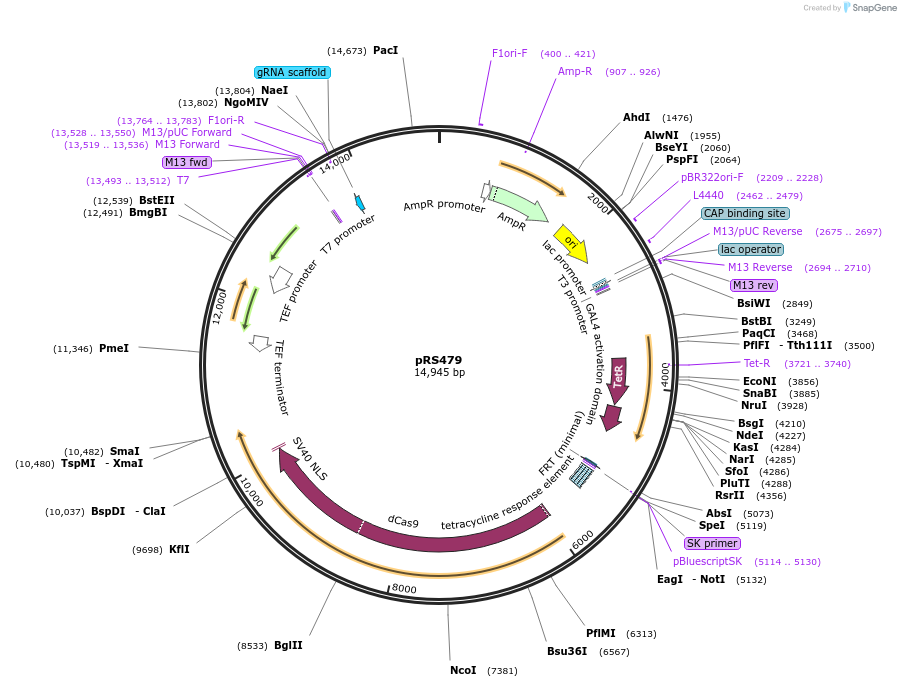
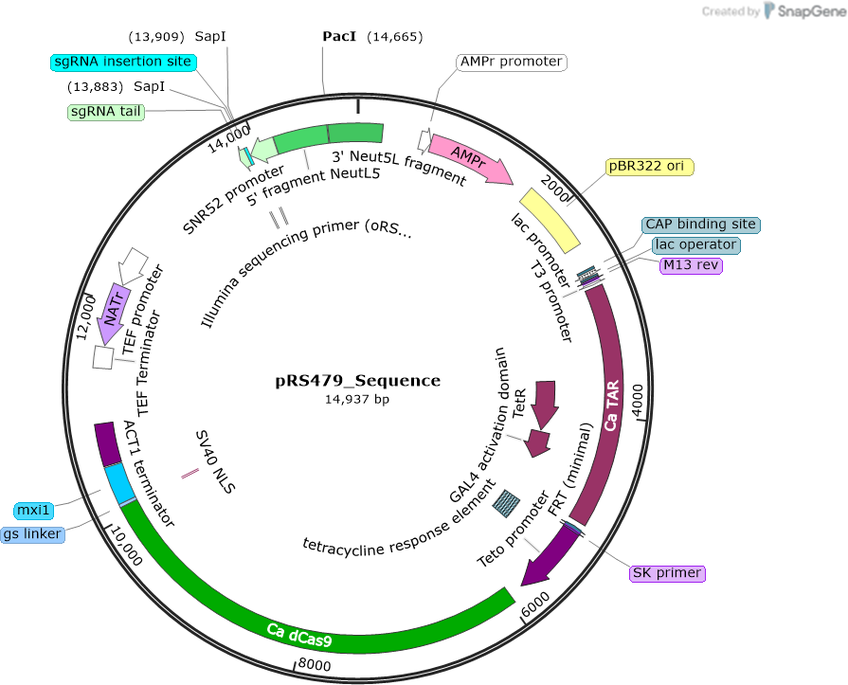
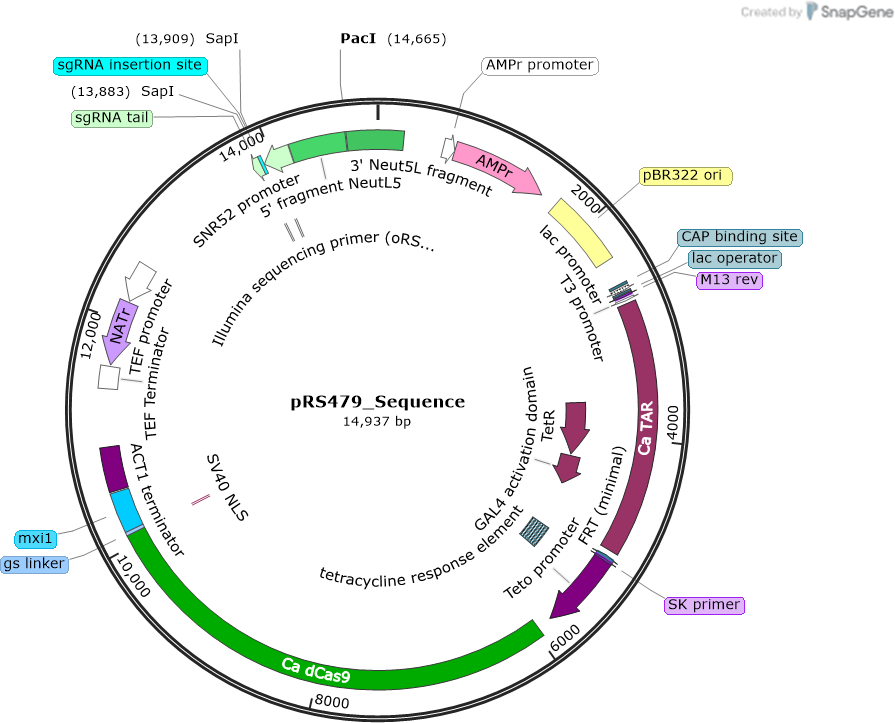
pRS479
(Plasmid
#250493)
-
Purpose(Empty Backbone) Regulatable CRISPRi plasmid for use in Candida albicans. Contains dCas9-Mxi1 controlled by a tetO promoter, NEUT5L integration site and the sgRNA cloning site
-
Depositing Lab
-
Sequence Information
Ordering
| Item | Catalog # | Description | Quantity | Price (USD) | |
|---|---|---|---|---|---|
| Plasmid | 250493 | Standard format: Plasmid sent in bacteria as agar stab | 1 | $94 | |
Backbone
-
Vector backbonepRS159
- Backbone size (bp) 14937
-
Modifications to backboneThe ACT1 promoter controlling dCAS9 expression was swapped out for a C. albicans optimized tetracycline regulated (tetO) promoter. Additionally, the tetracycline transactivator (TAR) was inserted into the plasmid.
-
Vector typeYeast Expression, CRISPR
-
Selectable markersNourseothricin
Growth in Bacteria
-
Bacterial Resistance(s)Ampicillin, 100 μg/mL
-
Growth Temperature30°C
-
Growth Strain(s)DH5alpha
-
