We narrowed to 11,999 results for: ARIA
-
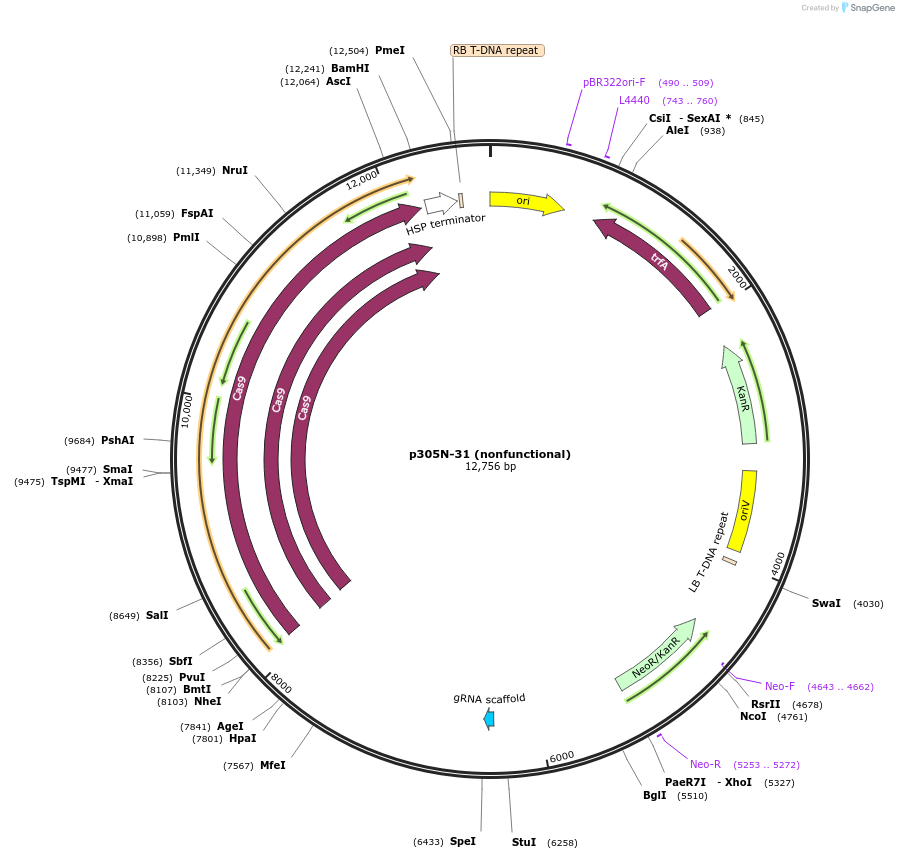
Plasmid#246314PurposeEvaluation of PtU6.2c4 promoter (nonfunctional) (Pol III promoter) sequence variation for CRISPR-Cas9 editing of mEGFP in a Nicotiana benthamiana reporter lineDepositorInsertPtU6.2c4 promoter (nonfunctional)
UseCRISPRExpressionPlantMutationdeletions: -A (-41 from TSS), -T (-30), -T (-29);…PromoterPtU6.2c4 promoter (nonfunctional)Available SinceOct. 30, 2025AvailabilityAcademic Institutions and Nonprofits only -
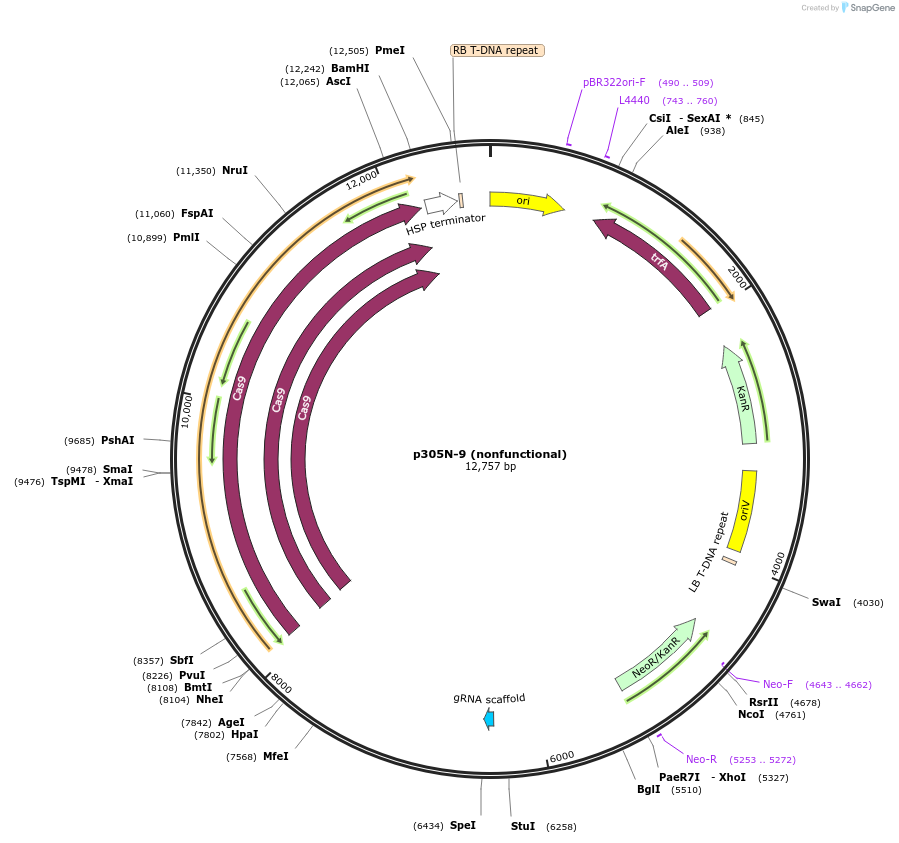
p305N-9 (nonfunctional)
