-
Depositing Lab
-
Publication
-
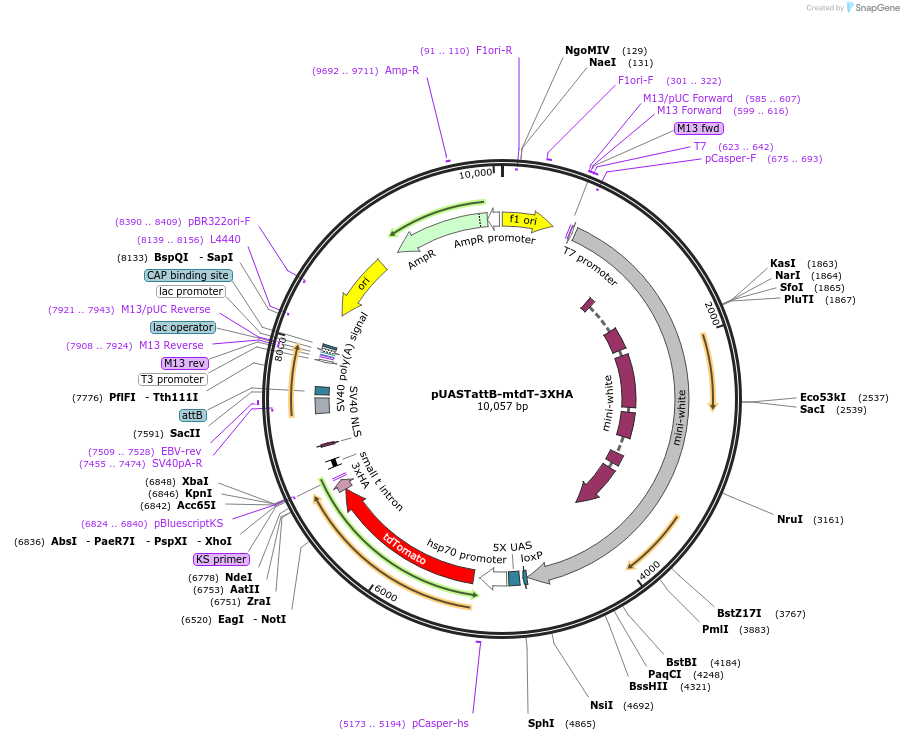
Sequence Information
Ordering
| Item | Catalog # | Description | Quantity | Price (USD) | |
|---|---|---|---|---|---|
| Plasmid | 24355 | Standard format: Plasmid sent in bacteria as agar stab | 1 | $94 | |
Backbone
-
Vector backbonepQuastattB
-
Backbone manufacturerBischof et al., 2007
- Backbone size w/o insert (bp) 8000
-
Vector typeInsect Expression
Growth in Bacteria
-
Bacterial Resistance(s)Ampicillin, 100 μg/mL
-
Growth Temperature37°C
-
Growth Strain(s)DH5alpha
-
Copy numberHigh Copy
Gene/Insert
-
Gene/Insert namemtDT-3XHA
-
Insert Size (bp)2000
-
Tags
/ Fusion Proteins
- mtdT (C terminal on backbone)
- 3XHA (C terminal on backbone)
Cloning Information
- Cloning method Restriction Enzyme
- 5′ cloning site BamHI (unknown if destroyed)
- 3′ cloning site XhoI (unknown if destroyed)
- 5′ sequencing primer T7
- 3′ sequencing primer HA-R
- (Common Sequencing Primers)
Resource Information
-
Reference
-
Articles Citing this Plasmid
Terms and Licenses
-
Academic/Nonprofit Terms
-
Industry Terms
- Not Available to Industry
Trademarks:
- Zeocin® is an InvivoGen trademark.
Depositor Comments
The N-terminal membrane tag on tdTomato contains
8 amino acids that direct myristoylation and palmitoylation (Muzumdar et al., 2007). 3 copies of the
HA tag were PCR amplified from pTHW (Drosophila Genomics Resource Center) and included a 5’
BsrGI restriction site and a 3’ EcoRI restriction site preceded by the TAA stop codon (5’
oligo:TTATGTACAAGTACCCATACGATGTTCCTGACTATGC; 3’ oligo:
TAAGAATTCTTAAGCGTAATCTGGAACGTCATATGGATAGG). pSN20-mtdT (Muzumdar et al.,
2007) was digested with BsrGI and EcoRI and the HA PCR fragment was ligated into this vector to
generate pSN20-mtdT-3xHA. This vector was then digested with XhoI and partially digested with
BamHI and cloned into pUASTattB (Bischof et al., 2007) to generate pUASTattB-mtdT-3xHA. The mtdT-3xHA reporter in vivo is as good as mCD8-GFP in labeling dendritic and axonal processes, or in labeling imaginal disc tissues.
However, it does not label neuronal cell bodies as well as mCD8-GFP as most of the mtdT-3xHA
signal is localized to the plasma membrane surface whereas mCD8-GFP, which also localizes to
intracellular membranes, allows for the cell soma to be better visualized.
These plasmids were created by your colleagues. Please acknowledge the Principal Investigator, cite the article in which the plasmids were described, and include Addgene in the Materials and Methods of your future publications.
-
For your Materials & Methods section:
pUASTattB-mtdT-3XHA was a gift from Liqun Luo (Addgene plasmid # 24355 ; http://n2t.net/addgene:24355 ; RRID:Addgene_24355) -
For your References section:
The Q system: a repressible binary system for transgene expression, lineage tracing, and mosaic analysis. Potter CJ, Tasic B, Russler EV, Liang L, Luo L. Cell. 2010 Apr 30. 141(3):536-48. 10.1016/j.cell.2010.02.025 PubMed 20434990