We narrowed to 42,359 results for: LAT
-
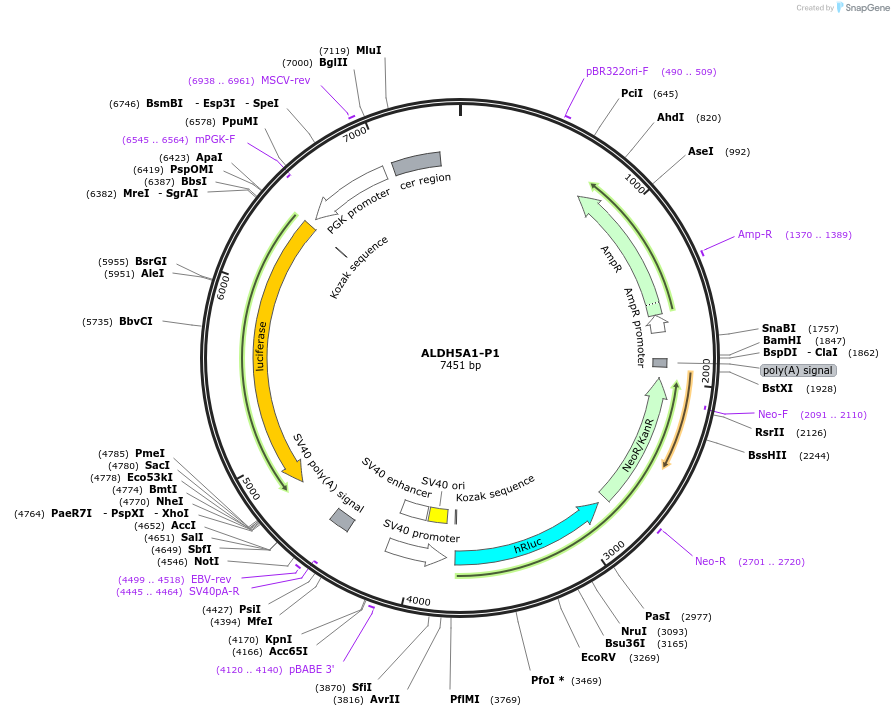
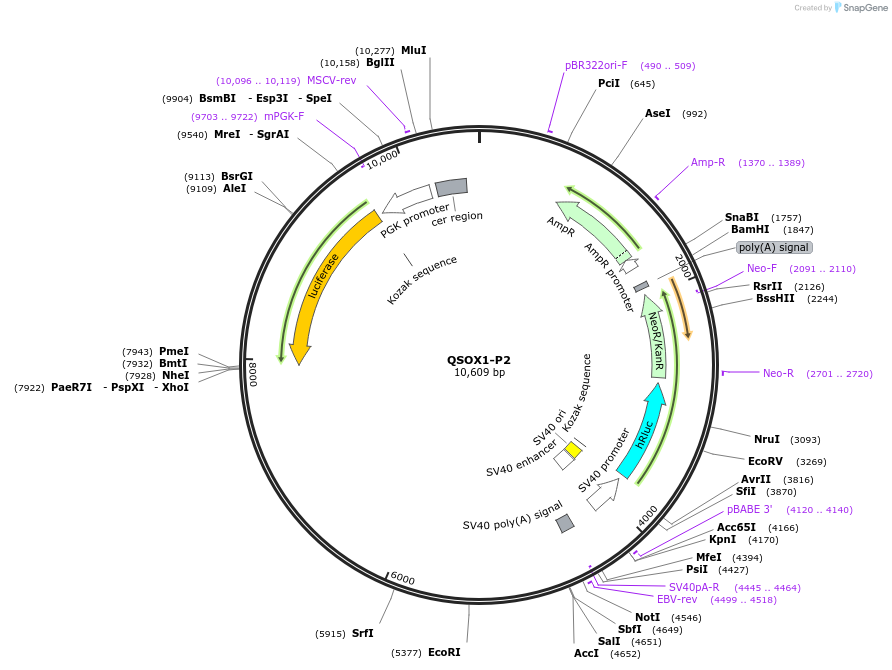
Plasmid#236020PurposeStudy the role and mechanism of the 3' UTR in regulating ALDH5A1-P1 gene function.DepositorInsertALDH5A1-P1 3'UTR (ALDH5A1 Human)
ExpressionMammalianAvailable SinceJan. 20, 2026AvailabilityAcademic Institutions and Nonprofits only -
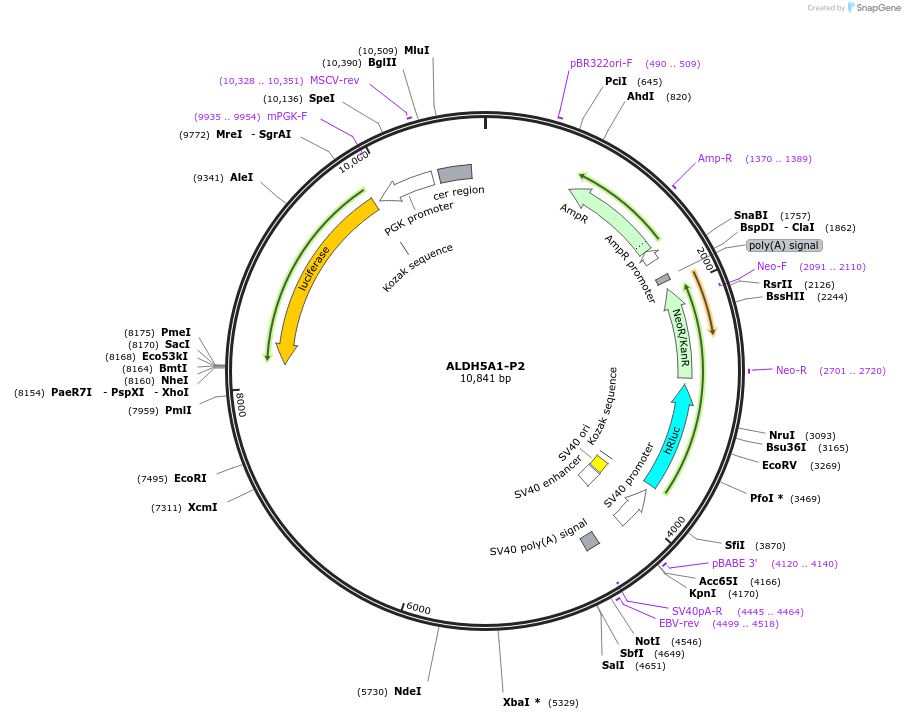
ALDH5A1-P2
Plasmid#236021PurposeStudy the role and mechanism of the 3' UTR in regulating ALDH5A1-P2 gene function.DepositorInsertALDH5A1-P2 3'UTR (ALDH5A1 Human)
ExpressionMammalianAvailable SinceJan. 20, 2026AvailabilityAcademic Institutions and Nonprofits only -
-
-
-
-
-
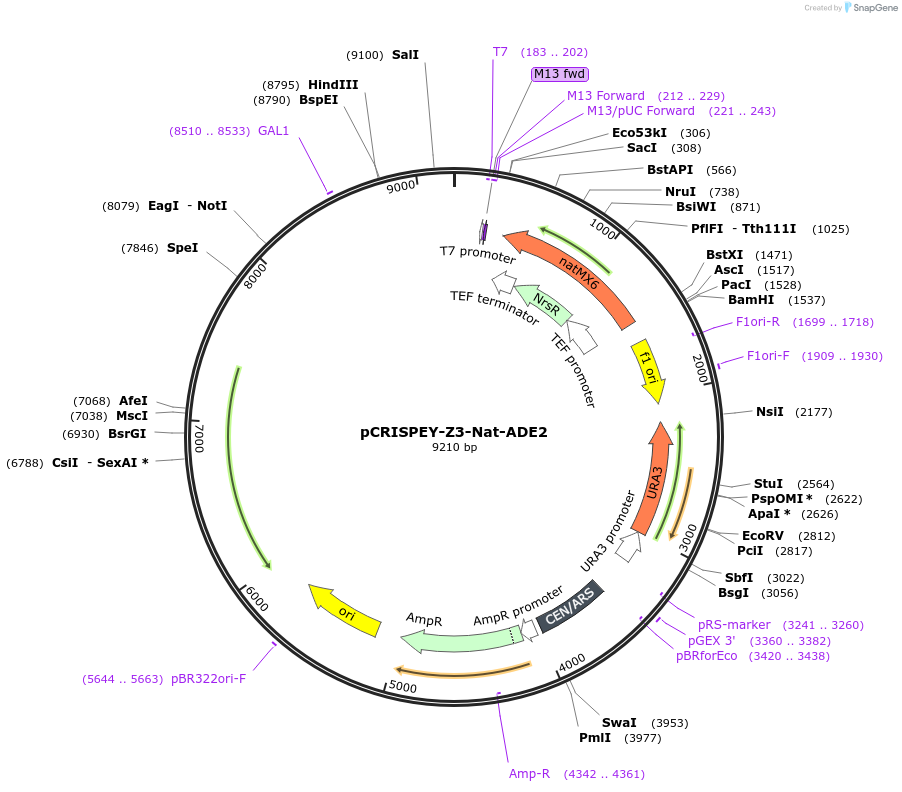
