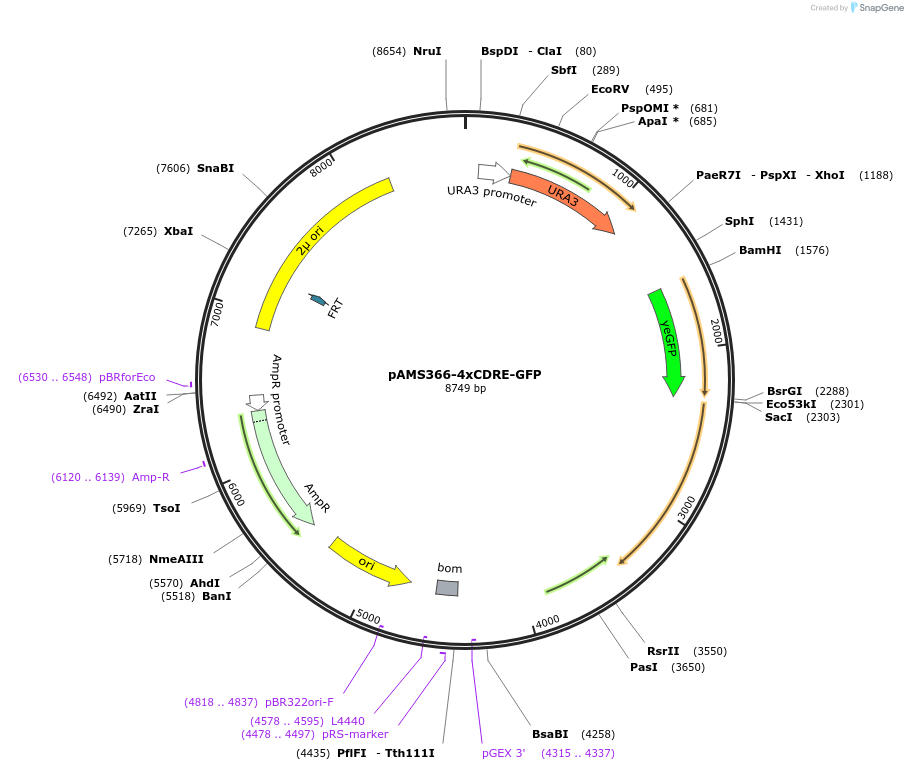
pAMS366-4xCDRE-GFP
(Plasmid
#138657)
-
Purposestable GFP reporter to monitor calcineurin activity via calcineurin-dependent response element (CDRE) in S. cerevisiae
-
Depositing Lab
-
Publication
-
Sequence Information
Ordering
| Item | Catalog # | Description | Quantity | Price (USD) | |
|---|---|---|---|---|---|
| Plasmid | 138657 | Standard format: Plasmid sent in bacteria as agar stab | 1 | $94 | |
Backbone
-
Vector backbonepAMS366
- Total vector size (bp) 8962
-
Vector typeyeast reporter plasmid
-
Selectable markersURA3
Growth in Bacteria
-
Bacterial Resistance(s)Ampicillin, 100 μg/mL
-
Growth Temperature37°C
-
Growth Strain(s)DH5alpha
-
Copy numberUnknown
Gene/Insert
-
Gene/Insert name4xCDRE-CYC1(PM)-yeGFP
-
SpeciesS. cerevisiae (budding yeast); A. victoria
- Promoter CYC1
Cloning Information
- Cloning method Restriction Enzyme
- 5′ cloning site BamHI (not destroyed)
- 3′ cloning site SacI (not destroyed)
- 5′ sequencing primer GTG GGT TTA GAT GAC AAG GGA GAC G
- 3′ sequencing primer CGT ACT GTG AGC CAG AGT TG
- (Common Sequencing Primers)
Resource Information
-
A portion of this plasmid was derived from a plasmid made by1. Stathopoulos AM, and Cyert MS (1997). Calcineurin acts through the CRZ1/TCN1-encoded transcription factor to regulate gene expression in yeast. Genes Dev. 11(24): 3432–3444. 9407035. and 2. Janke C, Magiera MM, Rathfelder N, Taxis C, Reber S, Maekawa H, Moreno-Borchart A, Doenges G, Schwob E, Schiebel E, and Knop M (2004). A versatile toolbox for PCR-based tagging of yeast genes: new fluorescent proteins, more markers and promoter substitution cassettes. Yeast. 21(11): 947–962. doi: 10.1002/yea.1142.
Terms and Licenses
-
Academic/Nonprofit Terms
-
Industry Terms
- Not Available to Industry
Trademarks:
- Zeocin® is an InvivoGen trademark.
Depositor Comments
1x CDRE (calcineurin-dependent response element) = caagcgcacagccaccgactggtg
For construction of pAMS366-4xCDRE-GFP, we used yeGFP from pYM25 (Janke
et al. 2004, sequence available from euroscarf), sequencing showed a
644C>G (Thr>Arg) mutation."
These plasmids were created by your colleagues. Please acknowledge the Principal Investigator, cite the article in which the plasmids were described, and include Addgene in the Materials and Methods of your future publications.
-
For your Materials & Methods section:
pAMS366-4xCDRE-GFP was a gift from Sabrina Büttner (Addgene plasmid # 138657 ; http://n2t.net/addgene:138657 ; RRID:Addgene_138657) -
For your References section:
Stable and destabilized GFP reporters to monitor calcineurin activity in Saccharomyces cerevisiae. Diessl , J., Nandy, A., Schug, C., Habernig, L., and Büttner, S.. Microbial Cell, 2020: in press