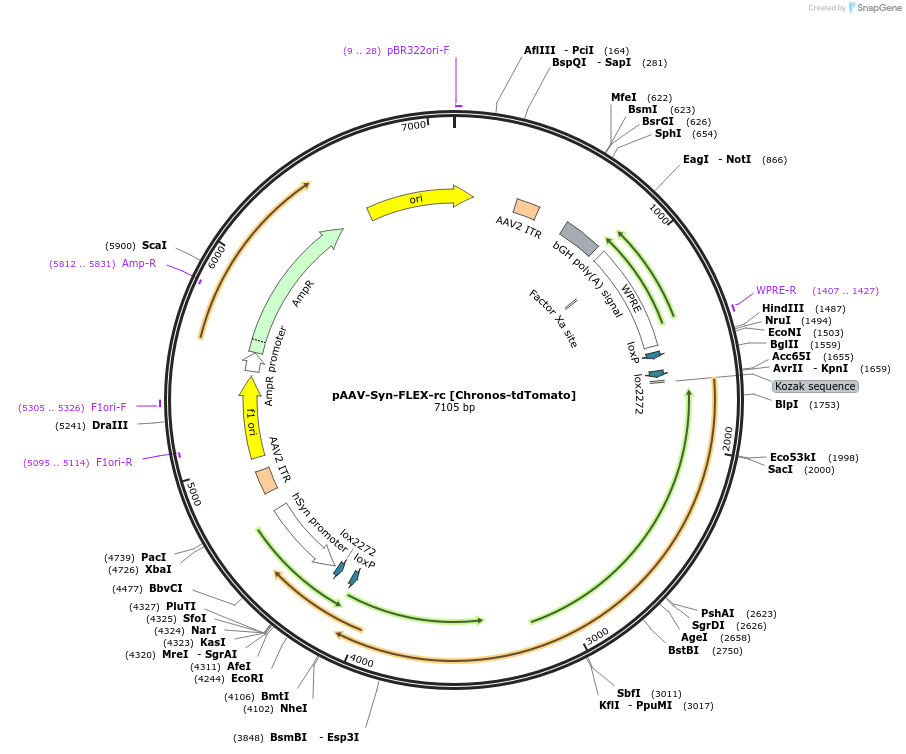
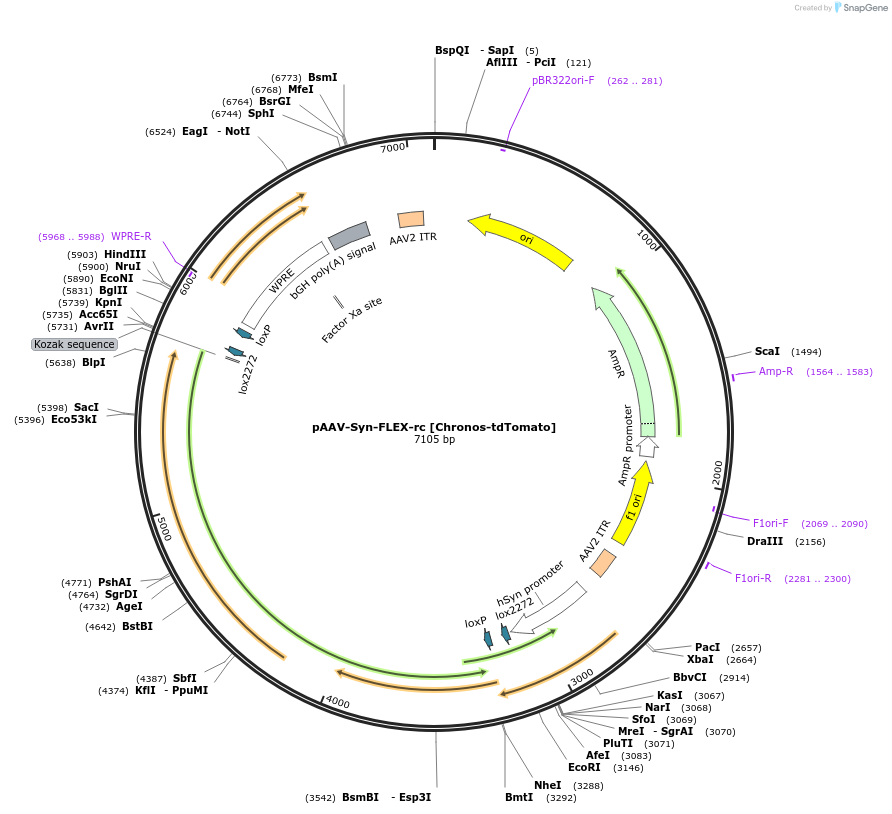
pAAV-Syn-FLEX-rc [Chronos-tdTomato]
(Plasmid
#84481)
-
PurposeAAV-mediated expression of Chronos-tdTomato under the Syn promoter, in floxed/reversed (Cre-dependent) manner. Using bGHpA signal. tdTomato is a codon diversified version.
