We narrowed to 19,179 results for: comp
-
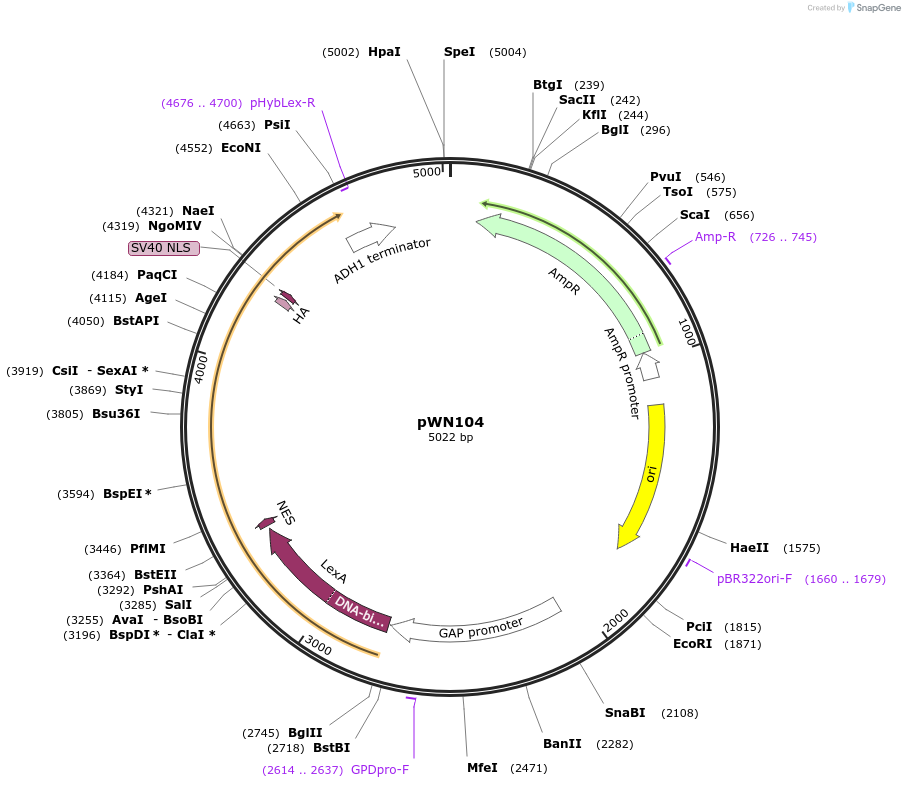
Plasmid#233861PurposeMoClo-YTK; ConLS/R1 cassette; NS3 systems; pTDH3-LexA-NES-NS3-V2-NLS-VP48-2XOaf1-tADH1DepositorInsertpTDH3-LexA-NES-NS3-V2-NLS-VP48-2XOaf1-tADH1
ExpressionYeastAvailable SinceMay 22, 2025AvailabilityAcademic Institutions and Nonprofits only -
pAL004
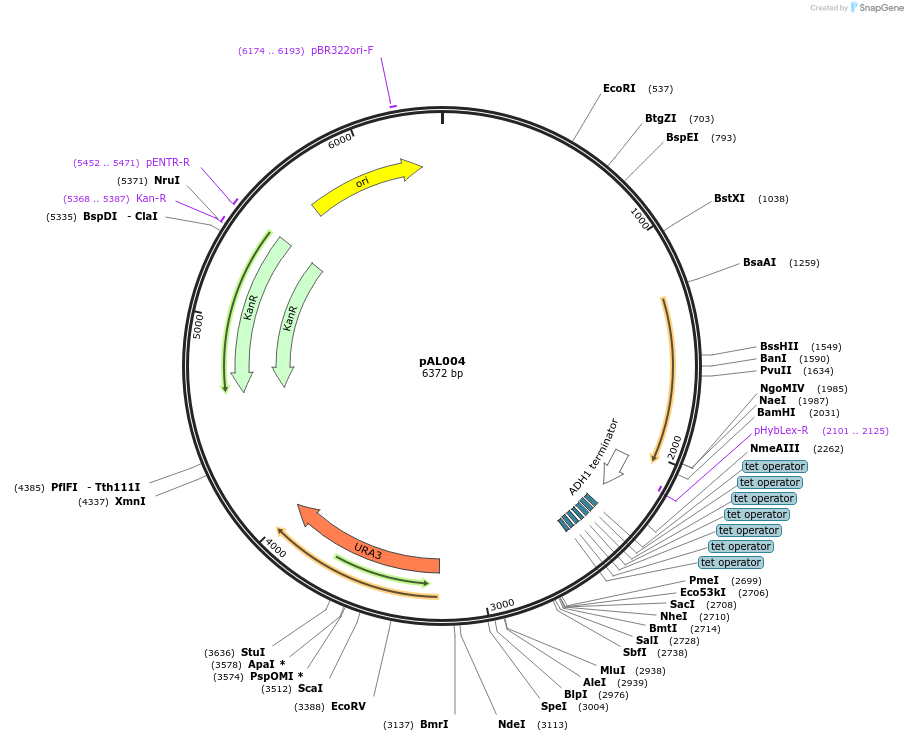
Plasmid#233863PurposeMoClo-YTK URA multigene plasmid with MCS; Tet systems; pRNR2-rtTA-v16-tADH1; tetO7-pLEU2m-MCS-tENO2DepositorInsertpRNR2-rtTA-v16-tADH1; tetO7-pLEU2m-MCS-tENO2