Plasmid#246292PurposeEvaluation of AtU6.29c13 promoter (nonfunctional) (Pol III promoter) sequence variation for CRISPR-Cas9 editing of mEGFP in a Nicotiana benthamiana reporter lineDepositorInsertAtU6.29c13 promoter (nonfunctional)
UseCRISPRExpressionPlantMutationdeletions: -A (-58 from TSS), -T (-57) within USEPromoterAtU6.29c13 promoter (nonfunctional)Available SinceOct. 30, 2025AvailabilityAcademic Institutions and Nonprofits only -
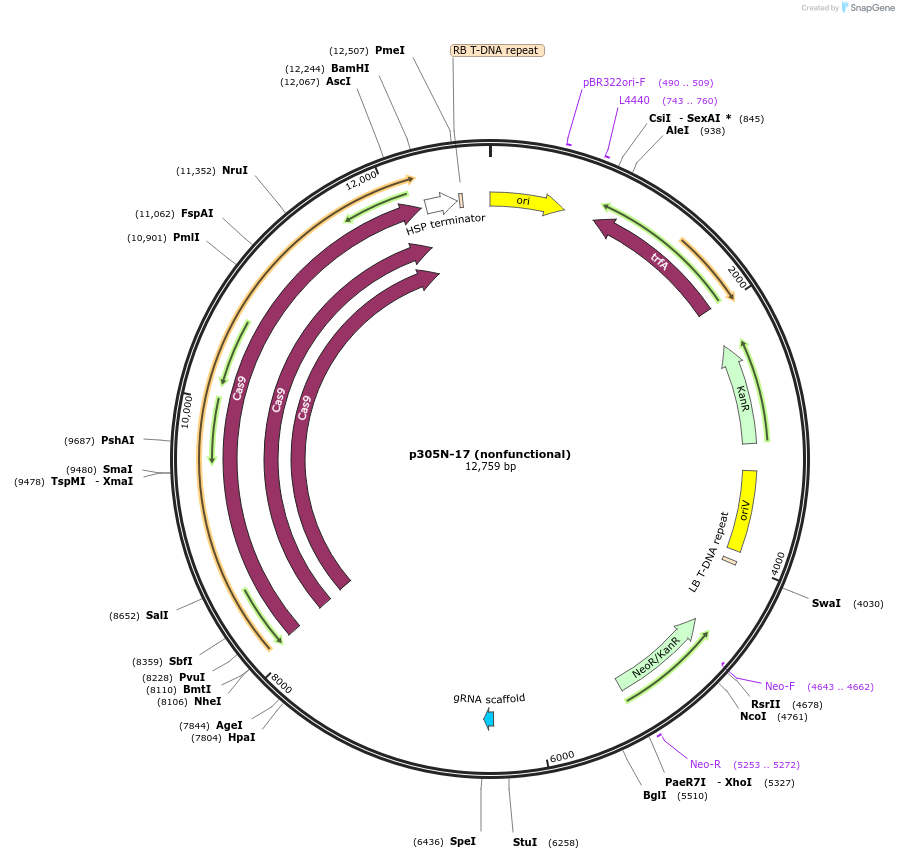
p305N-17 (nonfunctional)
Plasmid#246300PurposeEvaluation of HbU6.2m3 promoter (nonfunctional) (Pol III promoter) sequence variation for CRISPR-Cas9 