We narrowed to 17,788 results for: por
-
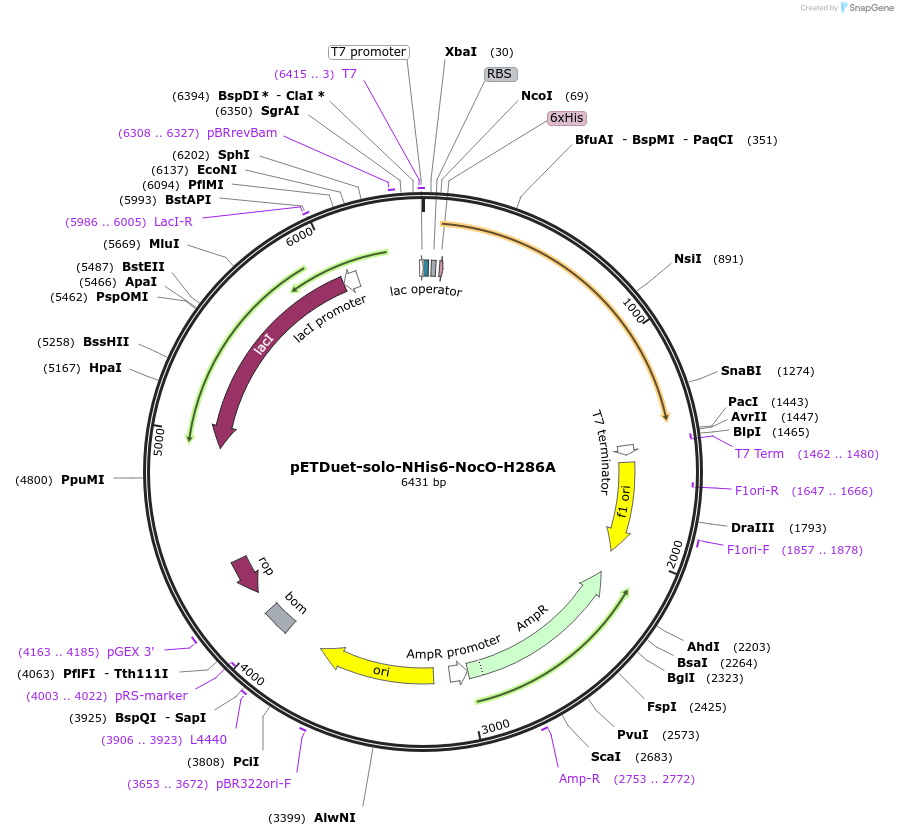
Plasmid#240409Purposeexpresses N-His tagged NocO H286A (diiron halogenase variant) from Nodularia sp. 06071 (mutation numbering based on amino acid position in native sequence), created from mutagenesis of WT plasmidDepositorInsertnocO
TagsHisExpressionBacterialMutationM1 of native sequence not included in insert, His…PromoterT7Available SinceJuly 10, 2025AvailabilityAcademic Institutions and Nonprofits only -
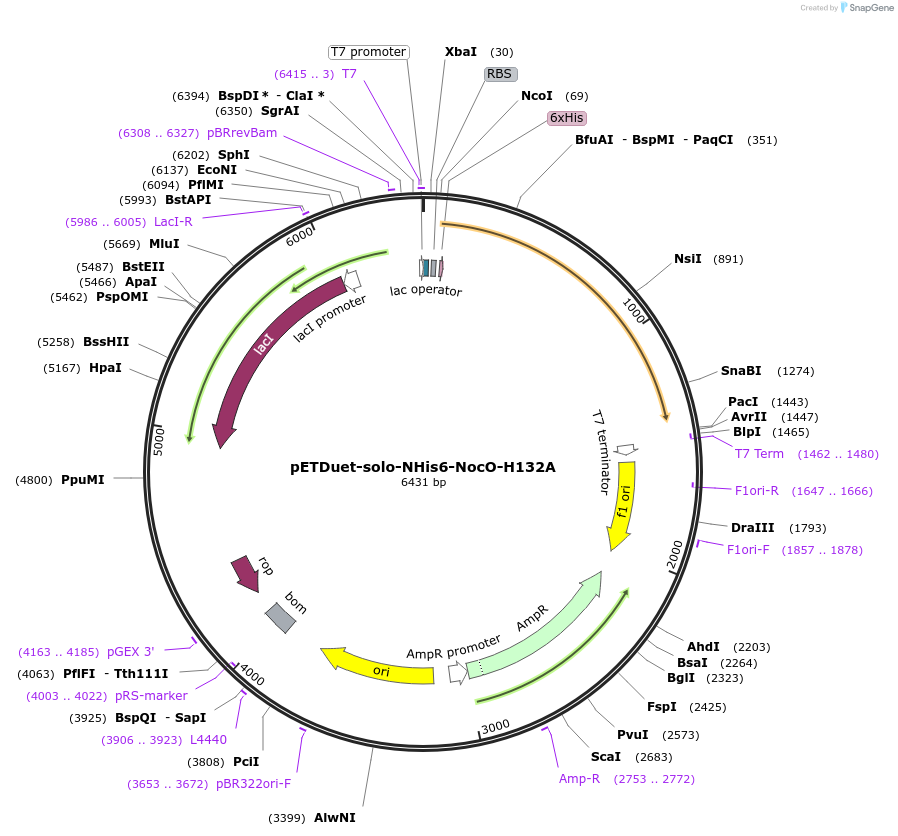
pETDuet-solo-NHis6-NocO-H132A