ExpressionYeastAvailable SinceMay 22, 2025AvailabilityAcademic Institutions and Nonprofits only -
pAL003
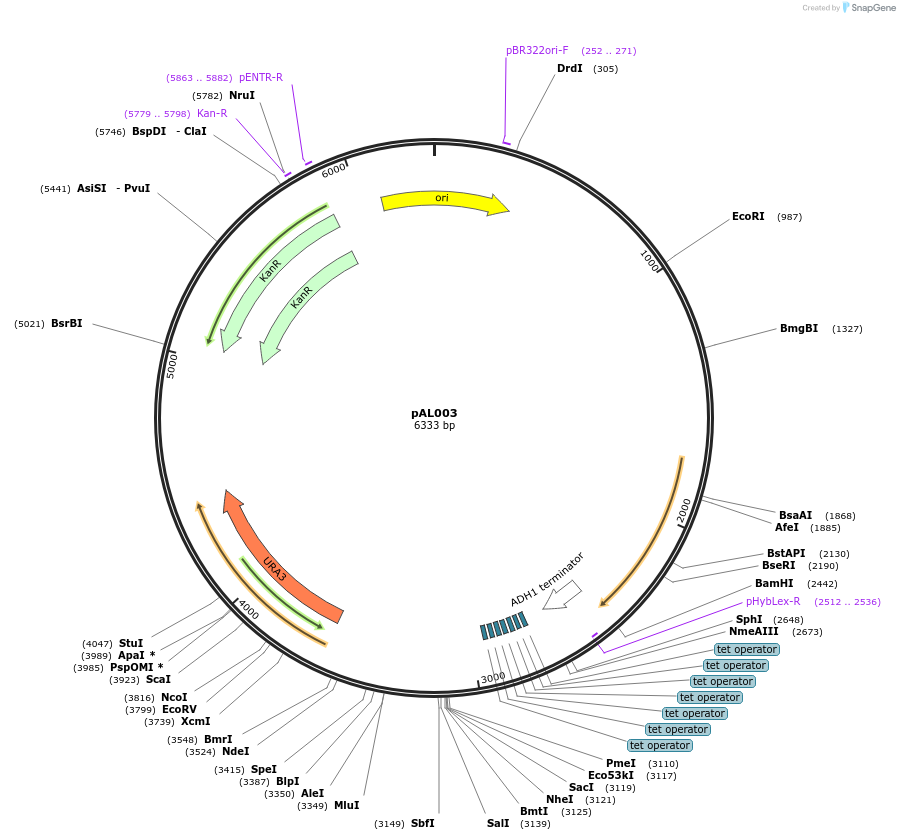
Plasmid#233864PurposeMoClo-YTK URA multigene plasmid with MCS; Tet systems; pRPL18B-rtTA3-tADH1; tetO7-pLEU2m-MCS-tENO2DepositorInsertpRPL18B-rtTA3-tADH1; tetO7-pLEU2m-MCS-tENO2
ExpressionYeastAvailable SinceMay 22, 2025AvailabilityAcademic Institutions and Nonprofits only -
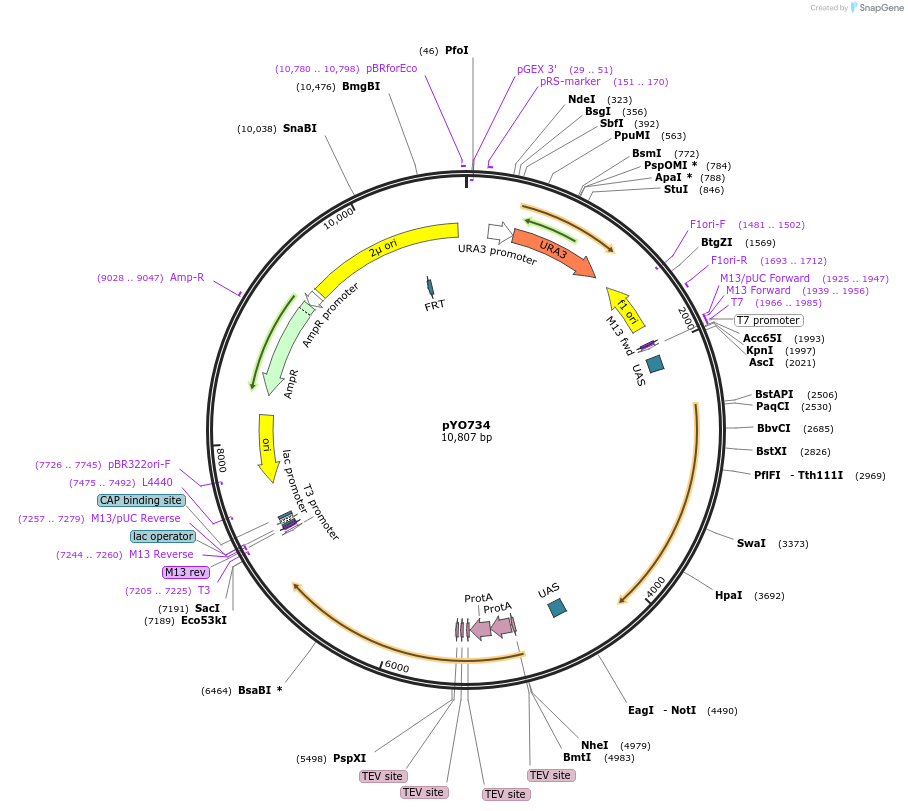
pYO734
