature Protocols 6.12 (2011): 1969–1980. PMC. Web. 11 Apr. 2017.
Wiles, Travis J. et al. “Combining Quantitative Genetic Footprinting and Trait Enrichment Analysis to Identify Fitness Determinants of a Bacterial Pathogen.” Ed. Danielle A. Garsin. PLoS Genetics 9.8 (2013): e1003716. PMC. Web. 11 Apr. 2017.
These plasmids were created by your colleagues. Please acknowledge the Principal Investigator, cite the article in which the plasmids were described, and include Addgene in the Materials and Methods of your future publications.
-
For your Materials & Methods section:
pSAM_AraC was a gift from Harry Mobley (Addgene plasmid # 91569 ; http://n2t.net/addgene:91569 ; RRID:Addgene_91569) -
For your References section:
Genome-wide transposon mutagenesis of Proteus mirabilis: Essential genes, fitness factors for catheter-associated urinary tract infection, and the impact of polymicrobial infection on fitness requirements. Armbruster CE, Forsyth-DeOrnellas V, Johnson AO, Smith SN, Zhao L, Wu W, Mobley HLT. PLoS Pathog. 2017 Jun 14;13(6):e1006434. doi: 10.1371/journal.ppat.1006434. PPATHOGENS-D-17-00798 [pii] PubMed 28614382