Plasmid#240404Purposeexpresses N-His tagged NocO H132A (diiron halogenase variant) from Nodularia sp. 06071 (mutation numbering based on amino acid position in native sequence), created from mutagenesis of WT plasmidDepositorInsertnocO
TagsHisExpressionBacterialMutationM1 of native sequence not included in insert, His…PromoterT7Available SinceJuly 10, 2025AvailabilityAcademic Institutions and Nonprofits only -
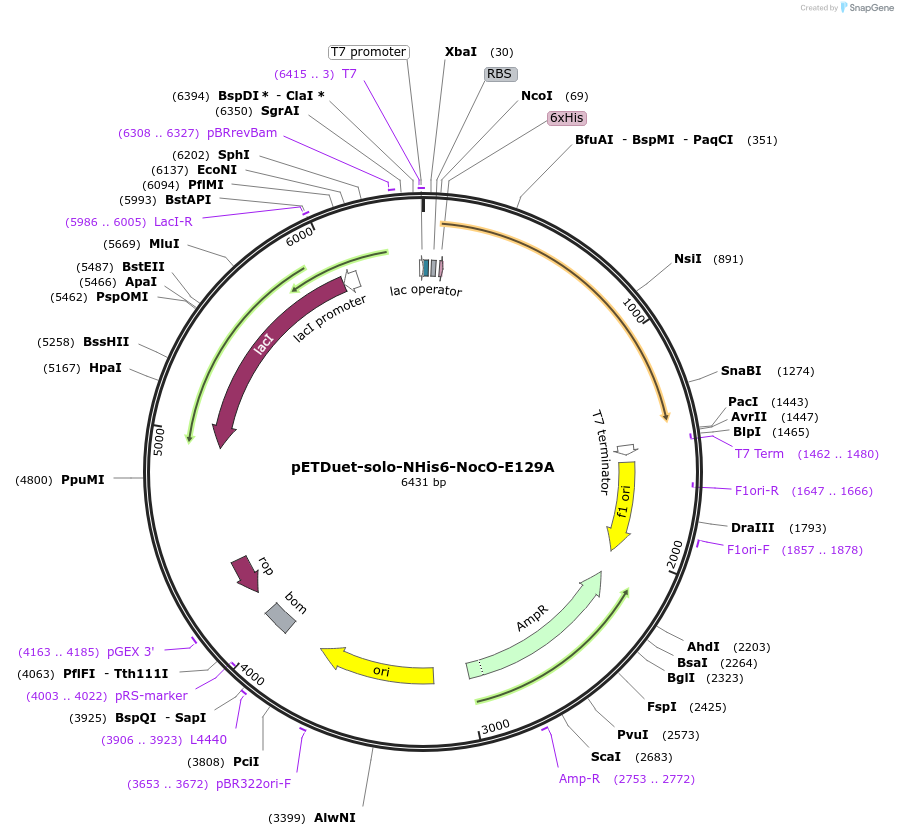
pETDuet-solo-NHis6-NocO-E129A
Plasmid#240403Purposeexpresses N-His tagged NocO E129A (diiron halogenase variant) from Nodularia sp. 06071 (mutation numbering based on amino acid position in native sequence), created from mutagenesis of WT plasmidDepositorInsertnocO
