-
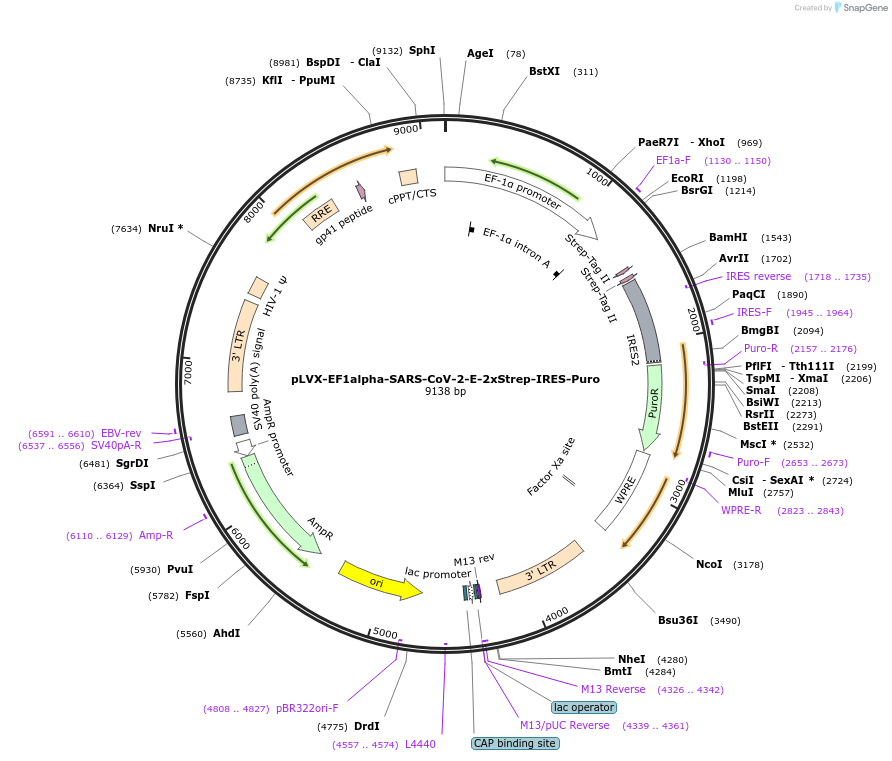
PurposeLentiviral expression of SARS-CoV-2 envelope (E) protein; transient expression and generate lentivirus
-
Depositing Lab
-
Sequence Information
Ordering
| Item | Catalog # | Description | Quantity | Price (USD) | |
|---|---|---|---|---|---|
| Plasmid | 141385 | Standard format: Plasmid sent in bacteria as agar stab | 1 | $94 * | |
* Log in to view industry pricing.
Backbone
-
Vector backbonepLVX-EF1alpha-IRES-Puro
-
Backbone manufacturerTakara Bio
- Backbone size w/o insert (bp) 8825
-
Vector typeMammalian Expression, Lentiviral
-
Selectable markersPuromycin
Growth in Bacteria
-
Bacterial Resistance(s)Ampicillin, 100 μg/mL
-
Growth Temperature30°C
-
Growth Strain(s)NEB Stable
-
Copy numberUnknown
Gene/Insert
-
Gene/Insert nameSARS-CoV-2-E
-
SpeciesSARS-CoV-2
-
Mutationhuman codon optimized
-
Entrez GeneE (a.k.a. GU280_gp04)
- Promoter EF1alpha
-
Tag
/ Fusion Protein
- 2xStrep (C terminal on insert)
Cloning Information
- Cloning method Unknown
Resource Information
-
Articles Citing this Plasmid
Terms and Licenses
-
Academic/Nonprofit Terms
-
Industry Terms
Trademarks:
- Zeocin® is an InvivoGen trademark.
Depositor Comments
Please visit https://www.biorxiv.org/content/10.1101/2020.03.22.002386v3 for bioRxiv preprint.
These plasmids were created by your colleagues. Please acknowledge the Principal Investigator, cite the article in which the plasmids were described, and include Addgene in the Materials and Methods of your future publications.
-
For your Materials & Methods section:
pLVX-EF1alpha-SARS-CoV-2-E-2xStrep-IRES-Puro was a gift from Nevan Krogan (Addgene plasmid # 141385 ; http://n2t.net/addgene:141385 ; RRID:Addgene_141385) -
For your References section:
A SARS-CoV-2 protein interaction map reveals targets for drug repurposing. Gordon DE, Jang GM, Bouhaddou M, Xu J, Obernier K, White KM, O'Meara MJ, Rezelj VV, Guo JZ, Swaney DL, Tummino TA, Huttenhain R, Kaake RM, Richards AL, Tutuncuoglu B, Foussard H, Batra J, Haas K, Modak M, Kim M, Haas P, Polacco BJ, Braberg H, Fabius JM, Eckhardt M, Soucheray M, Bennett MJ, Cakir M, McGregor MJ, Li Q, Meyer B, Roesch F, Vallet T, Mac Kain A, Miorin L, Moreno E, Naing ZZC, Zhou Y, Peng S, Shi Y, Zhang Z, Shen W, Kirby IT, Melnyk JE, Chorba JS, Lou K, Dai SA, Barrio-Hernandez I, Memon D, Hernandez-Armenta C, Lyu J, Mathy CJP, Perica T, Pilla KB, Ganesan SJ, Saltzberg DJ, Rakesh R, Liu X, Rosenthal SB, Calviello L, Venkataramanan S, Liboy-Lugo J, Lin Y, Huang XP, Liu Y, Wankowicz SA, Bohn M, Safari M, Ugur FS, Koh C, Savar NS, Tran QD, Shengjuler D, Fletcher SJ, O'Neal MC, Cai Y, Chang JCJ, Broadhurst DJ, Klippsten S, Sharp PP, Wenzell NA, Kuzuoglu-Ozturk D, Wang HY, Trenker R, Young JM, Cavero DA, Hiatt J, Roth TL, Rathore U, Subramanian A, Noack J, Hubert M, Stroud RM, Frankel AD, Rosenberg OS, Verba KA, Agard DA, Ott M, Emerman M, Jura N, von Zastrow M, Verdin E, Ashworth A, Schwartz O, d'Enfert C, Mukherjee S, Jacobson M, Malik HS, Fujimori DG, Ideker T, Craik CS, Floor SN, Fraser JS, Gross JD, Sali A, Roth BL, Ruggero D, Taunton J, Kortemme T, Beltrao P, Vignuzzi M, Garcia-Sastre A, Shokat KM, Shoichet BK, Krogan NJ. Nature. 2020 Jul;583(7816):459-468. doi: 10.1038/s41586-020-2286-9. Epub 2020 Apr 30. 10.1038/s41586-020-2286-9 PubMed 32353859