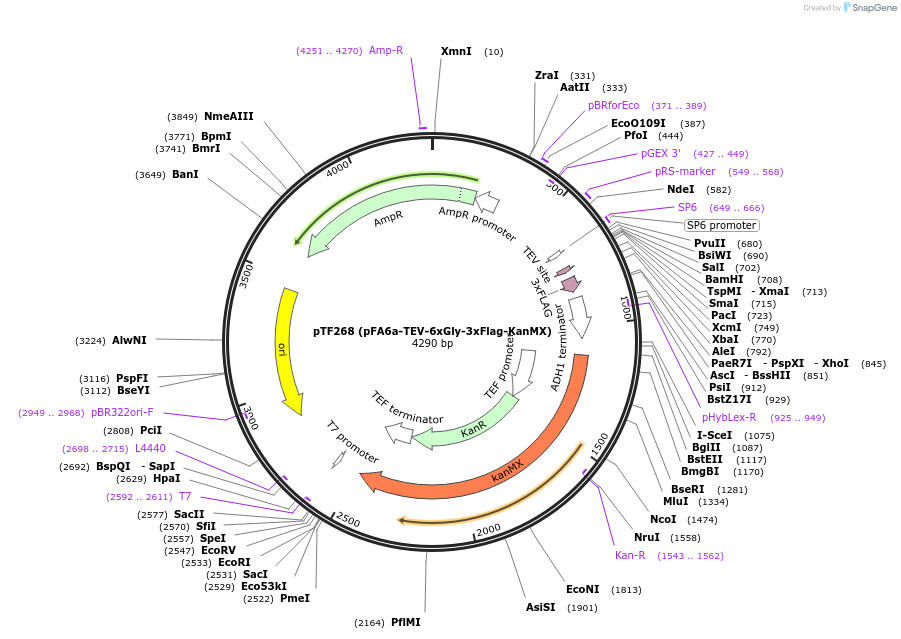
pTF268 (pFA6a-TEV-6xGly-3xFlag-KanMX)
(Plasmid
#44095)
-
Purpose(Empty Backbone)
-
Depositing Lab
-
Publication
-
Sequence Information
Ordering
| Item | Catalog # | Description | Quantity | Price (USD) | |
|---|---|---|---|---|---|
| Plasmid | 44095 | Standard format: Plasmid sent in bacteria as agar stab | 1 | $94 | |
Backbone
-
Vector backbonepFA6a-6xGLY-3xFLAG-kanMX6 (Addgene plasmid # 20754)
-
Backbone manufacturerMark Hochstrasser lab (Yale Univeristy, New Haven, CT) (PMID: 19243080)
- Backbone size (bp) 4255
-
Modifications to backbone6xGly/3xFlag replaced with TEV/6xGly/3xFlag
-
Vector typePCR-based yeast C-terminal tagging
-
Selectable markersG418
-
Tags
/ Fusion Proteins
- TEV protease site (C terminal on backbone)
- 6xGly linker (C terminal on backbone)
- 3xFlag (C terminal on backbone)
- ADH1 transcription terminator (C terminal on backbone)
- KanMX casette (C terminal on backbone)
Growth in Bacteria
-
Bacterial Resistance(s)Ampicillin, 100 μg/mL
-
Growth Temperature37°C
-
Growth Strain(s)DH5alpha
-
Copy numberUnknown
Cloning Information
- Cloning method Ligation Independent Cloning
- 5′ sequencing primer Sp6
- 3′ sequencing primer T7
- (Common Sequencing Primers)
Resource Information
-
A portion of this plasmid was derived from a plasmid made byBackbone pFA6a-6xGLY-3xFLAG-kanMX6 from Mark Hochstrasser lab (Yale Univeristy, New Haven, CT) via Addgene (Plasmid no. 20754)
-
Articles Citing this Plasmid
Terms and Licenses
-
Academic/Nonprofit Terms
-
Industry Terms
- Not Available to Industry
Trademarks:
- Zeocin® is an InvivoGen trademark.
Depositor Comments
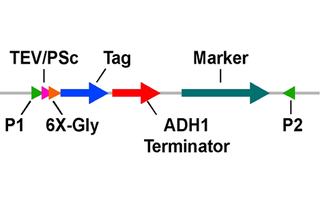
The depositing lab has constructed and deposited here 36 plasmids that can be used to generate linear PCR products to fuse to the 3' end of an ORF in the yeast genome and add (TEV or PreScission protease site) - 6X Gly linker - (12X His, 2X Strep, 3X FLAG, Protein A, or V5) - ADH1 transcription terminator - (KanMX, HphMX, or His3MX).
Six plasmids (Addgene plasmids 44062, 44068, 44074, 44080, 44086, and 44092) have no epitope/purification tag and so can be used to add an alternate tag by standard cloning into unique XhoI-KpnI-XbaI sites, if desired.
The other 30 plasmids are all of the possible combinations of the protease site - tag - marker options listed above. After fusion, the modified protein can then be followed with the epitope or purified using the tag and released from the matrix with the protease site.
The constructs were designed with one of two different protease sites to allow a complex with two different tags to be purified in two successive steps, using each protease to release the complex from each purification matrix.
The schematic diagram (see attached above) represents the format of the constructs and shows the proper scheme for producing the desired PCR products (see also below). If only epitope tagging is needed, the upstream primer can be adjusted to skip the protease site (see example below).
Construction:
DS5421 (pFA6a-6xGLY-3XFLAG-HIS3MX6 (Addgene Plasmid #20753), DS5422 (pFA6a-6xGLY-3XFLAG-KanMX) (Addgene Plasmid #20754), or DS5423 (pFA6a-6xGLY-3XFLAG-HphMX) (Addgene Plasmid #20755) (corresponding to each selection marker) was digested with AscI and PacI to release 3XFlag and replace it with TEV or PreScision sites followed by 6xGly and an XhoI-KpnI-XbaI linker.
Affinity tags were constructed by PCR or by annealing oligos to fit into the XhoI-XbaI gap. This includes 12xHis, 2xStrep II, 3xFLAG, PrtA (2 repeats as found in pBS1479), and V5. Each tag was inserted into each base vector, making 30 templates with all combinations of protease site, tag, and selectable marker.
PCR amplification:
These two primers (along with 50 nt of homology to the targets at the 5' ends) will amplify the desired product from any of the constructs (the reading frame for P1 is important – alternating codons bracketed).
P1 5’-(homology)...[CGA]CGG[ATC]CCC[GGG]TTA[ATT]AAC-3’
P2 5’-(homology)...GATATCATCGATGAATTCGAGCTCGTTT-3’
If only epitope tagging is needed, the upstream primer can be adjusted to skip the protease site. As an example, for the V5-tag-containing constructs, usage of the following primer (5’-(homology)... GGTGGAGGCTCTAGAGGTAAGCCAATTCCA-3’) in place of P1 will omit the protease cleavage site and add only 3xGly and V5 (GGGSR-V5-Stop).
These plasmids were created by your colleagues. Please acknowledge the Principal Investigator, cite the article in which the plasmids were described, and include Addgene in the Materials and Methods of your future publications.
-
For your Materials & Methods section:
pTF268 (pFA6a-TEV-6xGly-3xFlag-KanMX) was a gift from Tim Formosa (Addgene plasmid # 44095 ; http://n2t.net/addgene:44095 ; RRID:Addgene_44095)