TagsHisExpressionBacterialMutationM1 of native sequence not included in insert, Glu…PromoterT7Available SinceJuly 10, 2025AvailabilityAcademic Institutions and Nonprofits only -
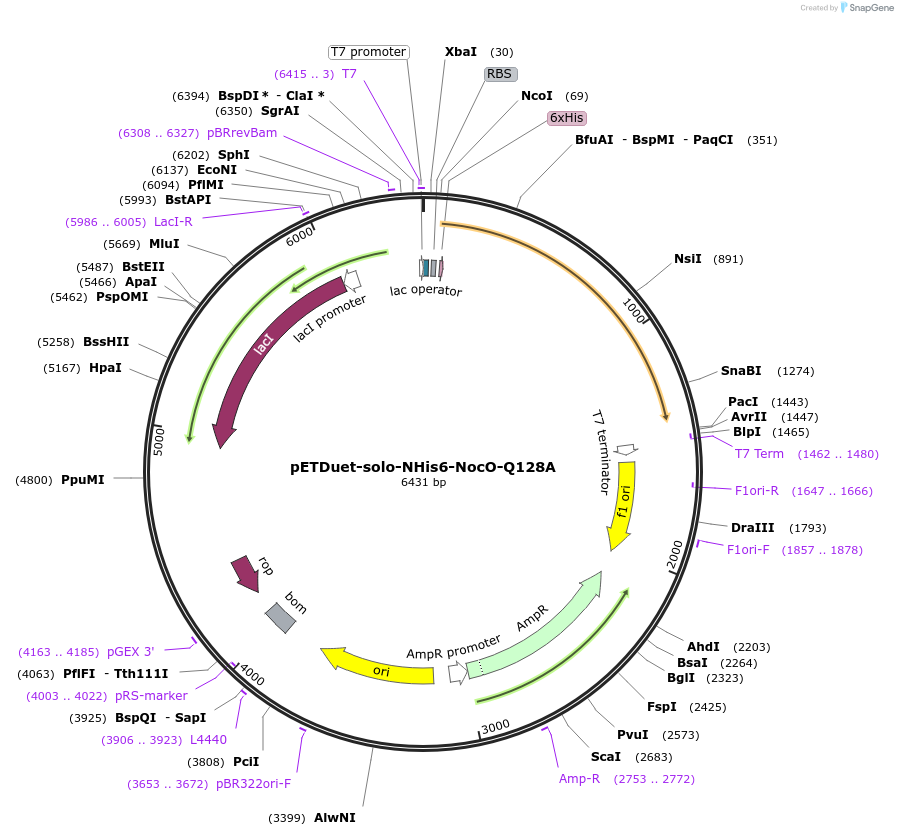
pETDuet-solo-NHis6-NocO-Q128A
Plasmid#240402Purposeexpresses N-His tagged NocO Q128A (diiron halogenase variant) from Nodularia sp. 06071 (mutation numbering based on amino acid position in native sequence), created from mutagenesis of WT plasmidDepositorInsertnocO
TagsHisExpressionBacterialMutationM1 of native sequence not included in insert, Glu…PromoterT7Available SinceJuly 10, 2025AvailabilityAcademic Institutions and Nonprofits only -