pCRISPEY-Z3-Nat-ADE2
Plasmid#232106PurposeExpresses estradiol-inducible CRISPR guide RNA and repair template for creating a frameshift mutation in ADE2 gene of yeast.DepositorInsertADE2 gRNA and repair template
UseCRISPRExpressionYeastMutationRepair template introduces C at nt 466 of ADE2 to…PromoterZ3Available SinceFeb. 21, 2025AvailabilityAcademic Institutions and Nonprofits only -
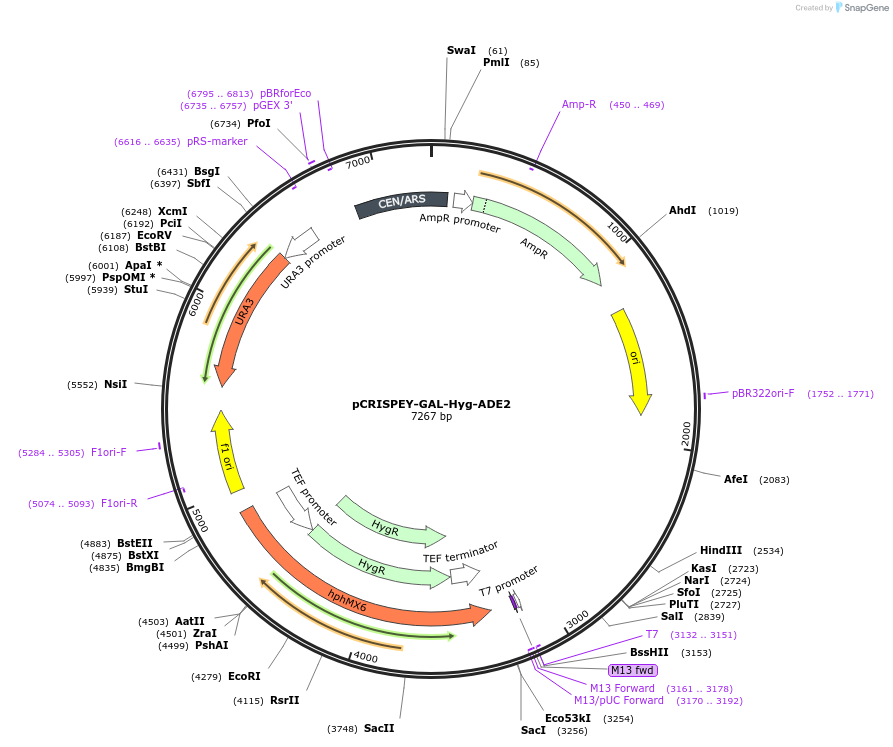
pCRISPEY-GAL-Hyg-ADE2
Plasmid#232104PurposeExpresses galactose-inducible CRISPR guide RNA and repair template for creating a frameshift mutation in ADE2 gene of yeast.DepositorInsertADE2 gRNA and repair template