Growth instructionsFor transformation into bacteria cells, the transformants must be grown on media containing both Ampicillin (100 μg/mL) and Nourseothricin (50 μg/mL).
-
Copy numberLow Copy
Cloning Information
- Cloning method Golden Gate
Resource Information
Terms and Licenses
-
Academic/Nonprofit Terms
-
Industry Terms
- Not Available to Industry
Trademarks:
- Zeocin® is an InvivoGen trademark.
Depositor Comments
Please visit https://doi.org/10.64898/2025.12.19.695556 for bioRxiv preprint.
These plasmids were created by your colleagues. Please acknowledge the Principal Investigator, cite the article in which the plasmids were described, and include Addgene in the Materials and Methods of your future publications.
-
For your Materials & Methods section:
pRS479 was a gift from Rebecca Shapiro (Addgene plasmid # 250493 ; http://n2t.net/addgene:250493 ; RRID:Addgene_250493) -
For your References section:
Pooled CRISPRi screening reveals fungal-specific drug target candidates. Wensing LF, Despres PC, Francis D, Fogal M, Hendriks A, Gervais NC, Fikry C, Adamu Bukari AR, Gerstein AC, Cuomo CA, Shapiro RS. Nat Microbiol. 2026 May 11. doi: 10.1038/s41564-026-02356-w. 10.1038/s41564-026-02356-w PubMed 42115739