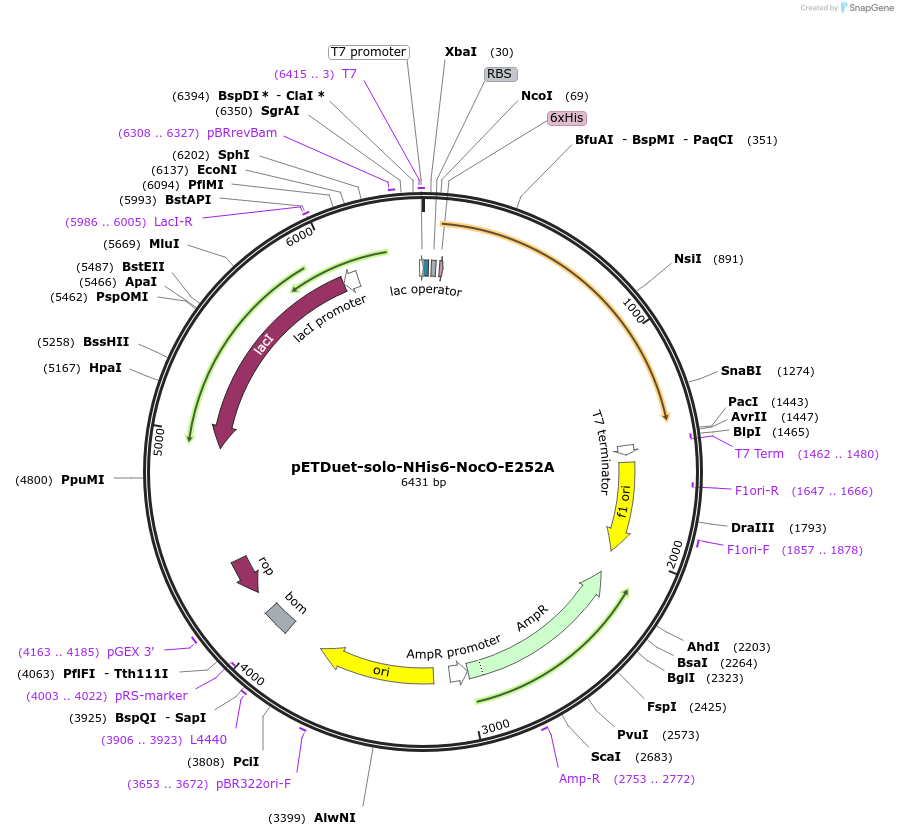
pETDuet-solo-NHis6-NocO-E252A
Plasmid#240405Purposeexpresses N-His tagged NocO E252A (diiron halogenase variant) from Nodularia sp. 06071 (mutation numbering based on amino acid position in native sequence), created from mutagenesis of WT plasmidDepositorInsertnocO
TagsHisExpressionBacterialMutationM1 of native sequence not included in insert, Glu…PromoterT7Available SinceJuly 10, 2025AvailabilityAcademic Institutions and Nonprofits only -
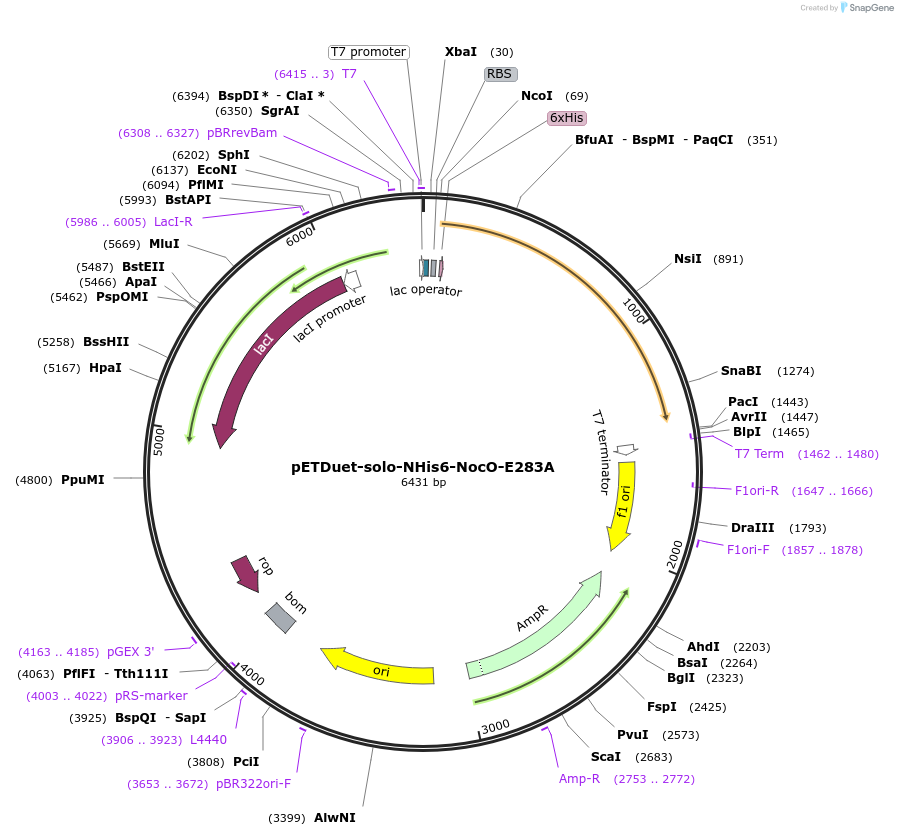
pETDuet-solo-NHis6-NocO-E283A
Plasmid#240408Purposeexpresses N-His tagged NocO E283A (diiron halogenase variant) from Nodularia sp. 