-
Purpose(Empty Backbone) 3'UTR segments of target genes can be inserted into this firefly luciferase reporter to test for their effects on protein production
-
Depositing Lab
-
Publication
-
Sequence Information
Ordering
| Item | Catalog # | Description | Quantity | Price (USD) | |
|---|---|---|---|---|---|
| Plasmid | 12178 | Standard format: Plasmid sent in bacteria as agar stab | 1 | $94 | |
Backbone
-
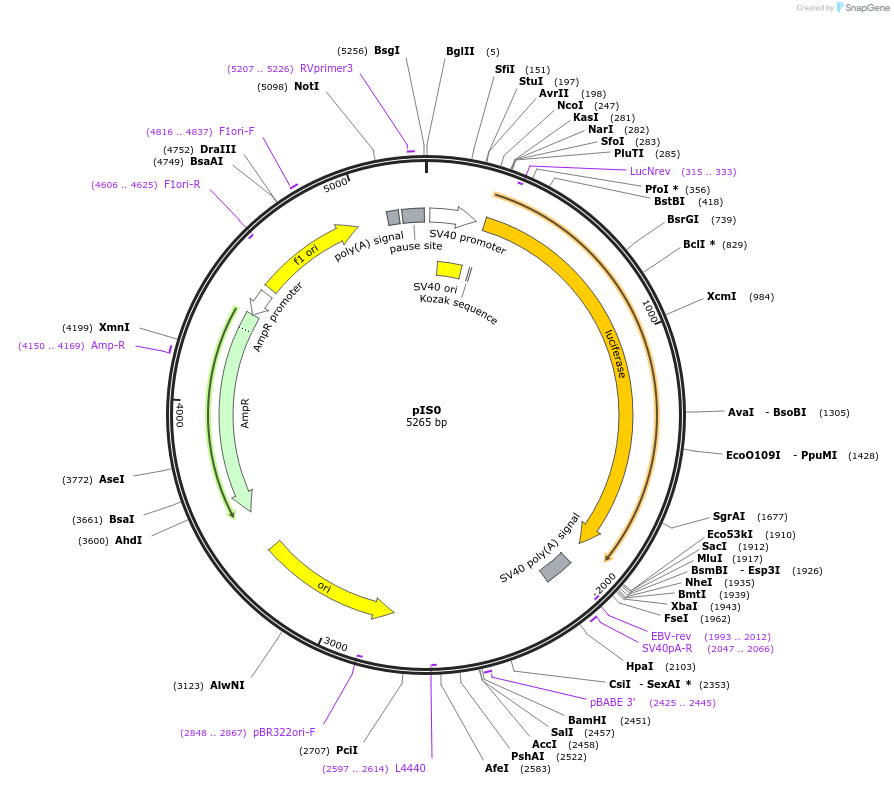
Vector backbonepIS0
-
Backbone manufacturerBartel Lab
- Backbone size (bp) 5264
-
Vector typeMammalian Expression, Luciferase
Growth in Bacteria
-
Bacterial Resistance(s)Ampicillin, 100 μg/mL
-
Growth Temperature37°C
-
Growth Strain(s)DH5alpha
-
Copy numberHigh Copy
Gene/Insert
-
Gene/Insert nameNone
Cloning Information
- Cloning method Restriction Enzyme
- 5′ sequencing primer n/a
- 3′ sequencing primer EBV rev primer (GTGGTTTGTCCAAACTCATC)
- (Common Sequencing Primers)
Resource Information
-
Articles Citing this Plasmid
Terms and Licenses
-
Academic/Nonprofit Terms
-
Industry Terms
- Not Available to Industry
Trademarks:
- Zeocin® is an InvivoGen trademark.
Depositor Comments
The firefly luciferase vector was modified from pGL3 Control Vector (Promega), such that a short sequence containing multiple cloning sites (5'-AGCTCTATACGCGTCTCAAGCTTACTGCTAGC GT-3') was inserted into the XbaI site immediately downstream from the stop codon.
3'UTR segments of target genes can be inserted into this vector to test for their effects on mRNA stability.
These plasmids were created by your colleagues. Please acknowledge the Principal Investigator, cite the article in which the plasmids were described, and include Addgene in the Materials and Methods of your future publications.
-
For your Materials & Methods section:
pIS0 was a gift from David Bartel (Addgene plasmid # 12178 ; http://n2t.net/addgene:12178 ; RRID:Addgene_12178) -
For your References section:
MicroRNA-directed cleavage of HOXB8 mRNA. Yekta S, Shih IH, Bartel DP. Science. 2004 Apr 23. 304(5670):594-6. 10.1126/science.1097434 PubMed 15105502