-
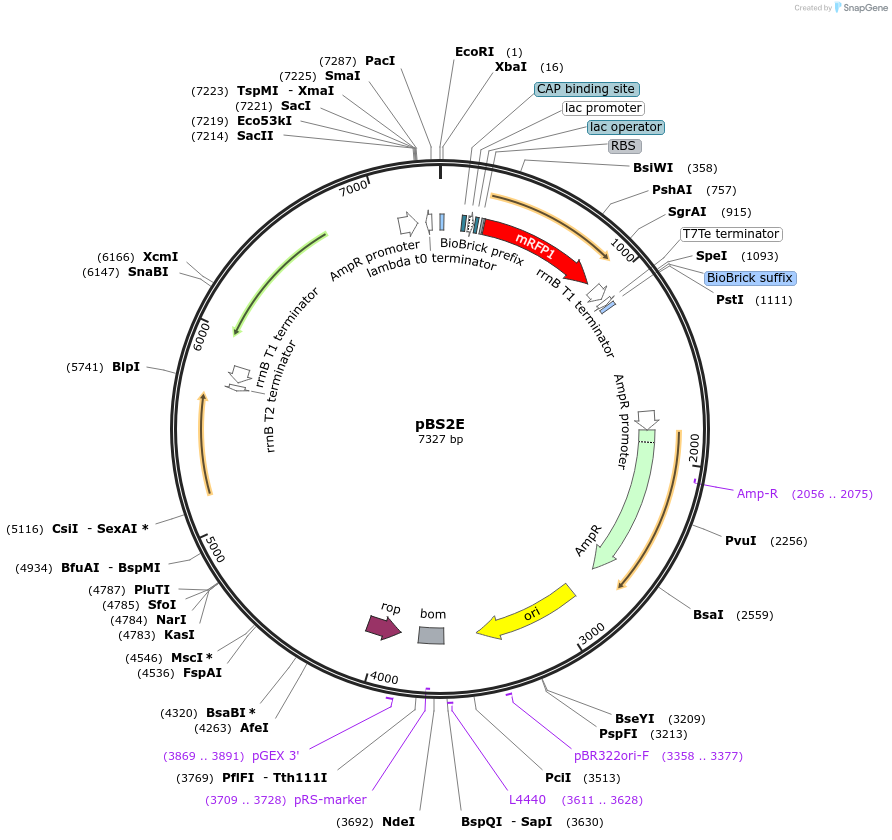
Purpose(Empty Backbone) Empty vector, integration at lacA, ampr, mlsr
-
Depositing Lab
-
Sequence Information
Ordering
| Item | Catalog # | Description | Quantity | Price (USD) | |
|---|---|---|---|---|---|
| Plasmid | 55169 | Standard format: Plasmid sent in bacteria as agar stab | 1 | $94 | |
Backbone
-
Vector backbonepAX01
-
Backbone manufacturerHartl et al 2001
-
Modifications to backbonepAX01 was cut with SacI. The 6.3 kb fragment was religated to remove the xylR-PxylA-fragment. The PstI site in bla was removed. The vector was cut with XbaI and the 6 kb fragment religated to reduce the amount of forbidden restriction sites. The lost terminator of erm was replaced by the PCR-amplified terminator with BsaI-overhangs. The vector was opened with XbaI and ligated with the PCR product cut with BsaI (XbaI-compatible overhang). The correct direction was checked by sequencing and removal of the XbaI-site was confirmed. Finally, the MCS was amplified from pSB1C3 with PstI and NsiI overhangs, cut with PstI and NsiI and ligated into the PstI-cut vector. Correct orientation of the insert was confirmed by restriction digest and sequencing. The remaining NgoMIV sites were removed by subsequent site-directed mutagenesis.
-
Vector typeSynthetic Biology ; Bacillus BioBrick Box
- Promoter none
-
Selectable markerserm: specifies resistance to macrolide, lincosamide and streptogramin B antibiotics if induced by erythromycin
Growth in Bacteria
-
Bacterial Resistance(s)Ampicillin, 100 μg/mL
-
Growth Temperature37°C
-
Growth Strain(s)DH5alpha
-
Copy numberUnknown
Cloning Information
- Cloning method Restriction Enzyme
- 5′ sequencing primer na
- (Common Sequencing Primers)
Resource Information
-
Article Citing this Plasmid
Terms and Licenses
-
Academic/Nonprofit Terms
-
Industry Terms
- Not Available to Industry
Trademarks:
- Zeocin® is an InvivoGen trademark.
Depositor Comments
This is an “empty” vector that lacks promoters and reporter genes. The integrative part contains the flanking homology regions, a resistance cassette for selection in B. subtilis and the multiple cloning site (MCS), containing an rfp-cassette flanked by the restriction sites EcoRI, NotI, XbaI (upstream) and SpeI, NotI and PstI (downstream). They allow cloning in BioBrick standard with selection for white colonies as a result of the removal of the rfp-insert, which – if still present – leads to formation of red colonies in E. coli.
For sequencing of inserts, use the following primers:
fwd: GGCAACCGAGCGTTCTG
rev: CTGACAGCGTTTCGATCC
These plasmids were created by your colleagues. Please acknowledge the Principal Investigator, cite the article in which the plasmids were described, and include Addgene in the Materials and Methods of your future publications.
-
For your Materials & Methods section:
pBS2E was a gift from Thorsten Mascher (Addgene plasmid # 55169 ; http://n2t.net/addgene:55169 ; RRID:Addgene_55169) -
For your References section:
The Bacillus BioBrick Box: generation and evaluation of essential genetic building blocks for standardized work with Bacillus subtilis. Radeck J, Kraft K, Bartels J, Cikovic T, Durr F, Emenegger J, Kelterborn S, Sauer C, Fritz G, Gebhard S, Mascher T. J Biol Eng. 2013 Dec 2;7(1):29. doi: 10.1186/1754-1611-7-29. 10.1186/1754-1611-7-29 PubMed 24295448