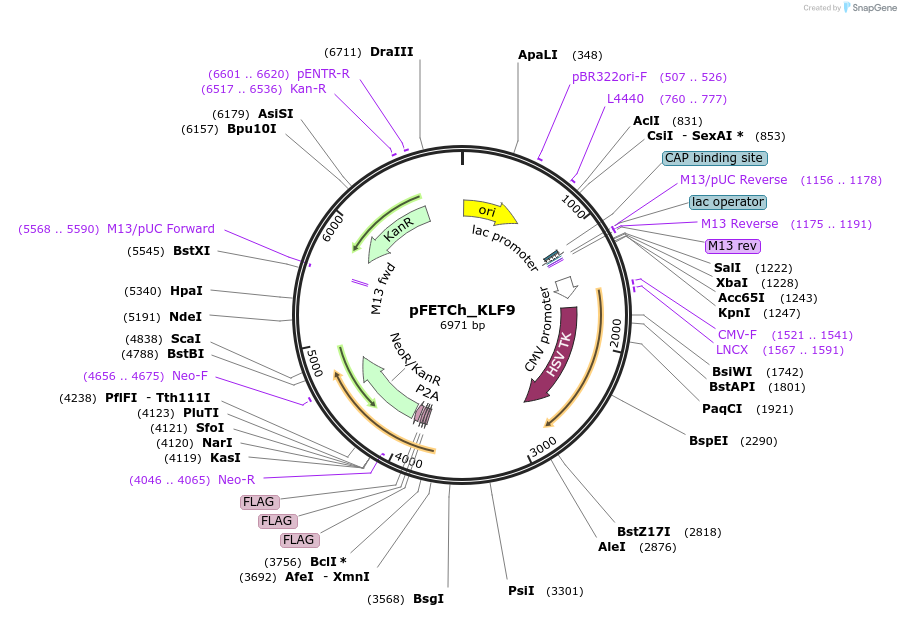
pFETCh_KLF9
(Plasmid
#86286)
-
PurposeDonor vector for 3' FLAG tag of human KLF9
-
Depositing Labs
-
Publication
-
Sequence Information
Ordering
| Item | Catalog # | Description | Quantity | Price (USD) | |
|---|---|---|---|---|---|
| Plasmid | 86286 | Standard format: Plasmid sent in bacteria as agar stab | 1 | $94 | |
Backbone
-
Vector backbonepFETCh_Donor
-
Vector typeCRISPR
Growth in Bacteria
-
Bacterial Resistance(s)Kanamycin, 50 μg/mL
-
Growth Temperature37°C
-
Growth Strain(s)DH5alpha
-
Copy numberHigh Copy
Gene/Insert
-
Gene/Insert nameKLF9 homology arms
-
SpeciesH. sapiens (human)
-
Entrez GeneKLF9 (a.k.a. BTEB, BTEB1)
-
Tag
/ Fusion Protein
- 3XFLAG-P2A-NeoR (C terminal on insert)
Cloning Information
- Cloning method Gibson Cloning
- 5′ sequencing primer acgcctgtgaaaccgtacta
- 3′ sequencing primer aactgttgggaagggcgatc
- (Common Sequencing Primers)
Terms and Licenses
-
Academic/Nonprofit Terms
-
Industry Terms
- Not Available to Industry
Trademarks:
- Zeocin® is an InvivoGen trademark.
Depositor Comments
This plasmid was co-transfected along with PX458_KLF9_1 and PX458_KLF9_2 into HepG2 cells, and genomic editing was verified by PCR. The donor nucleotide sequence in the final two codons of the KLF9 gene are modified to discourage sgRNA recognition, while retaining the native amino acid sequence. We performed ChIP-seq on this cell line using anti-FLAG Sigma F1804 and obtained data that passed our quality tests. Immunoprecipitation with anti-FLAG Sigma F1804 followed by mass spectrometry identified KLF9. All released data, including ChIP-seq files and validation data, are available at the ENCODE web portal: https://www.encodeproject.org/search/?searchTerm=flag&type=Experiment&lab.title=Richard+Myers%2C+HAIB . Note: This plasmid is part of the Mendenhall and Meyers CRISPR-based Tagging system to add a tag (currently using FLAG) to endogenous proteins. Please see https://www.addgene.org/crispr/tagging/ for more details.
These plasmids were created by your colleagues. Please acknowledge the Principal Investigator, cite the article in which the plasmids were described, and include Addgene in the Materials and Methods of your future publications.
-
For your Materials & Methods section:
pFETCh_KLF9 was a gift from Eric Mendenhall & Richard M. Myers (Addgene plasmid # 86286 ; http://n2t.net/addgene:86286 ; RRID:Addgene_86286) -
For your References section:
CETCh-seq: CRISPR epitope tagging ChIP-seq of DNA-binding proteins. Savic D, Partridge EC, Newberry KM, Smith SB, Meadows SK, Roberts BS, Mackiewicz M, Mendenhall EM, Myers RM. Genome Res. 2015 Sep 9. 10.1101/gr.193540.115 PubMed 26355004