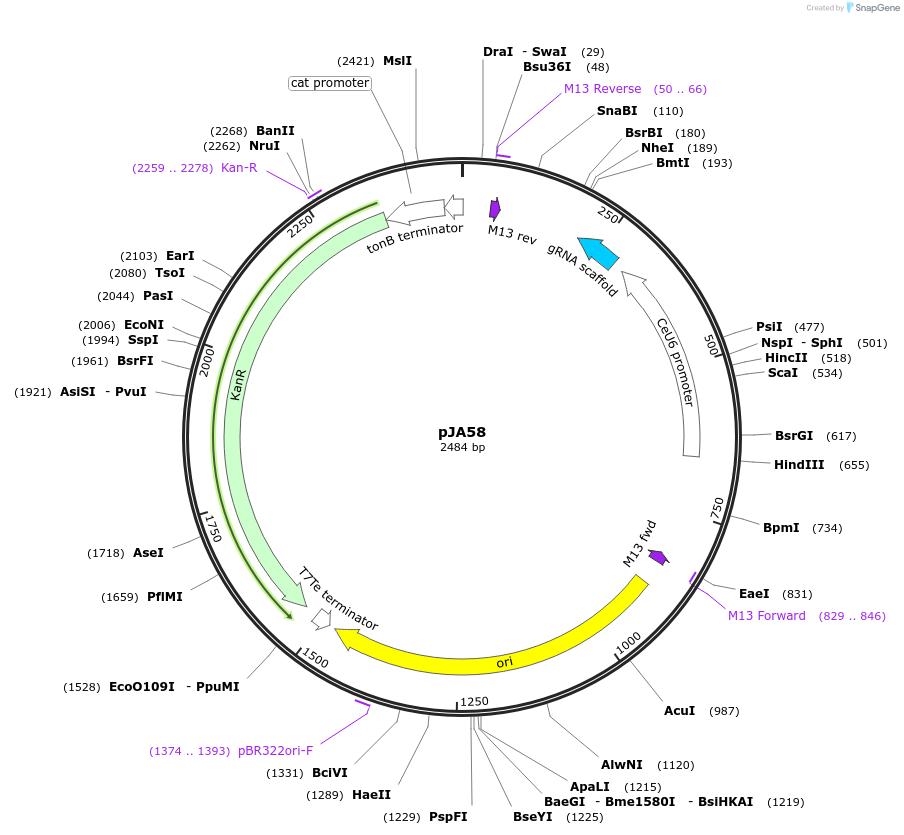
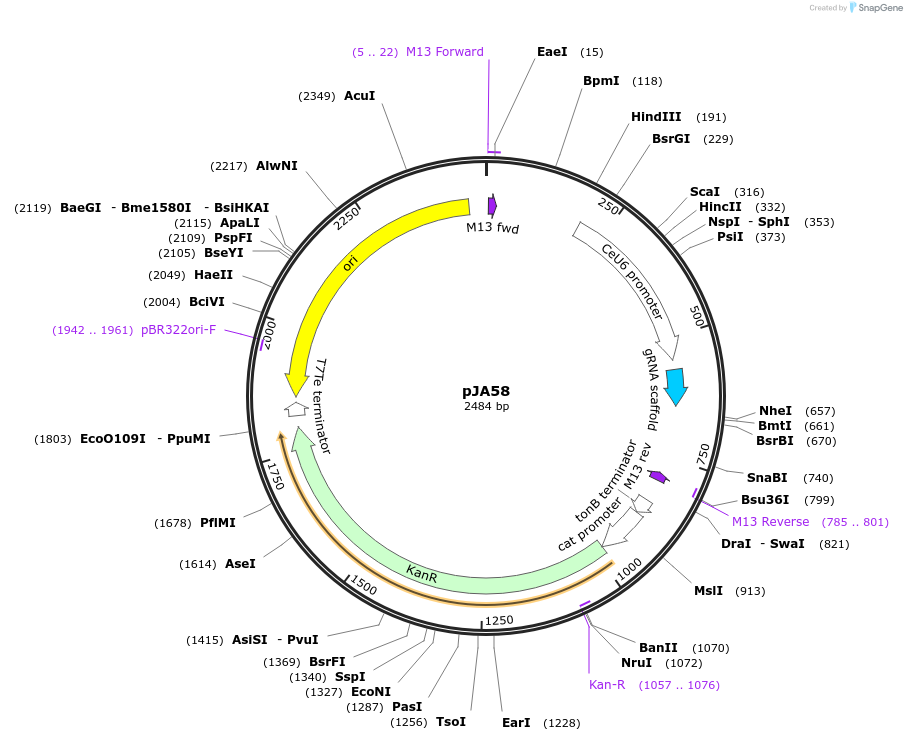
-
PurposegRNA for cleavage at dpy-10(cn64) locus in C elegans
-
Depositing Lab
-
Sequence Information
Ordering
| Item | Catalog # | Description | Quantity | Price (USD) | |
|---|---|---|---|---|---|
| Plasmid | 59933 | Standard format: Plasmid sent in bacteria as agar stab | 1 | $94 | |
Backbone
-
Vector backbonepRB1017
-
Vector typeCRISPR
Growth in Bacteria
-
Bacterial Resistance(s)Kanamycin, 50 μg/mL
-
Growth Temperature37°C
-
Growth Strain(s)Top10
-
Growth instructionsFrom depositor: These gRNA vectors tende to concatamerize. They may appear normal by digest or sequencing, but by running undigested vector out on a gel can you determine whether concatamerization has occurred. Note that the catamers may still function normally.
-
Copy numberHigh Copy
Gene/Insert
-
Gene/Insert namedpy-10(cn64) gRNA
-
SpeciesC. elegans (nematode)
-
Insert Size (bp)19
- Promoter U6
Cloning Information
- Cloning method Restriction Enzyme
- 5′ cloning site BsaI (destroyed during cloning)
- 3′ cloning site BsaI (destroyed during cloning)
- 5′ sequencing primer M13 For
- 3′ sequencing primer M13 Rev
- (Common Sequencing Primers)
Resource Information
-
Articles Citing this Plasmid
Terms and Licenses
-
Academic/Nonprofit Terms
-
Industry Terms
- Not Available to Industry
Trademarks:
- Zeocin® is an InvivoGen trademark.
These plasmids were created by your colleagues. Please acknowledge the Principal Investigator, cite the article in which the plasmids were described, and include Addgene in the Materials and Methods of your future publications.
-
For your Materials & Methods section:
pJA58 was a gift from Andrew Fire (Addgene plasmid # 59933 ; http://n2t.net/addgene:59933 ; RRID:Addgene_59933) -
For your References section:
Efficient Marker-Free Recovery of Custom Genetic Modifications with CRISPR/Cas9 in Caenorhabditis elegans. Arribere JA, Bell RT, Fu BX, Artiles KL, Hartman PS, Fire AZ. Genetics. 2014 Aug 26. pii: genetics.114.169730. 10.1534/genetics.114.169730 PubMed 25161212