UseCRISPRExpressionYeastMutationRepair template introduces C at nt 466 of ADE2 to…PromoterGAL7Available SinceFeb. 21, 2025AvailabilityAcademic Institutions and Nonprofits only -
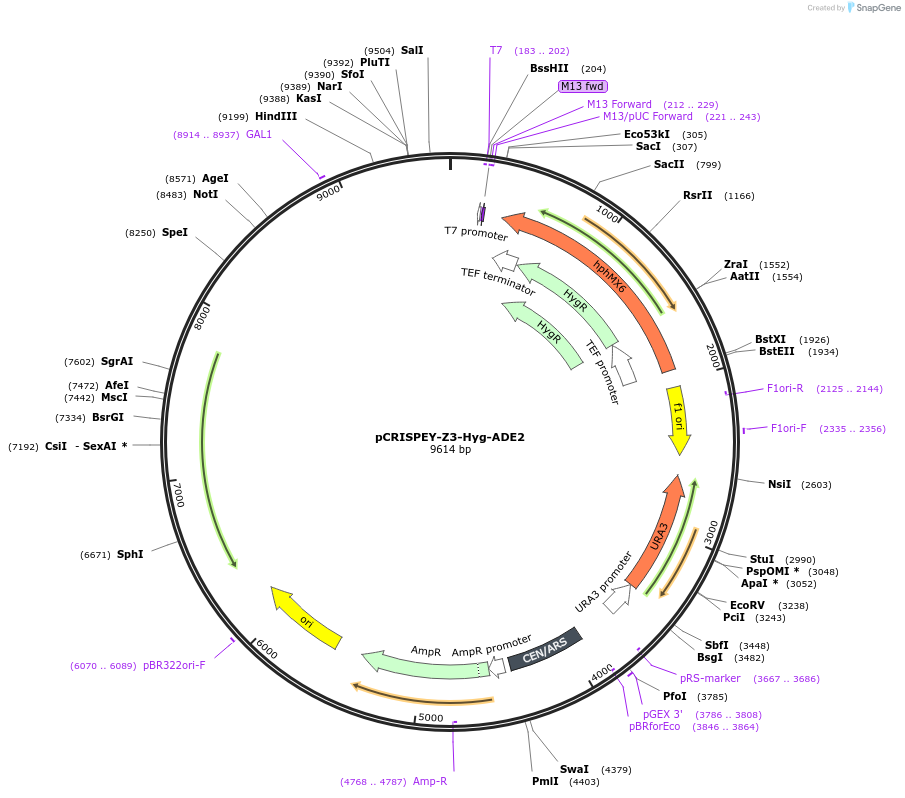
pCRISPEY-Z3-Hyg-ADE2
Plasmid#232107PurposeExpresses estradiol-inducible CRISPR guide RNA and repair template for creating a frameshift mutation in ADE2 gene of yeast.DepositorInsertADE2 gRNA and repair template
UseCRISPRExpressionYeastMutationRepair template introduces C at nt 466 of ADE2 to…PromoterZ3Available SinceFeb. 21, 2025AvailabilityAcademic Institutions and Nonprofits only -
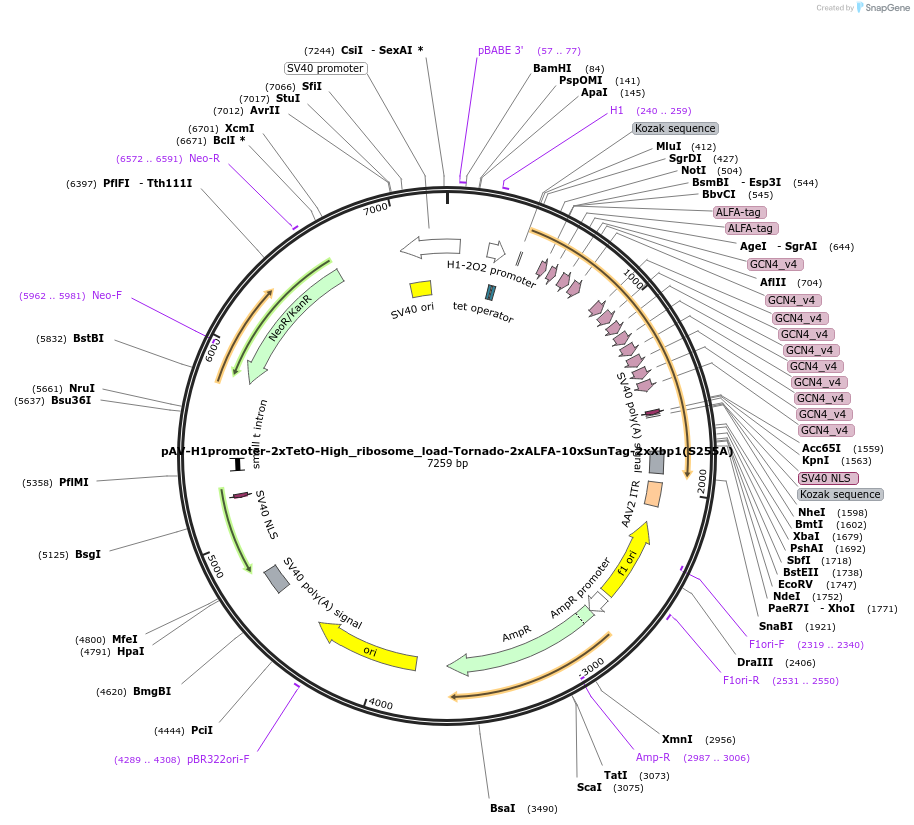
pAV-H1promoter-2xTetO-High_ribosome_load-Tornado-2xALFA-10xSunTag-2xXbp1(S255A)