06071 (mutation numbering based on amino acid position in native sequence), created from mutagenesis of WT plasmidDepositorInsertnocO
TagsHisExpressionBacterialMutationM1 of native sequence not included in insert, Glu…PromoterT7Available SinceJuly 10, 2025AvailabilityAcademic Institutions and Nonprofits only -
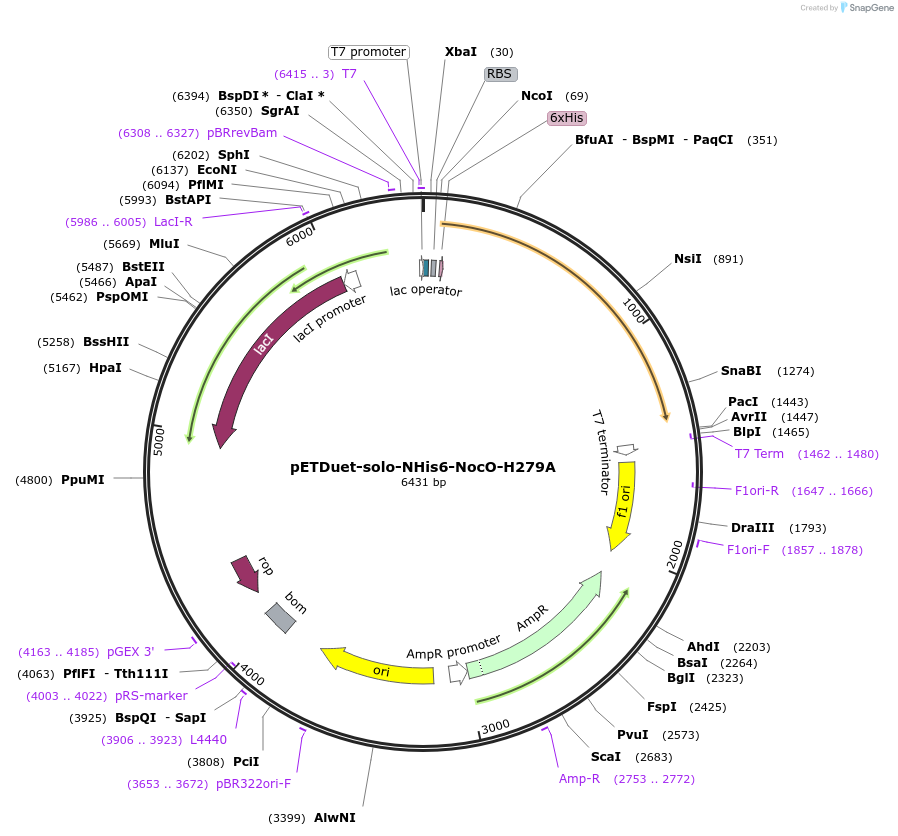
pETDuet-solo-NHis6-NocO-H279A
Plasmid#240406Purposeexpresses N-His tagged NocO H279A (diiron halogenase variant) from Nodularia sp. 06071 (mutation numbering based on amino acid position in native sequence), created from mutagenesis of WT plasmidDepositorInsertnocO