Plasmid#235753PurposeProtein expression of ScVPS30 and ZZ-ScVPS38DepositorAvailable SinceMay 14, 2025AvailabilityAcademic Institutions and Nonprofits only -
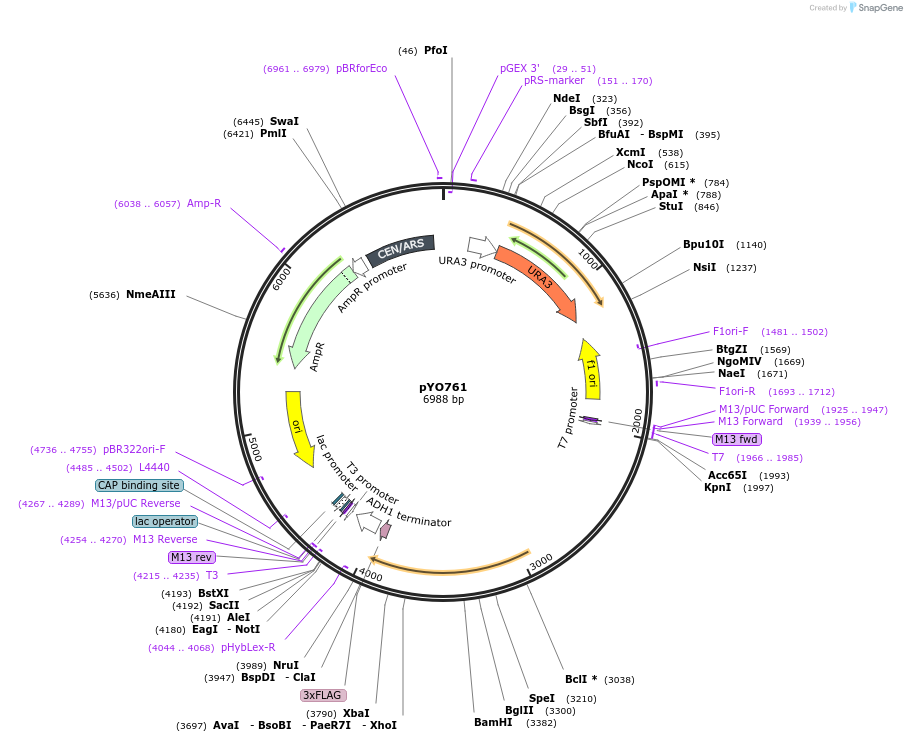
pYO761
Plasmid#235760PurposeExpression of ScVPS38 delta BARA2 (320-440)-x3FLAG under its own promoterDepositorInsertVSP38 delta BARA2
Tags3xFLAGExpressionYeastMutationaa 320-440 deletedPromoterScVPS38 promoterAvailable SinceMay 13, 2025AvailabilityAcademic Institutions and Nonprofits only -
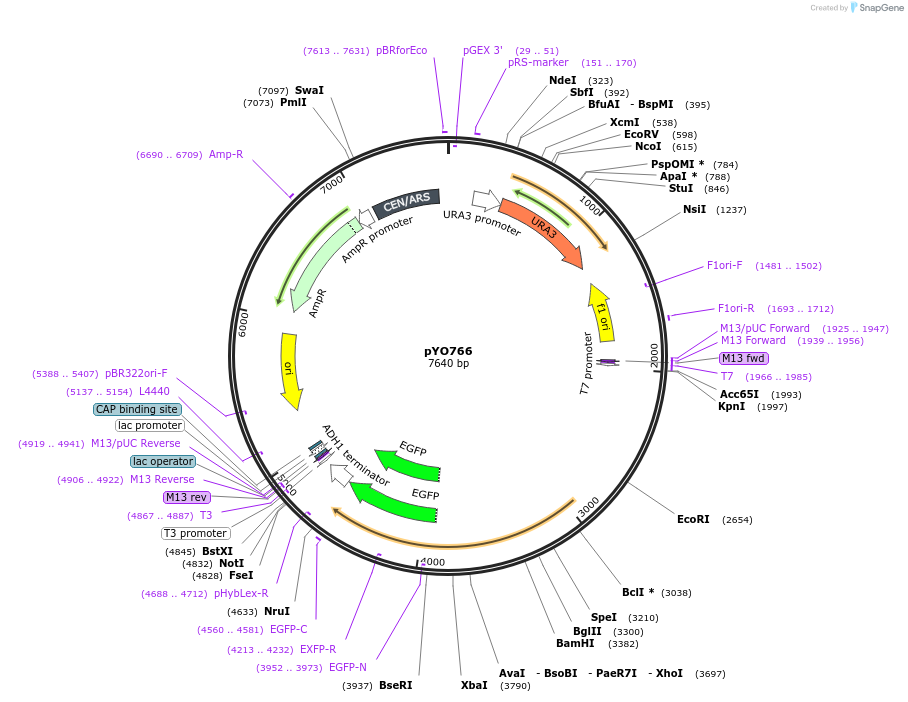
pYO766