editing of mEGFP in a Nicotiana benthamiana reporter lineDepositorInsertHbU6.2m3 promoter (nonfunctional)
UseCRISPRExpressionPlantMutationC-to-T (-29 from TSS), G-to-A (-24)PromoterHbU6.2m3 promoter (nonfunctional)Available SinceOct. 30, 2025AvailabilityAcademic Institutions and Nonprofits only -
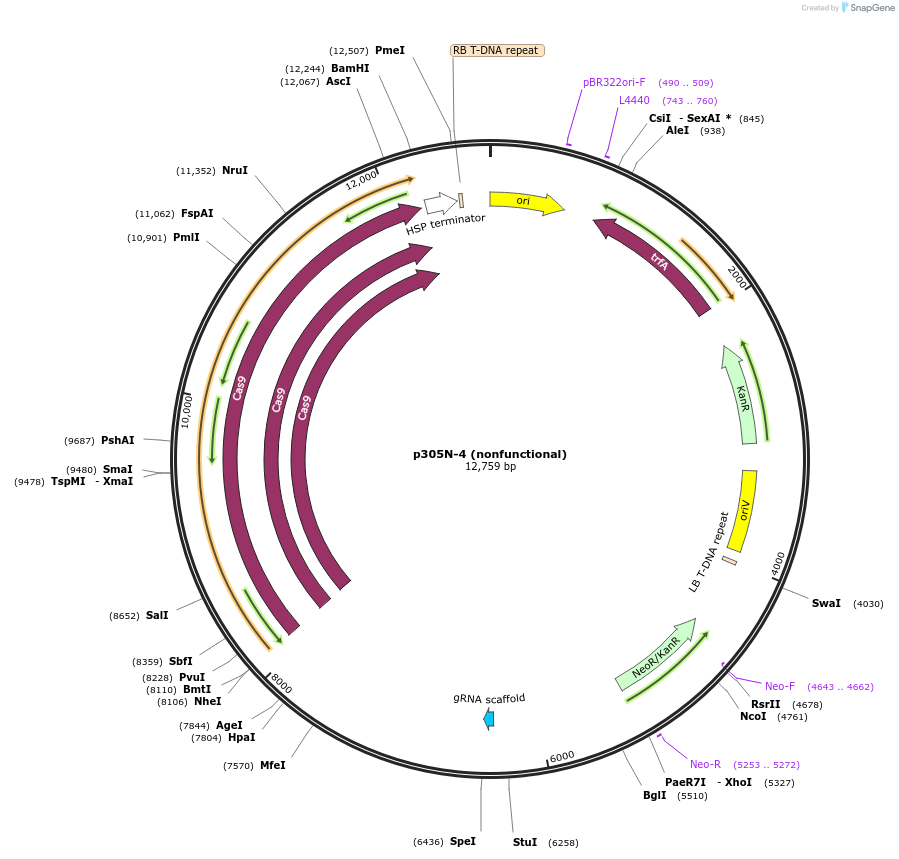
p305N-4 (nonfunctional)
Plasmid#246287PurposeEvaluation of AtU6.1c1 promoter (nonfunctional) (Pol III promoter) sequence variation for CRISPR-Cas9 editing of mEGFP in a Nicotiana benthamiana reporter lineDepositorInsertAtU6.1c1 promoter (nonfunctional)
