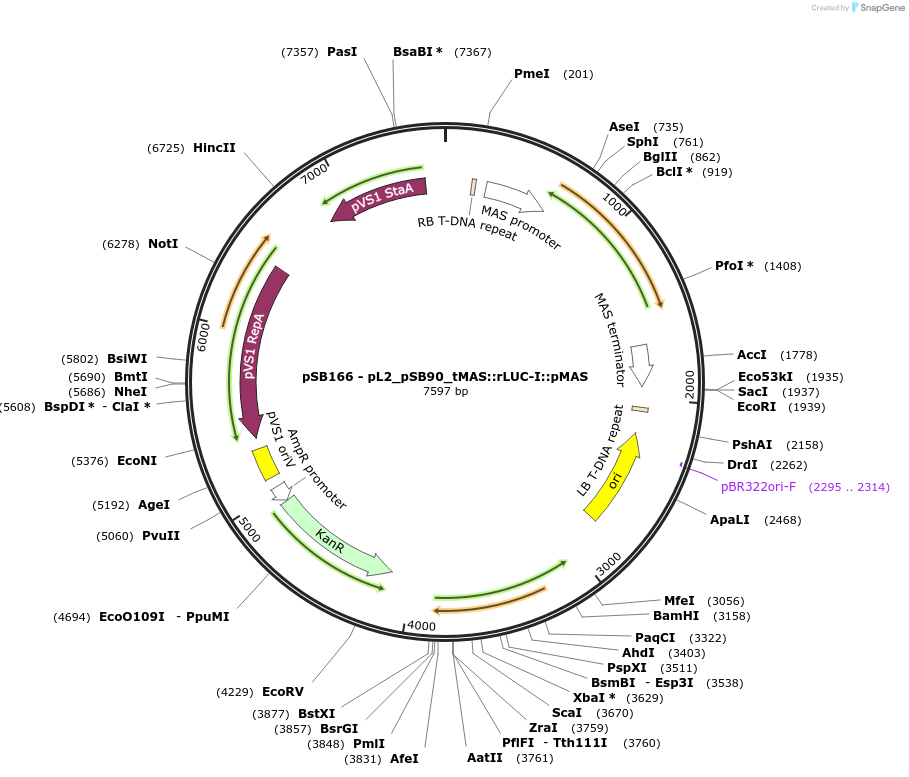
pSB166 - pL2_pSB90_tMAS::rLUC-I::pMAS
(Plasmid
#123199)
-
Purposebinary plant vector for transient expression of a Renilla luciferase (rLUC, with intron)
-
Depositing Labs
-
Sequence Information
Ordering
| Item | Catalog # | Description | Quantity | Price (USD) | |
|---|---|---|---|---|---|
| Plasmid | 123199 | Standard format: Plasmid sent in bacteria as agar stab | 1 | $94 | |
Backbone
-
Vector backbonepSB90
-
Backbone manufacturerModified from pAGM4723 (Weber et al., 2011; DOI:10.1371/journal.pone.0016765) by the addition of the VirGN54D gene
- Backbone size w/o insert (bp) 5851
-
Vector typePlant Expression, Luciferase
Growth in Bacteria
-
Bacterial Resistance(s)Kanamycin, 50 μg/mL
-
Growth Temperature37°C
-
Growth Strain(s)DH5alpha
-
Copy numberHigh Copy
Gene/Insert
-
Gene/Insert nametMAS::rLUC-I::pMAS
-
SpeciesSynthetic
-
Insert Size (bp)1702
Cloning Information
- Cloning method Restriction Enzyme
- 5′ cloning site Unknown (unknown if destroyed)
- 3′ cloning site Unknown (unknown if destroyed)
- 5′ sequencing primer n/a
- (Common Sequencing Primers)
Resource Information
Terms and Licenses
-
Academic/Nonprofit Terms
-
Industry Terms
- Not Available to Industry
Trademarks:
- Zeocin® is an InvivoGen trademark.
Depositor Comments
Vector for transient expression of a Renilla luciferase (rLUC, intron for avoiding expression in Agrobacteria). Small insert size and VirGN54D in the vector backbone facilitate strong transient expression in plants.
These plasmids were created by your colleagues. Please acknowledge the Principal Investigator, cite the article in which the plasmids were described, and include Addgene in the Materials and Methods of your future publications.
-
For your Materials & Methods section:
pSB166 - pL2_pSB90_tMAS::rLUC-I::pMAS was a gift from Erin Cram & Carolyn Lee-Parsons (Addgene plasmid # 123199 ; http://n2t.net/addgene:123199 ; RRID:Addgene_123199) -
For your References section:
EASI Transformation: An Efficient Transient Expression Method for Analyzing Gene Function in Catharanthus roseus Seedlings. Mortensen S, Bernal-Franco D, Cole LF, Sathitloetsakun S, Cram EJ, Lee-Parsons CWT. Front Plant Sci. 2019 Jun 11;10:755. doi: 10.3389/fpls.2019.00755. eCollection 2019. 10.3389/fpls.2019.00755 PubMed 31263474