Plasmid#235763PurposeExpression of ScVPS38 delta BARA2 (320-440)-EGFP under its own promoterDepositorInsertVSP38 delta BARA2
TagsEGFPExpressionYeastMutationaa 320-440 deletedPromoterScVPS38 promoterAvailable SinceMay 13, 2025AvailabilityAcademic Institutions and Nonprofits only -
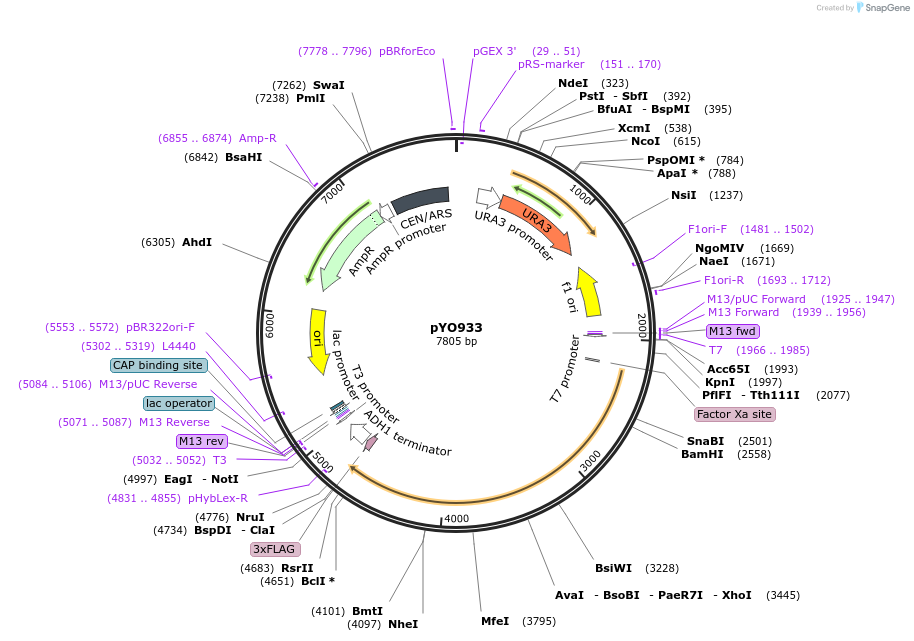
pYO933
Plasmid#235765PurposeExpression of ScVPS34 delta C154-V202-3xFLAG under its own promoterDepositorInsertVPS34 delta C154-V202
