-
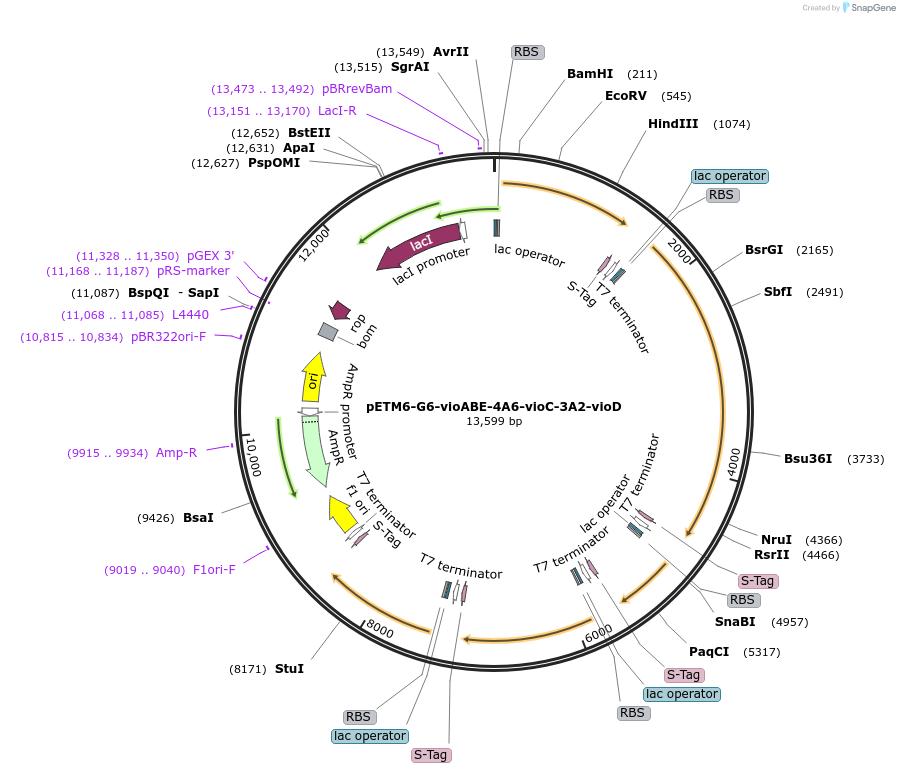
PurposeViolacein pathway (vioABECD) in monocistronic configuration, transcriptionally driven by orthogonal T7-lac promoter variants G6 (vioABE), 4A6 (vioC), and 3A2 (vioD) for orthogonal flux redirection.
-
Depositing Lab
-
Publication
-
Sequence Information
Ordering
| Item | Catalog # | Description | Quantity | Price (USD) | |
|---|---|---|---|---|---|
| Plasmid | 73440 | Standard format: Plasmid sent in bacteria as agar stab | 1 | $94 | |
Backbone
-
Vector backbonepETM6
-
Backbone manufacturerKoffas Lab, Addgene Plasmid #49795
- Backbone size w/o insert (bp) 5147
-
Vector typeBacterial Expression, CRISPR, Synthetic Biology
Growth in Bacteria
-
Bacterial Resistance(s)Ampicillin, 100 μg/mL
-
Growth Temperature37°C
-
Growth Strain(s)DH5alpha
-
Copy numberHigh Copy
Gene/Insert
-
Gene/Insert namePG6-vioABE-P4A6-vioC-P3A2-vioD
-
SpeciesPseudoalteromonas luteoviolacea strain B
- Promoter PG6, P4A6, and P3A2 (orthogonal T7-lac variants)
Cloning Information
- Cloning method Unknown
- 5′ sequencing primer pBRrevBam
- (Common Sequencing Primers)
Resource Information
-
Articles Citing this Plasmid
Terms and Licenses
-
Academic/Nonprofit Terms
-
Industry Terms
- Not Available to Industry
Trademarks:
- Zeocin® is an InvivoGen trademark.
Depositor Comments
Coexpression with G6 CRISPR repressor shuts off all colored metabolite production. Coexpression with 4A6 CRISPR repressor switches carbon flux from violacein production (violet culture) to proviolacein (blue-violet culture). Coexpression with with 3A2 CRISPR repressor switches carbon flux from violacein production (violet culture) to deoxyviolacein (red-violet).
These plasmids were created by your colleagues. Please acknowledge the Principal Investigator, cite the article in which the plasmids were described, and include Addgene in the Materials and Methods of your future publications.
-
For your Materials & Methods section:
pETM6-G6-vioABE-4A6-vioC-3A2-vioD was a gift from Mattheos Koffas (Addgene plasmid # 73440 ; http://n2t.net/addgene:73440 ; RRID:Addgene_73440) -
For your References section:
Rapid generation of CRISPR/dCas9-regulated, orthogonally repressible hybrid T7-lac promoters for modular, tuneable control of metabolic pathway fluxes in Escherichia coli. Cress BF, Jones JA, Kim DC, Leitz QD, Englaender JA, Collins SM, Linhardt RJ, Koffas MA. Nucleic Acids Res. 2016 Apr 13. pii: gkw231. 10.1093/nar/gkw231 PubMed 27079979