Plasmid#231596PurposeExpress high ribosome loading circular RNA translation reporter 2xALFATag-10xSunTag-2xXbp1(S255A) pause site under H1 promoterDepositorInsertKozak sequence-Tornado-2xALFATag-10xSunTag-2xXbp1(S255A) pause sequence
ExpressionMammalianAvailable SinceJan. 31, 2025AvailabilityAcademic Institutions and Nonprofits only -
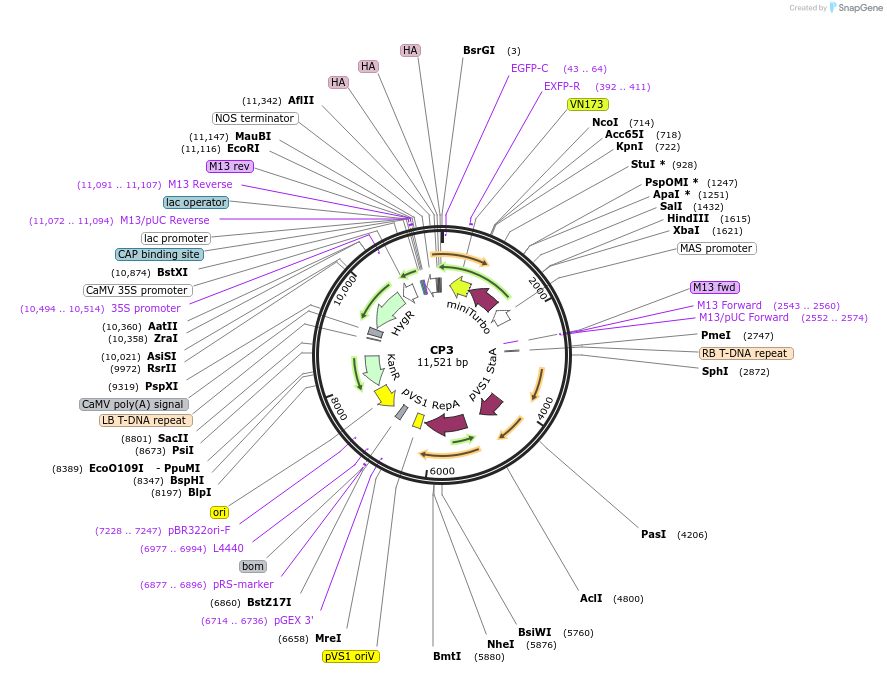
CP3
Plasmid#223335PurposeMapping of cytosol-facing chloroplast outer membrane proximity proteome by proximity-dependent biotinylation in living Arabidopsis cellsDepositorInsertToc64-miniTurboID-EGFP-3HA
TagsminiTurboID-EGFP-3HAExpressionPlantPromoterCaMV35SAvailable SinceJan. 21, 2025AvailabilityAcademic Institutions and Nonprofits only -
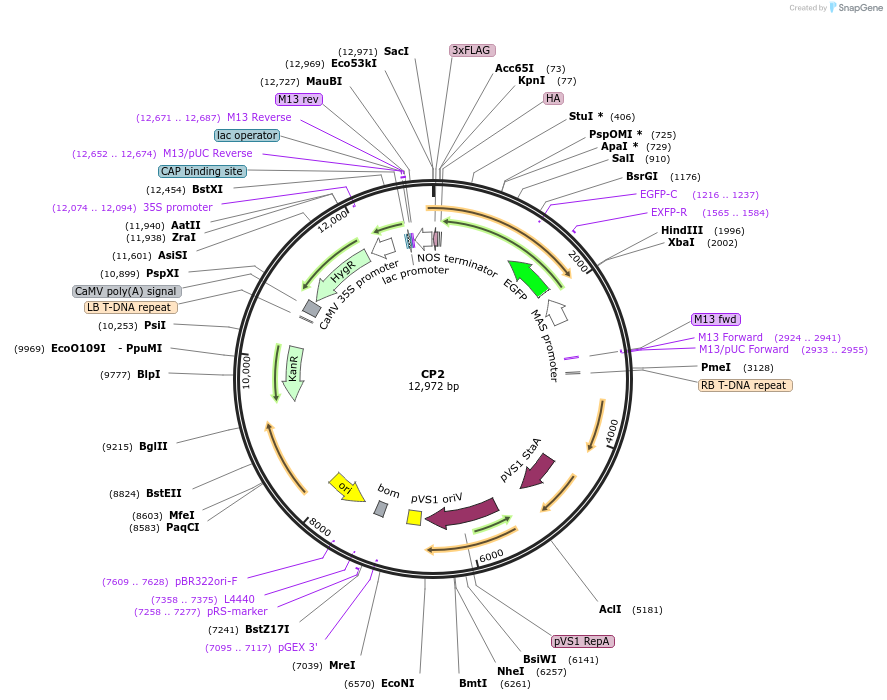