Tags3xFLAGExpressionYeastMutationaa 154-202 deletedPromoterScVPS34 promoterAvailable SinceMay 13, 2025AvailabilityAcademic Institutions and Nonprofits only -
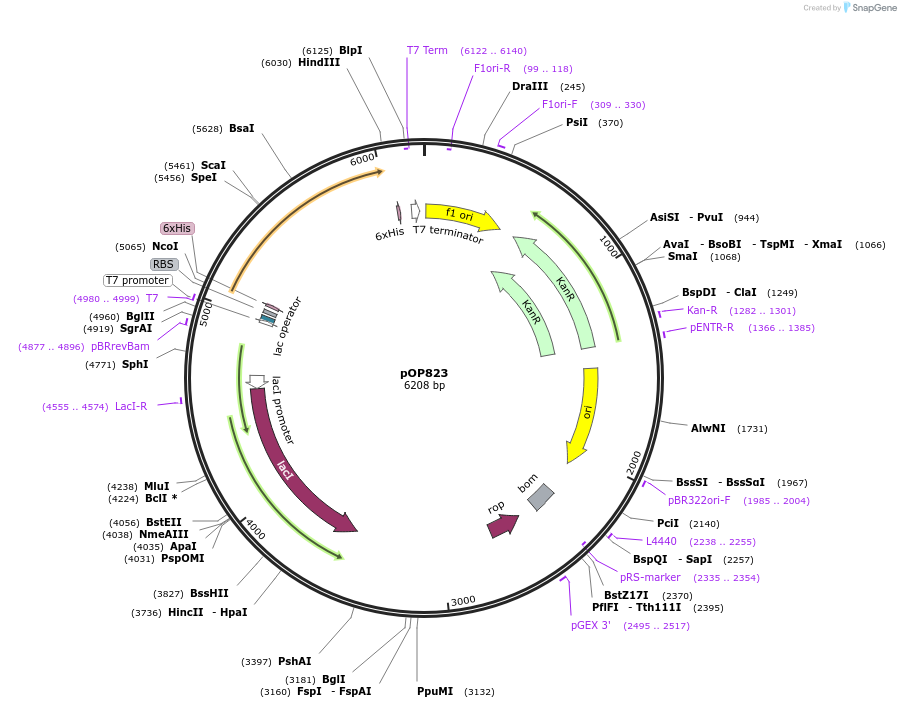
pOP823
Plasmid#235718PurposeProtein expression of His6-SUMO-HsRAB5A (Q79L C19S,C63S) 1-212 in bacteria for maleimide conjugationDepositorAvailable SinceMay 13, 2025AvailabilityAcademic Institutions and Nonprofits only -
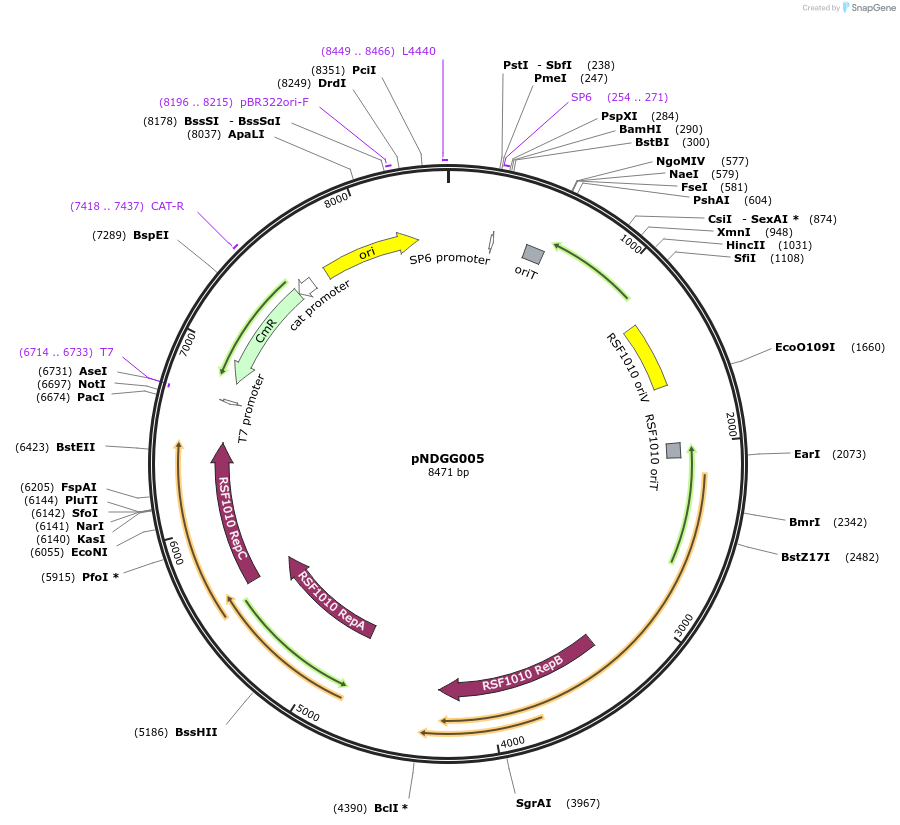
pNDGG005
Plasmid#231309PurposeVector part for BEVA; BEVA2.0 Position 3 RSF1010 broad host range origin of replication with oriT, CmR.DepositorInsertBEVA2.0 