UseCRISPRExpressionPlantMutationG-to-C at -15 from TSSPromoterAtU6.1c1 promoter (nonfunctional)Available SinceOct. 30, 2025AvailabilityAcademic Institutions and Nonprofits only -
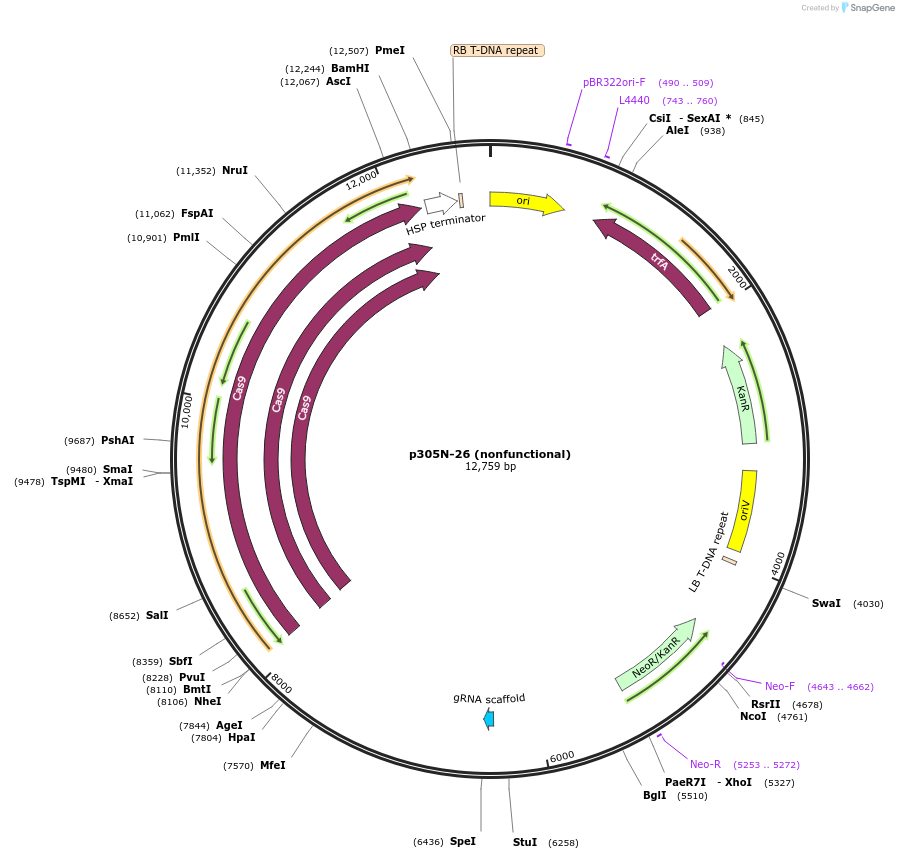
p305N-26 (nonfunctional)
Plasmid#246309PurposeEvaluation of MtU6.6m7 promoter (nonfunctional) (Pol III promoter) sequence variation for CRISPR-Cas9 editing of mEGFP in a Nicotiana benthamiana reporter lineDepositorInsertMtU6.6m7 promoter (nonfunctional)
UseCRISPRExpressionPlantMutationC-to-A (-63 from TSS), T-to-G (-57)PromoterMtU6.6m7 promoter (nonfunctional)Available SinceOct. 30, 2025AvailabilityAcademic Institutions and Nonprofits only -
