-
Depositing Lab
-
Publication
-
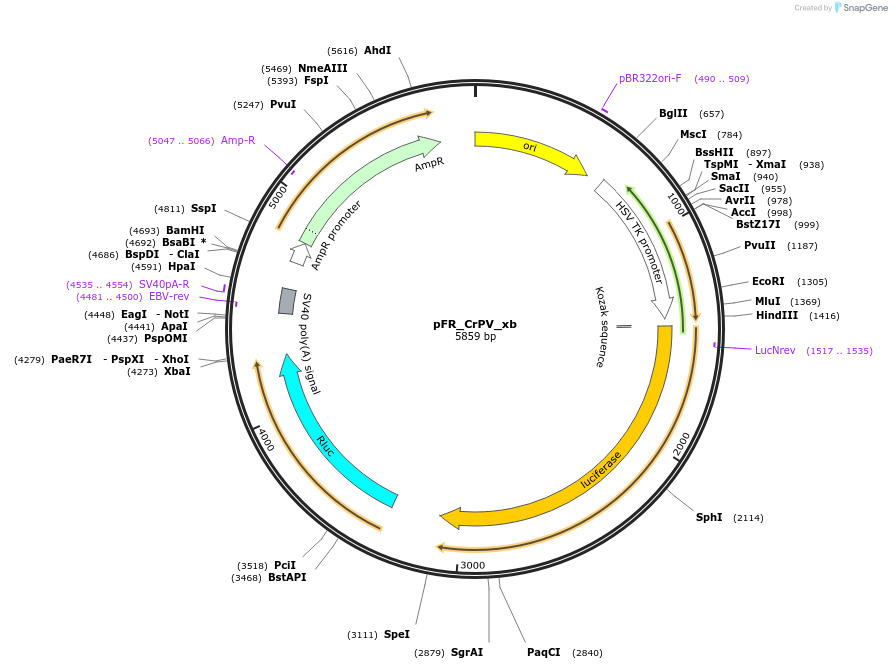
Sequence Information
Ordering
| Item | Catalog # | Description | Quantity | Price (USD) | |
|---|---|---|---|---|---|
| Plasmid | 11509 | Standard format: Plasmid sent in bacteria as agar stab | 1 | $94 | |
Backbone
-
Vector backbonepRL-TK
-
Backbone manufacturerPromega
- Backbone size w/o insert (bp) 4045
-
Vector typeLuciferase
Growth in Bacteria
-
Bacterial Resistance(s)Ampicillin, 100 μg/mL
-
Growth Temperature37°C
-
Growth Strain(s)DH5alpha
-
Copy numberUnknown
Gene/Insert
-
Gene/Insert nameFirefly luciferase, Cricket Paralysis Virus IRES sequence, Renilla, and 3'UTR short RNA binding sites
Cloning Information
- Cloning method Restriction Enzyme
- 5′ cloning site see comments (unknown if destroyed)
- 3′ cloning site see comments (unknown if destroyed)
- 5′ sequencing primer na
- 3′ sequencing primer EBV-rev
- (Common Sequencing Primers)
Resource Information
-
Articles Citing this Plasmid
Terms and Licenses
-
Academic/Nonprofit Terms
-
Industry Terms
- Not Available to Industry
Trademarks:
- Zeocin® is an InvivoGen trademark.
Depositor Comments
The CrPV IRES sequence was cloned by splint ligation of two synthetic 100mers, and cloned into the 5' UTR of pRL-TKx6; the firefly luciferase ORF was then inserted into the resulting construct. Six short RNA binding sites for CXCR4 are also present.
These plasmids were created by your colleagues. Please acknowledge the Principal Investigator, cite the article in which the plasmids were described, and include Addgene in the Materials and Methods of your future publications.
-
For your Materials & Methods section:
pFR_CrPV_xb was a gift from Phil Sharp (Addgene plasmid # 11509 ; http://n2t.net/addgene:11509 ; RRID:Addgene_11509) -
For your References section:
Short RNAs Repress Translation after Initiation in Mammalian Cells. Petersen CP, Bordeleau ME, Pelletier J, Sharp PA. Mol Cell. 2006 Feb 17. 21(4):533-42. 10.1016/j.molcel.2006.01.031 PubMed 16483934