CP2
Plasmid#223334PurposeMapping of cytosol-facing chloroplast outer membrane proximity proteome by proximity-dependent biotinylation in living Arabidopsis cellsDepositorInsertToc64-EGFP-TurboID-3HA
TagsEGFP-TurboID-3HAExpressionPlantPromoterCaMV35SAvailable SinceJan. 21, 2025AvailabilityAcademic Institutions and Nonprofits only -
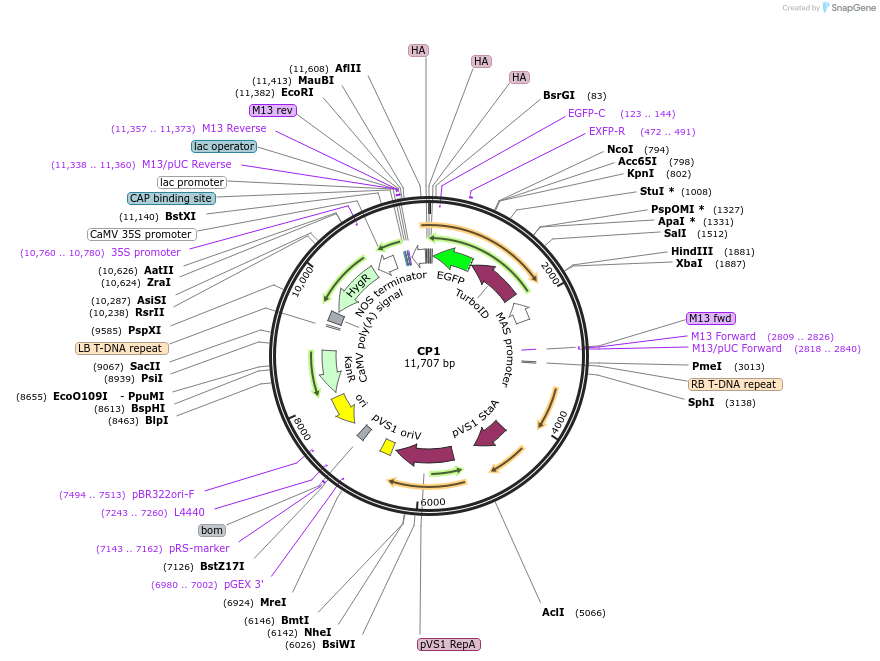
CP1
Plasmid#223333PurposeMapping of cytosol-facing chloroplast outer membrane proximity proteome by proximity-dependent biotinylation in living Arabidopsis cellsDepositorInsertToc64-TurboID-EGFP-3HA
TagsTurboID-EGFP-3HAExpressionPlantPromoterCaMV35SAvailable SinceJan. 21, 2025AvailabilityAcademic Institutions and Nonprofits only -
-