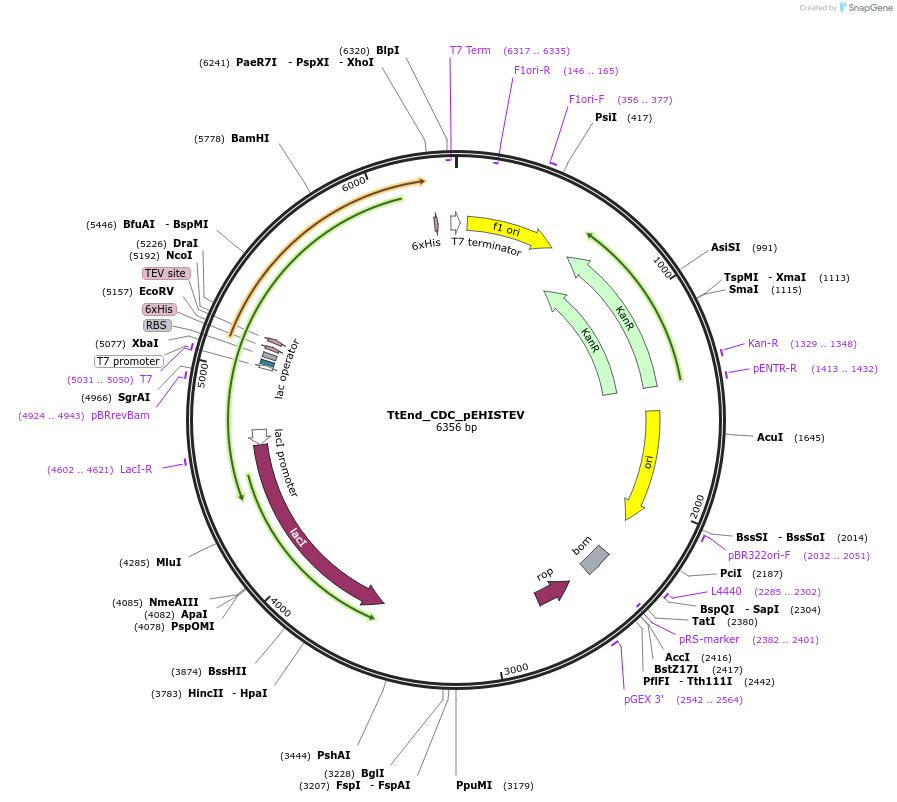
TtEnd_CDC_pEHISTEV
Plasmid#179736PurposeTtEnd_CDC_pEHISTEV expresses tuncated variant of TtEnd5A (186-531 amni caids) comprsisng of cataytic domain and a linker at C-terminal.DepositorInsertTtEnd5A
Tags6xHisExpressionBacterialMutation185 Amino acids from N-terminal were deleted.PromoterT7Available SinceOct. 14, 2025AvailabilityAcademic Institutions and Nonprofits only -
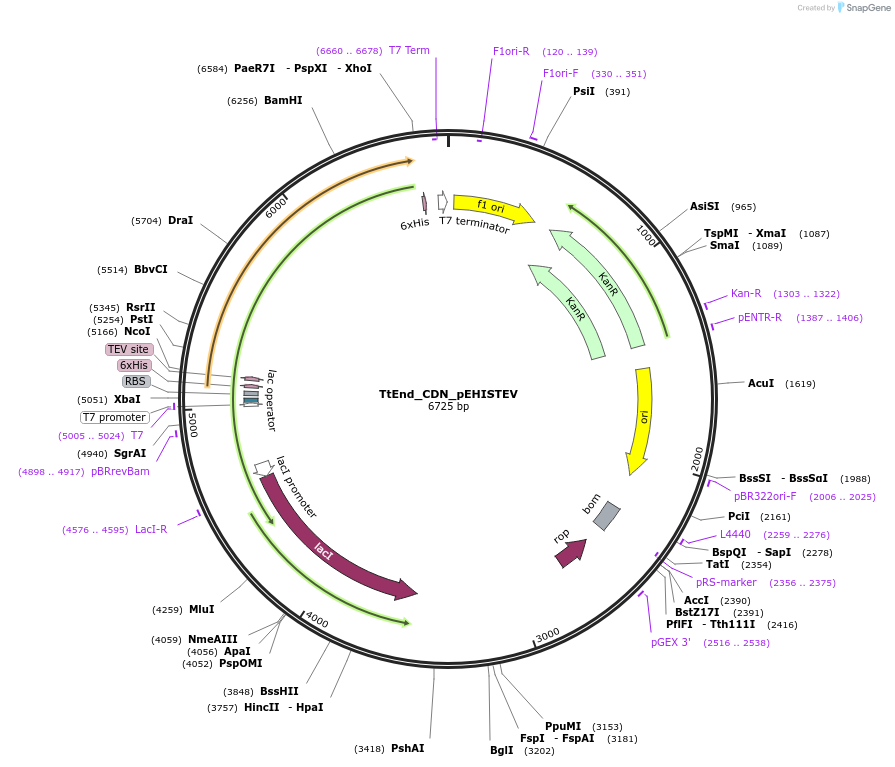
TtEnd_CDN_pEHISTEV
Plasmid#179738PurposeTtEnd_CDN_pEHISTEV expresses tuncated variant of TtEnd5A (19-486 amnino acids) comprsisng of cataytic domain and a linker at N-terminal.DepositorInsertTtEnd5A