-
Depositing Lab
-
Sequence Information
Ordering
| Item | Catalog # | Description | Quantity | Price (USD) | |
|---|---|---|---|---|---|
| Plasmid | 84481 | Standard format: Plasmid sent in bacteria as agar stab | 1 | $94 | |
Want a viral vector made from this plasmid?
Make a packaging request and we'll get back to you.
Please log in to submit a packaging request.
-
SerotypeSelect serotype for details See details about
-
PricingSelect serotype and quantity $ USD for preparation of µL virus + $34 USD for plasmid.
-
How this works
- Place a request for a quantity of 2 (0.2 mL), 10 (1 mL), 25 (2.5 mL), or 50 (5 mL). Our all-inclusive pricing includes DNA production and QC.
- Addgene will quickly confirm that we can produce a high-quality prep for you.
- Track your request and place an order from within your account. Payment information must be added before we can begin processing your order.
- Receive your prep in 6–9 weeks after the MTA is approved by your organization.
- Learn more about our Packaged on Request Service.
Backbone
-
Vector backboneAAV-Syn-FLEX
-
Backbone manufactureroriginal from Stratagene
- Backbone size w/o insert (bp) 4684
- Total vector size (bp) 7105
-
Vector typeMammalian Expression, AAV
Growth in Bacteria
-
Bacterial Resistance(s)Ampicillin, 100 μg/mL
-
Growth Temperature37°C
-
Growth Strain(s)NEB Stable
-
Growth instructionsCan use DH5alpha at 37°C or Stbl3 at 30°C. Carbenicillin is preferred over ampicillin. In DH5alpha this plasmid may act more like a high copy plasmid, although in Stbl3 it may act more like a low copy plasmid.
-
Copy numberLow Copy
Gene/Insert
-
Gene/Insert nameChronos-tdTomato
-
Alt nameStigeoclonium helveticum channelrhodopsin-GFP
-
Alt nameChR90-tdTomato
-
SpeciesStigeoclonium helveticum
-
Insert Size (bp)2421
-
GenBank IDKF992040
- Promoter human synapsin promoter
-
Tag
/ Fusion Protein
- tdTomato (C terminal on insert)
Cloning Information
- Cloning method Restriction Enzyme
- 5′ cloning site NheI (not destroyed)
- 3′ cloning site KpnI (not destroyed)
- 5′ sequencing primer gcacgggcgcgaccatctgc
- 3′ sequencing primer TAGCGTAAAAGGAGCAACATAG
- (Common Sequencing Primers)
Resource Information
-
Article Citing this Plasmid
Terms and Licenses
-
Academic/Nonprofit Terms
-
Industry Terms
- Not Available to Industry
Trademarks:
- Zeocin® is an InvivoGen trademark.
Depositor Comments
The plasmid is fully sequenced in the coding sequence regions (opsin-fluorophore and important flanking regions). Multiple digestions were done to verify the vector structure. The construct and the virus were both tested in vitro.
These plasmids were created by your colleagues. Please acknowledge the Principal Investigator, cite the article in which the plasmids were described, and include Addgene in the Materials and Methods of your future publications.
-
For your Materials & Methods section:
pAAV-Syn-FLEX-rc [Chronos-tdTomato] was a gift from Edward Boyden (Addgene plasmid # 84481 ; http://n2t.net/addgene:84481 ; RRID:Addgene_84481) -
For your References section:
Independent optical excitation of distinct neural populations. Klapoetke NC, Murata Y, Kim SS, Pulver SR, Birdsey-Benson A, Cho YK, Morimoto TK, Chuong AS, Carpenter EJ, Tian Z, Wang J, Xie Y, Yan Z, Zhang Y, Chow BY, Surek B, Melkonian M, Jayaraman V, Constantine-Paton M, Wong GK, Boyden ES. Nat Methods. 2014 Mar;11(3):338-46. doi: 10.1038/nmeth.2836. Epub 2014 Feb 9. 10.1038/nmeth.2836 PubMed 24509633