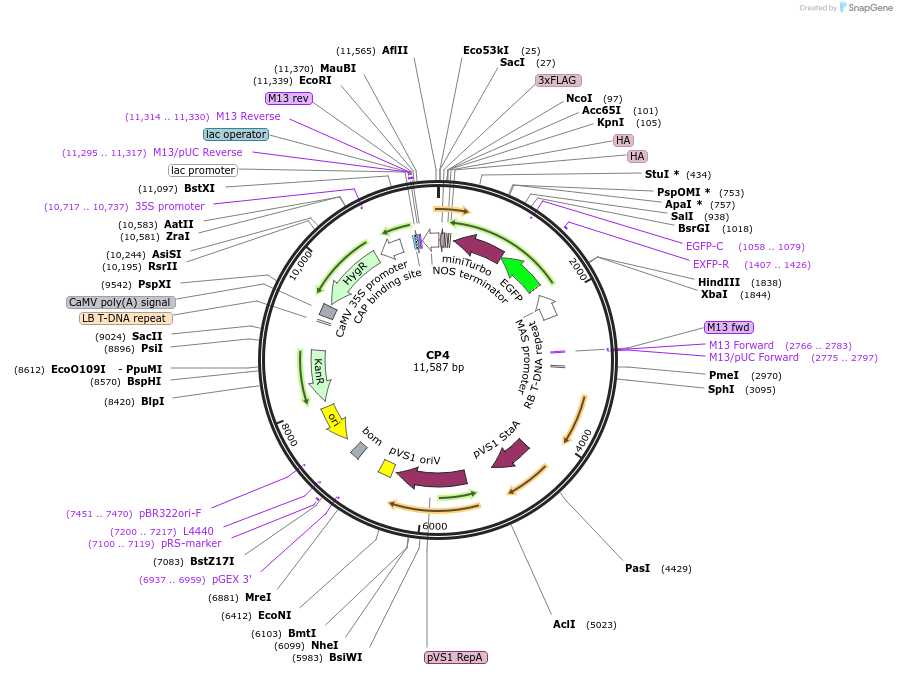
CP4
Plasmid#223336PurposeMapping of cytosol-facing chloroplast outer membrane proximity proteome by proximity-dependent biotinylation in living Arabidopsis cellsDepositorInsertToc64-EGFP-miniTurboID-3HA
TagsEGFP-miniTurboID-3HAExpressionPlantPromoterCaMV35SAvailable SinceDec. 9, 2024AvailabilityAcademic Institutions and Nonprofits only -
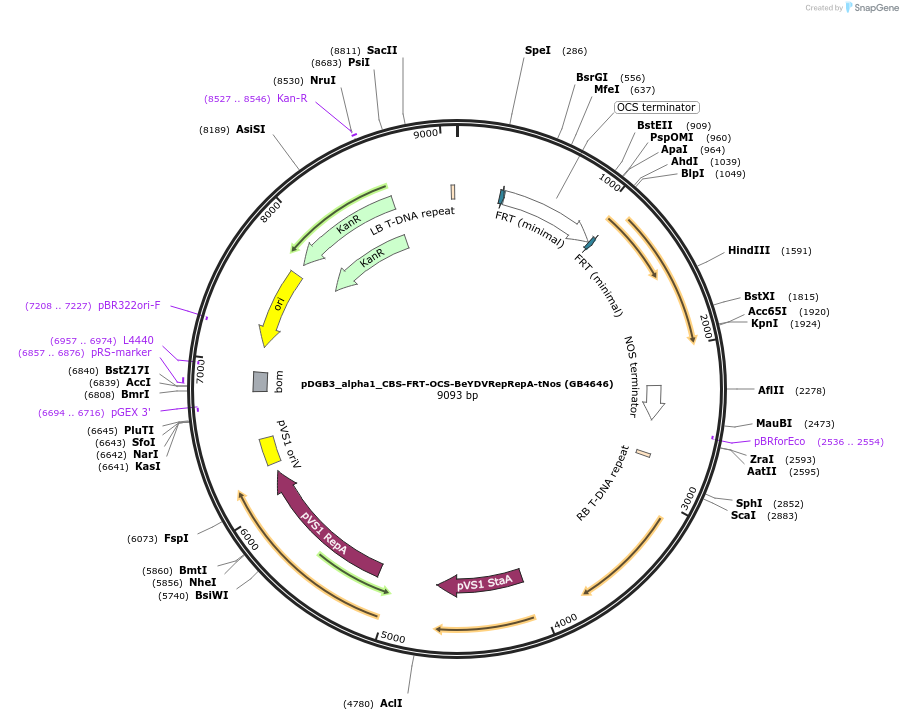
pDGB3_alpha1_CBS-FRT-OCS-BeYDVRepRepA-tNos (GB4646)
Plasmid#225438PurposeTranscriptional unit for copper- and flipase-regulated BeYDV Rep/RepA. OCSt, flanked by FRT sites, is embedded in the 5'UTR of miniDFR.DepositorInsertCBS-FRT-OCS-BeYDVRepRepA-tNos