Tags6xHisExpressionBacterialMutation45 Amino acids from C-terminal were deleted.PromoterT7Available SinceOct. 14, 2025AvailabilityAcademic Institutions and Nonprofits only -
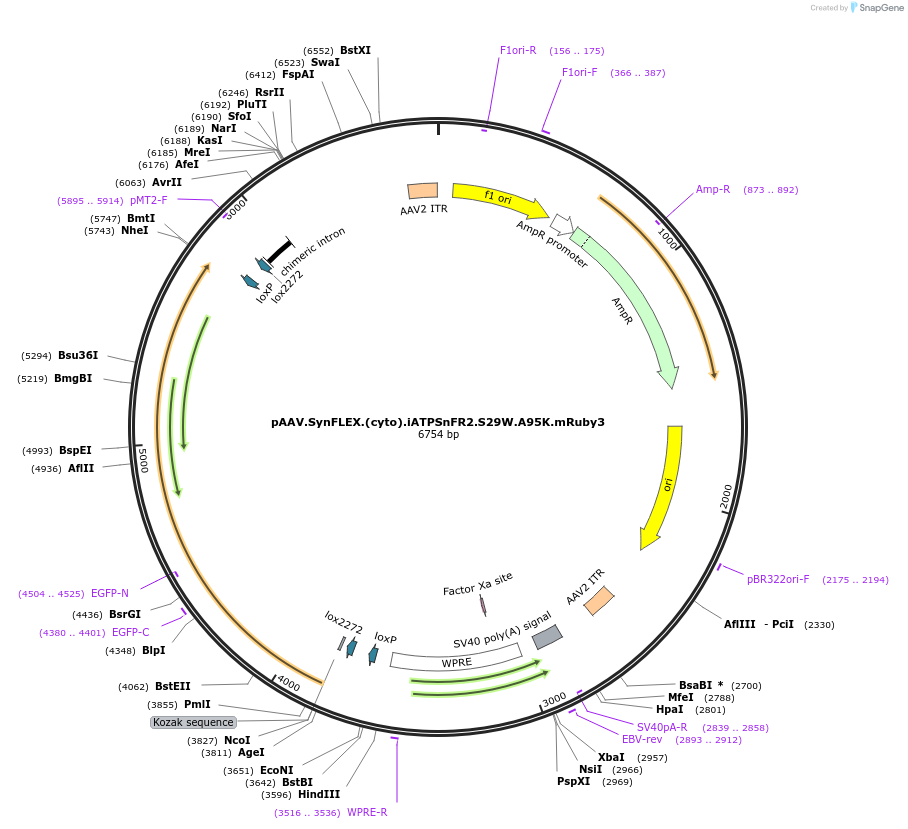
pAAV.SynFLEX.(cyto).iATPSnFR2.S29W.A95K.mRuby3
Plasmid#244911PurposeiATPSnFR2.S29W.A95K variant with non-responsive mRuby3DepositorInsertiATPSnFR2.S29W.A95K.mRuby3
UseAAVMutationS29W.A95KAvailable SinceOct. 8, 2025AvailabilityAcademic Institutions and Nonprofits only -
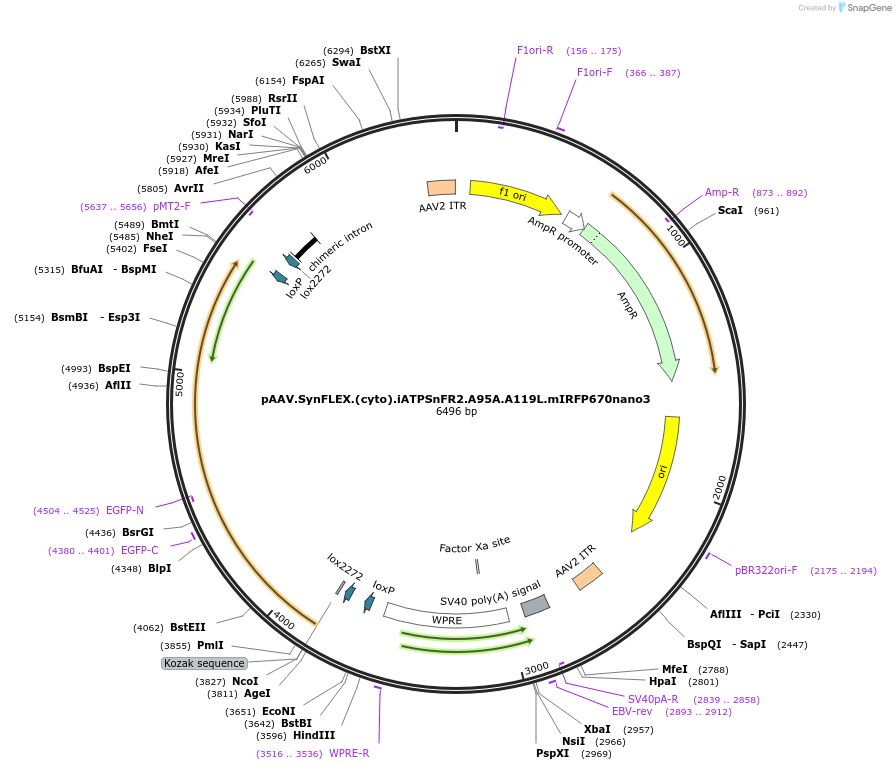