TagsHisExpressionBacterialMutationM1 of native sequence not included in insert, His…PromoterT7Available SinceJuly 10, 2025AvailabilityAcademic Institutions and Nonprofits only -
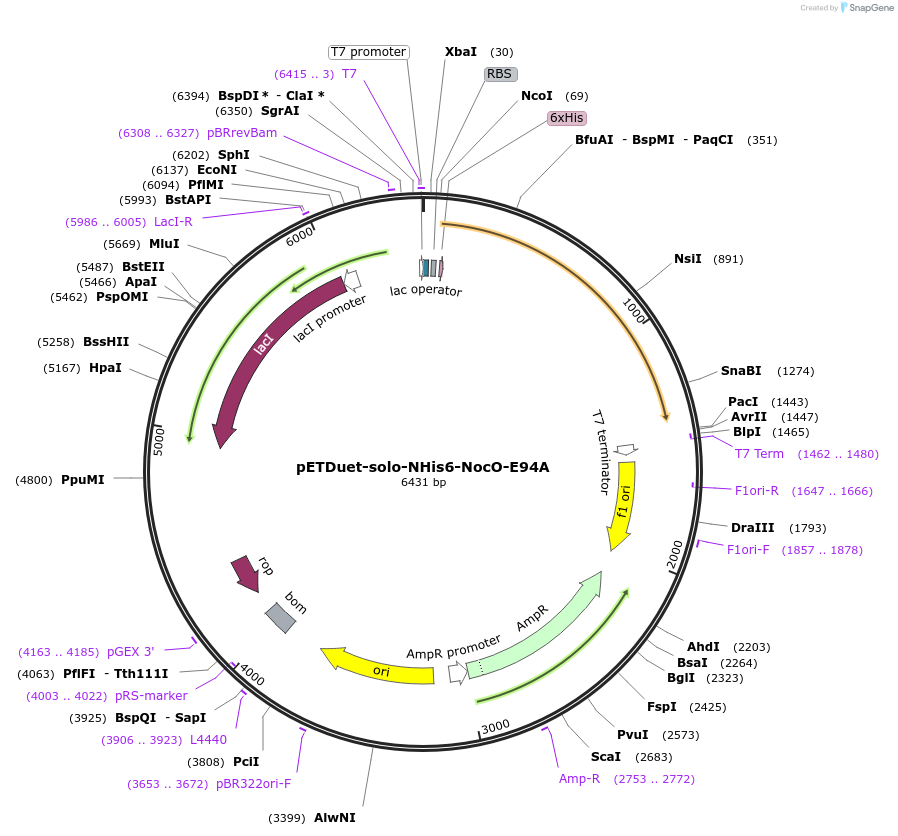
pETDuet-solo-NHis6-NocO-E94A
Plasmid#240401Purposeexpresses N-His tagged NocO E94A (diiron halogenase variant) from Nodularia sp. 06071 (mutation numbering based on amino acid position in native sequence), created from mutagenesis of WT plasmidDepositorInsertnocO
TagsHisExpressionBacterialMutationM1 of native sequence not included in insert, Glu…PromoterT7Available SinceJuly 10, 2025AvailabilityAcademic Institutions and Nonprofits only -
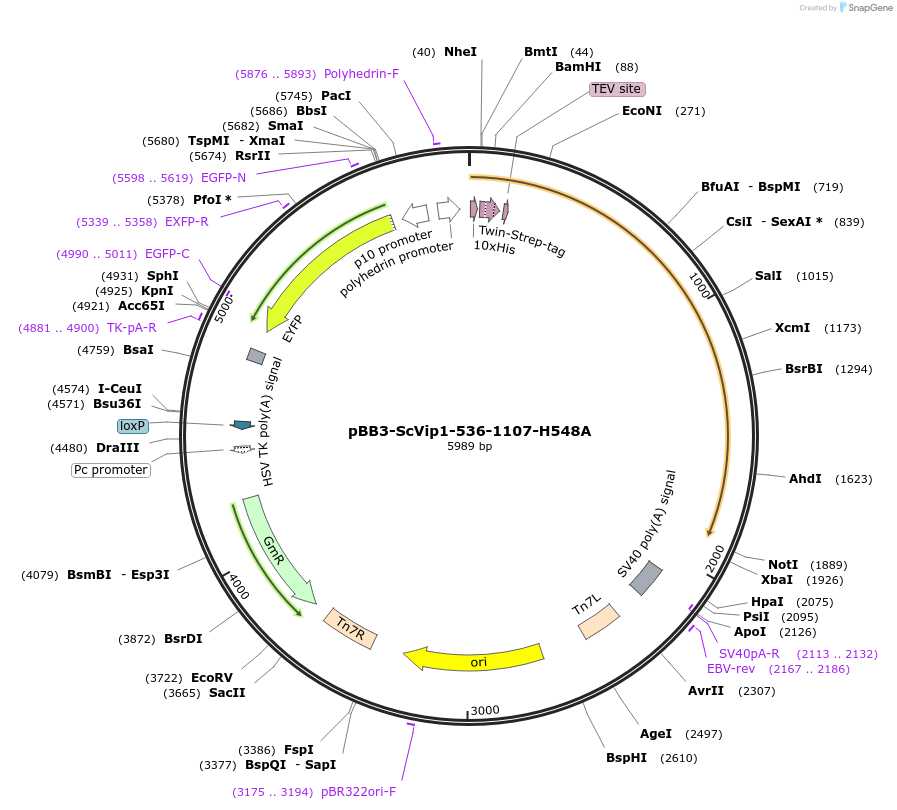