UseSynthetic BiologyMutationBsaI and BsmBI sites removedAvailable SinceOct. 3, 2024AvailabilityAcademic Institutions and Nonprofits only -
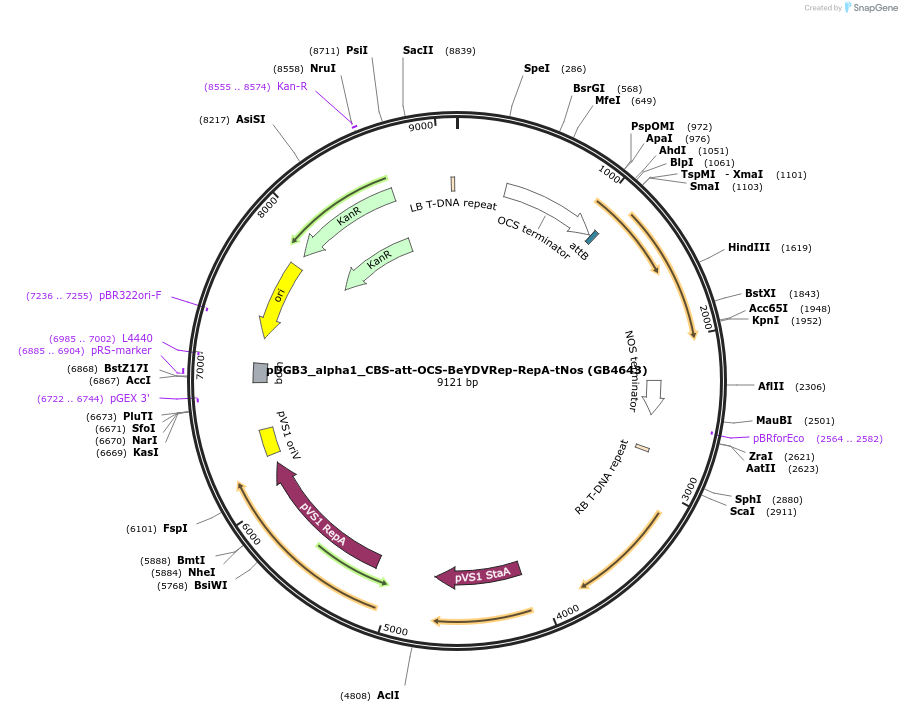
pDGB3_alpha1_CBS-att-OCS-BeYDVRep-RepA-tNos (GB4643)
Plasmid#225437PurposeTranscriptional unit for the copper and PhiC31 regulated expression of BeYDV Rep/RepA.DepositorInsertCBS-att-OCS-BeYDVRep/RepA-tNos
UseSynthetic BiologyMutationBsaI and BsmBI sites removedAvailable SinceOct. 3, 2024AvailabilityAcademic Institutions and Nonprofits only -
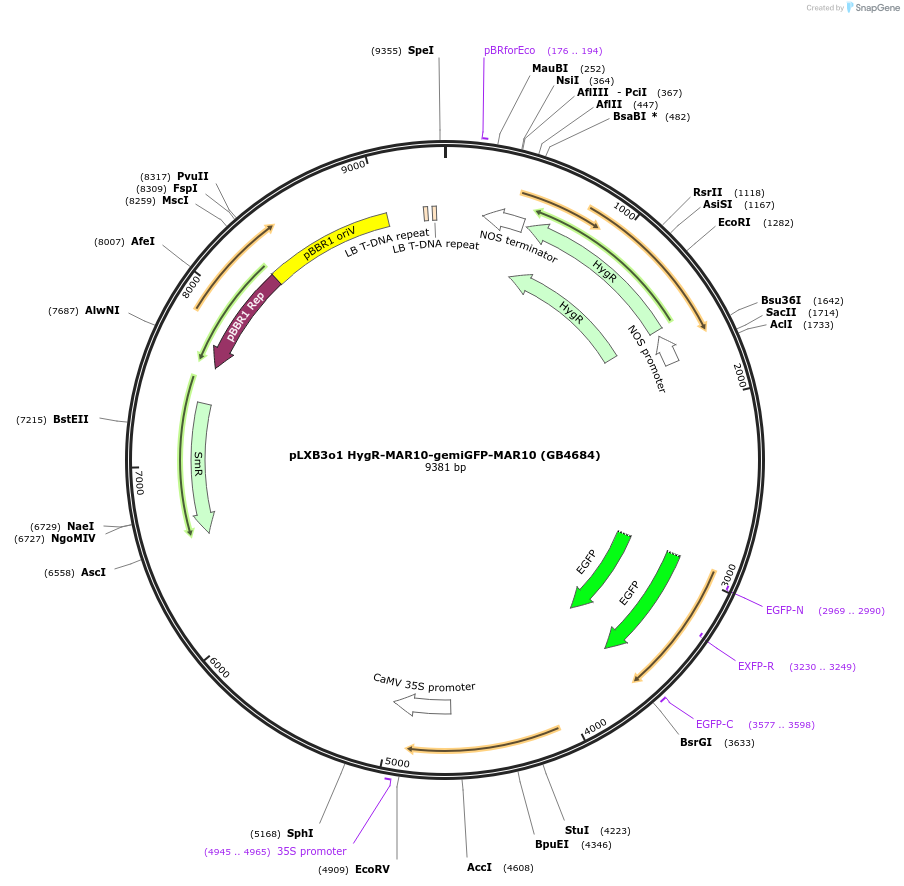
pLXB3o1 HygR-MAR10-gemiGFP-MAR10 (GB4684)
Plasmid#225439PurposeModule for stable transformation of Geminino 1.0-eGFP flanked by two MAR10 insulators and next to a Hygromycin resistance cassette.DepositorInsertHygR-MAR10-gemiGFP-MAR10