Position 3 RSF1010 broad host range origin of replication with oriT, CmR.
ExpressionBacterialAvailable SinceMay 8, 2025AvailabilityAcademic Institutions and Nonprofits only -
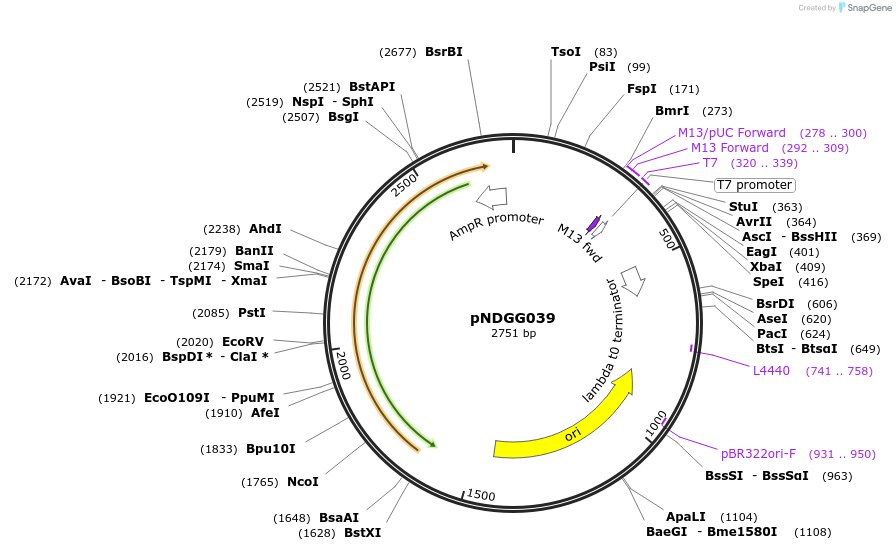
pNDGG039
Plasmid#231339PurposeTerminator; BEVA2.0 T15m terminator with DF extension for CIDAR MoClo Golden Gate Assembly, DF module. Cloned via BpiI GG Assembly of PCR amplicon into POGG006, SpR.DepositorInsertBEVA2.0 T15m terminator with DF extension for CIDAR MoClo Golden Gate Assembly, DF module.
