-
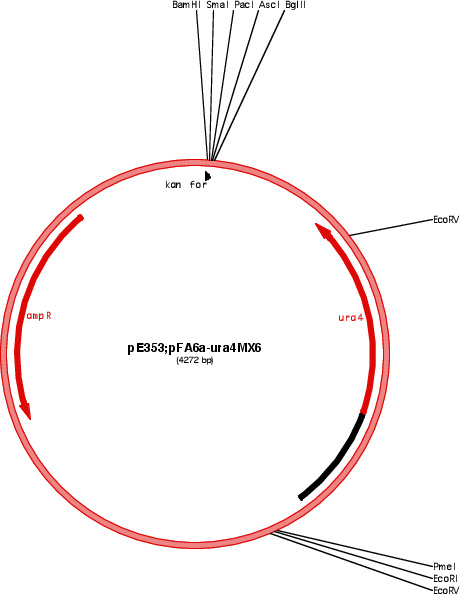
Purpose(Empty Backbone) gene deletion in S. pombe
-
Depositing Lab
-
Sequence Information
Ordering
| Item | Catalog # | Description | Quantity | Price (USD) | |
|---|---|---|---|---|---|
| Plasmid | 49184 | Standard format: Plasmid sent in bacteria as agar stab | 1 | $94 | |
Backbone
-
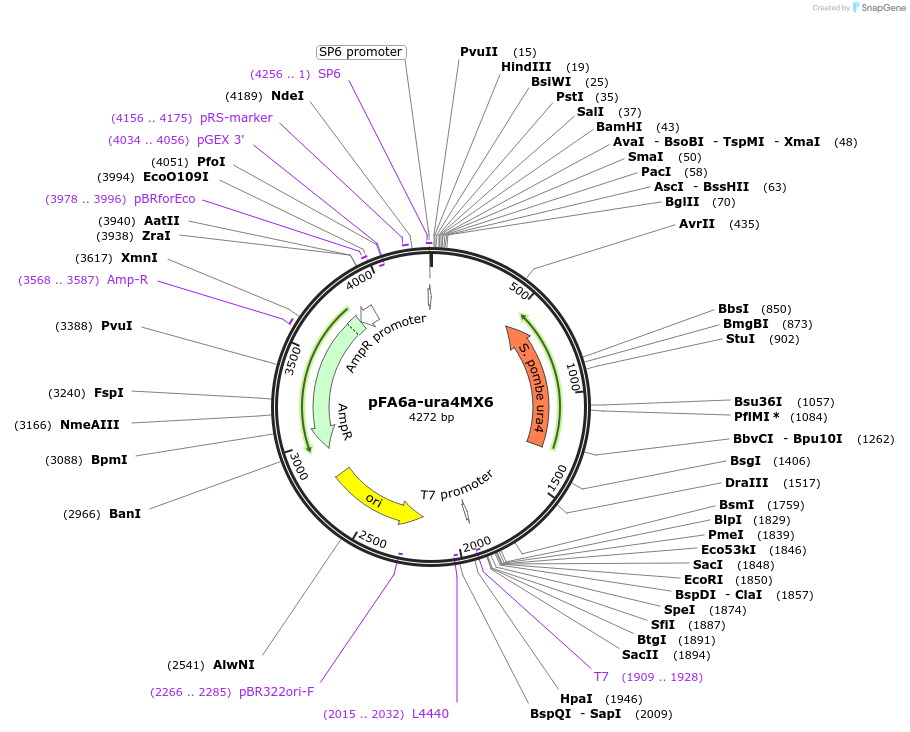
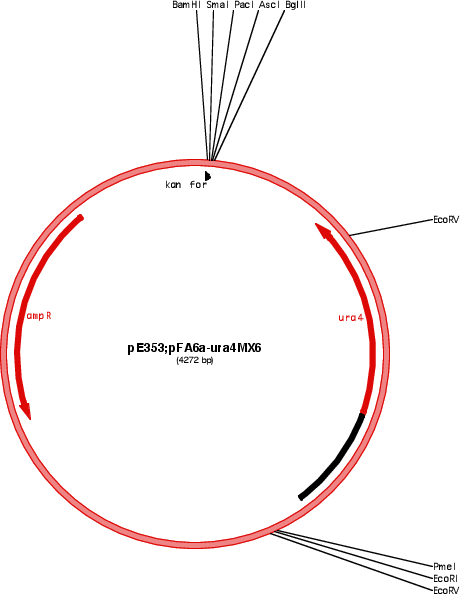
Vector backbonepFA6a
-
Vector typeYeast Expression
-
Selectable markersura4
Growth in Bacteria
-
Bacterial Resistance(s)Ampicillin, 100 μg/mL
-
Growth Temperature37°C
-
Growth Strain(s)DH5alpha
-
Copy numberUnknown
Terms and Licenses
-
Academic/Nonprofit Terms
-
Industry Terms
- Not Available to Industry
Trademarks:
- Zeocin® is an InvivoGen trademark.
Depositor Comments
Note that there are two adenines in Addgene's T7 quality control sequence that introduce a HindIII site and an AflII site into the plasmid. Note that these discrepancies with the depositor sequence do NOT affect the function of the plasmid. The vector is used as a template for PCR to amplify DNA cassettes to modify chromosomal gene loci. The additional adenines are not within the primer sequences to amplify DNA cassettes for gene manipulation.
These plasmids were created by your colleagues. Please acknowledge the Principal Investigator, cite the article in which the plasmids were described, and include Addgene in the Materials and Methods of your future publications.
-
For your Materials & Methods section:
pFA6a-ura4MX6 was a gift from Eishi Noguchi (Addgene plasmid # 49184 ; http://n2t.net/addgene:49184 ; RRID:Addgene_49184) -
For your References section:
New vectors for epitope tagging and gene disruption in Schizosaccharomyces pombe. Gadaleta MC, Iwasaki O, Noguchi C, Noma K, Noguchi E. Biotechniques. 2013 Nov;55(5):257-63. doi: 10.2144/000114100. 10.2144/000114100 PubMed 24215641