UseSynthetic BiologyMutationBsaI and BsmBI sites removedAvailable SinceOct. 3, 2024AvailabilityAcademic Institutions and Nonprofits only -
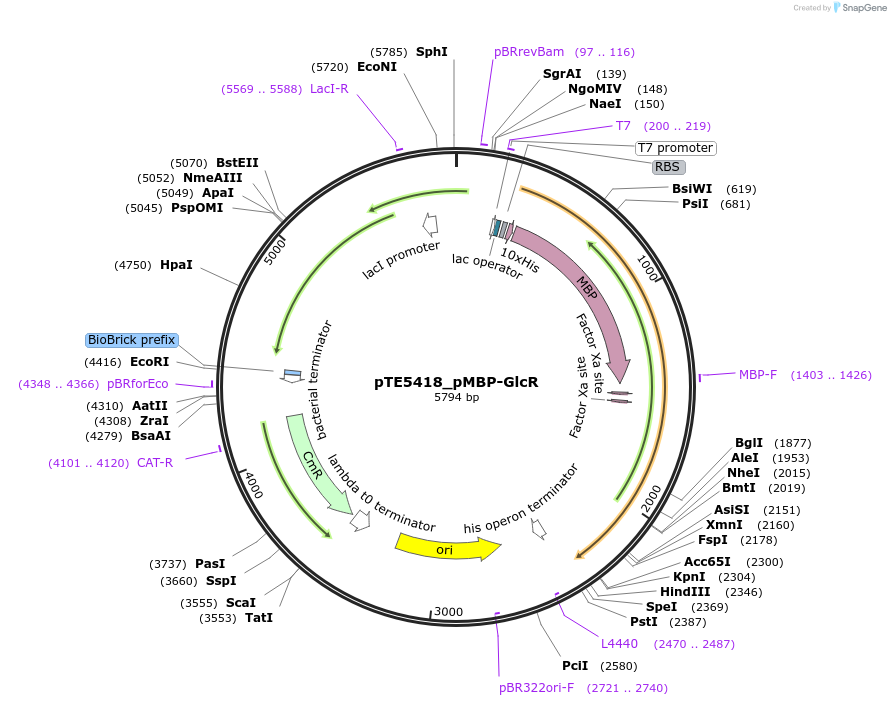
pTE5418_pMBP-GlcR
Plasmid#221192PurposeExpression vector for 10x-MBP-GlcR (MGlcR). GlcR is a glycolate-responsive transcriptional repressor and the gene product of pden4400 from Paracoccus denitrificans, codon optimized for E. coli.DepositorInsertglcR (pden4400 from Paracoccus denitrificans)
Tags10x His tag with E. coli maltose binding protein …ExpressionBacterialPromoterT7 promoter and T7 promoter-lacOAvailable SinceSept. 25, 2024AvailabilityAcademic Institutions and Nonprofits only