pAAV.SynFLEX.(cyto).iATPSnFR2.A95A.A119L.mIRFP670nano3
Plasmid#244917PurposeiATPSnFR2.A95A.A119L variant with non-responsive mIRFP670nano3DepositorInsertiATPSnFR2.A95A.A119L.mIRFP670nano3
UseAAVMutationA95A.A119LAvailable SinceOct. 8, 2025AvailabilityAcademic Institutions and Nonprofits only -
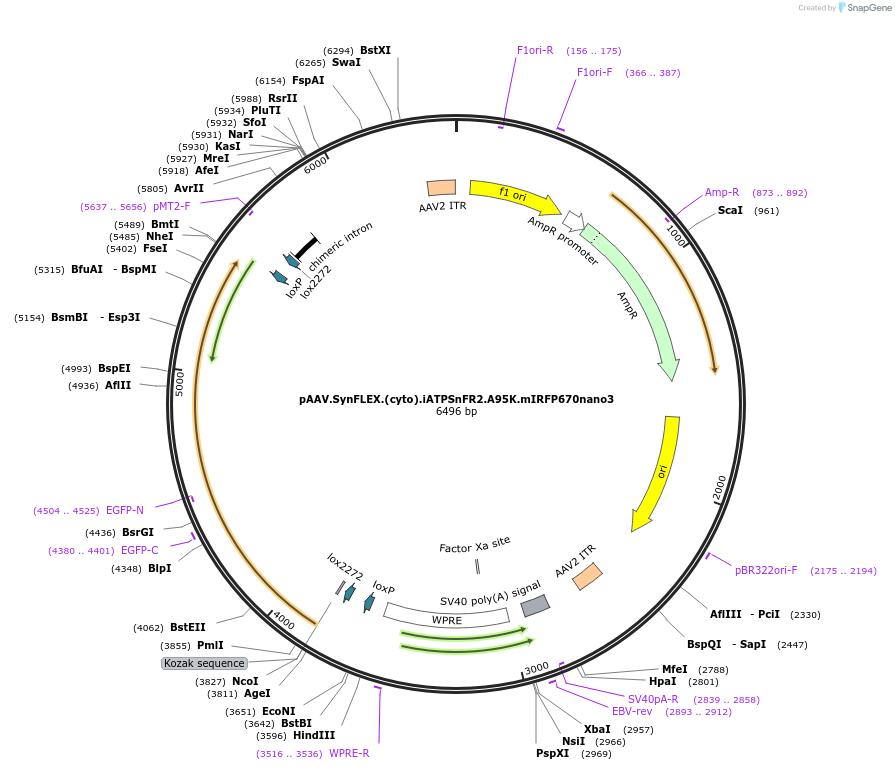
pAAV.SynFLEX.(cyto).iATPSnFR2.A95K.mIRFP670nano3
Plasmid#244916PurposeiATPSnFR2.A95K variant with non-responsive mIRFP670nano3DepositorInsertiATPSnFR2.A95K.mIRFP670nano3
UseAAVMutationA95KAvailable SinceOct. 8, 2025AvailabilityAcademic Institutions and Nonprofits only