pBB3-ScVip1-536-1107-H548A
Plasmid#238319PurposeScVip1-536-1107-H548A protein expression in insect cellsDepositorInsertVIP1 (VIP1 Budding Yeast)
TagsHis10-Tag, Twin-Strep-tag, TEVExpressionInsectMutationScVip1-536-1107-H548AAvailable SinceJune 26, 2025AvailabilityAcademic Institutions and Nonprofits only -
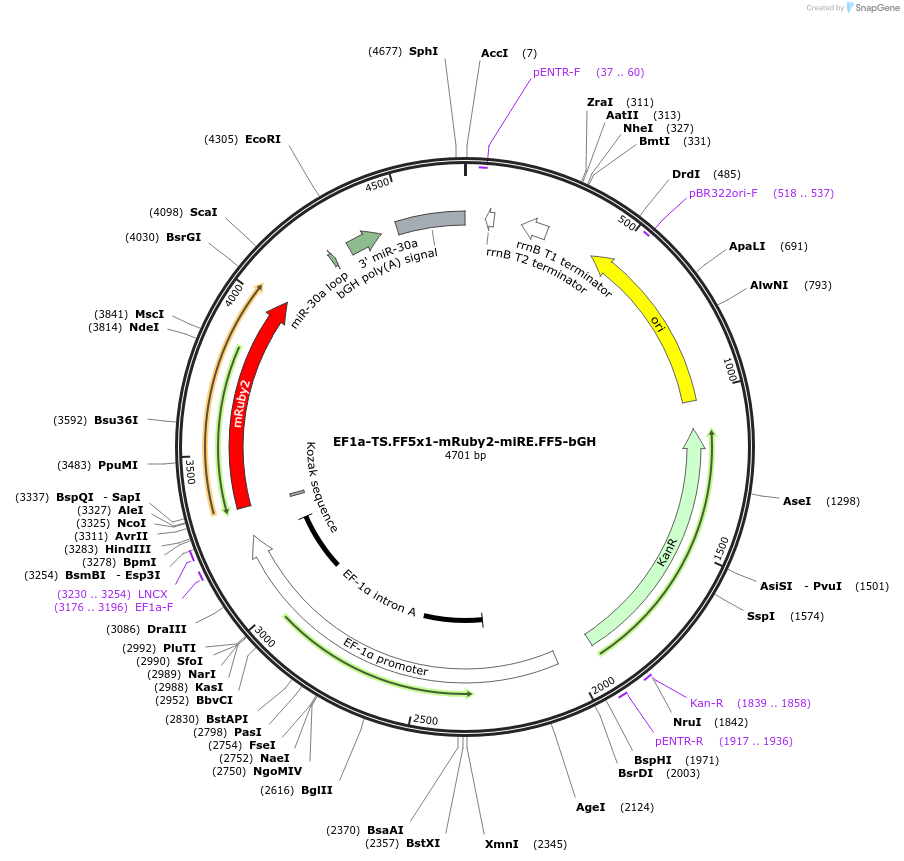
EF1a-TS.FF5x1-mRuby2-miRE.FF5-bGH
Plasmid#235259PurposeComMAND closed-loop circuit regulating mRuby2 with miRE-FF5 design 3DepositorInsertmRuby2
UseSynthetic BiologyExpressionMammalianPromoterEF1a (human EF1a)Available SinceJune 16, 2025AvailabilityAcademic Institutions and Nonprofits only