ExpressionBacterialAvailable SinceMay 8, 2025AvailabilityAcademic Institutions and Nonprofits only -
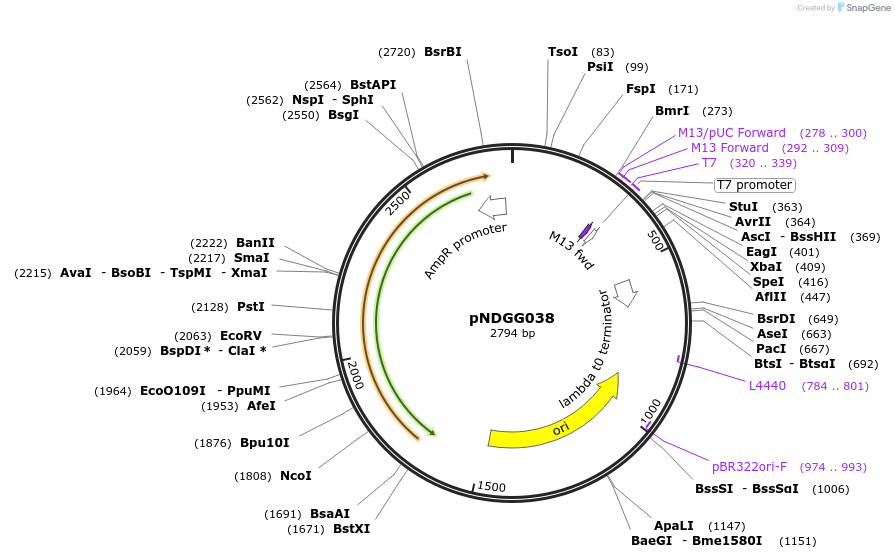
pNDGG038
Plasmid#231338PurposeTerminator; BEVA2.0 T12m terminator with DF extension for CIDAR MoClo Golden Gate Assembly, DF module. Cloned via BpiI GG Assembly of PCR amplicon into POGG006, SpR.DepositorInsertBEVA2.0 T12m terminator with DF extension for CIDAR MoClo Golden Gate Assembly, DF module.
ExpressionBacterialAvailable SinceMay 8, 2025AvailabilityAcademic Institutions and Nonprofits only -
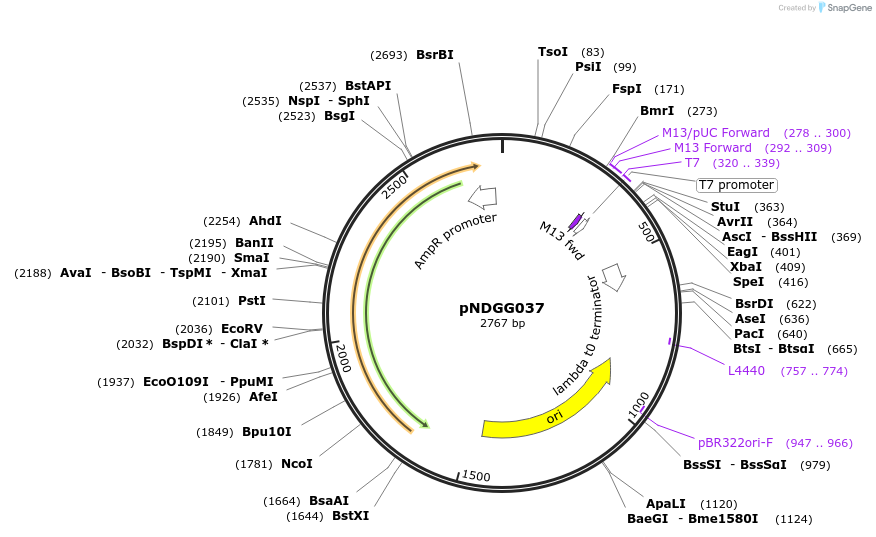
pNDGG037
Plasmid#231337PurposeTerminator; BEVA2.0 T2m terminator with DF extension for CIDAR MoClo Golden Gate Assembly, DF module. Cloned via BpiI GG Assembly of PCR amplicon into POGG006, SpR.DepositorInsertBEVA2.0 T2m terminator with DF extension for CIDAR MoClo Golden Gate Assembly, DF module.
ExpressionBacterialAvailable SinceMay 8, 2025AvailabilityAcademic Institutions and Nonprofits only -
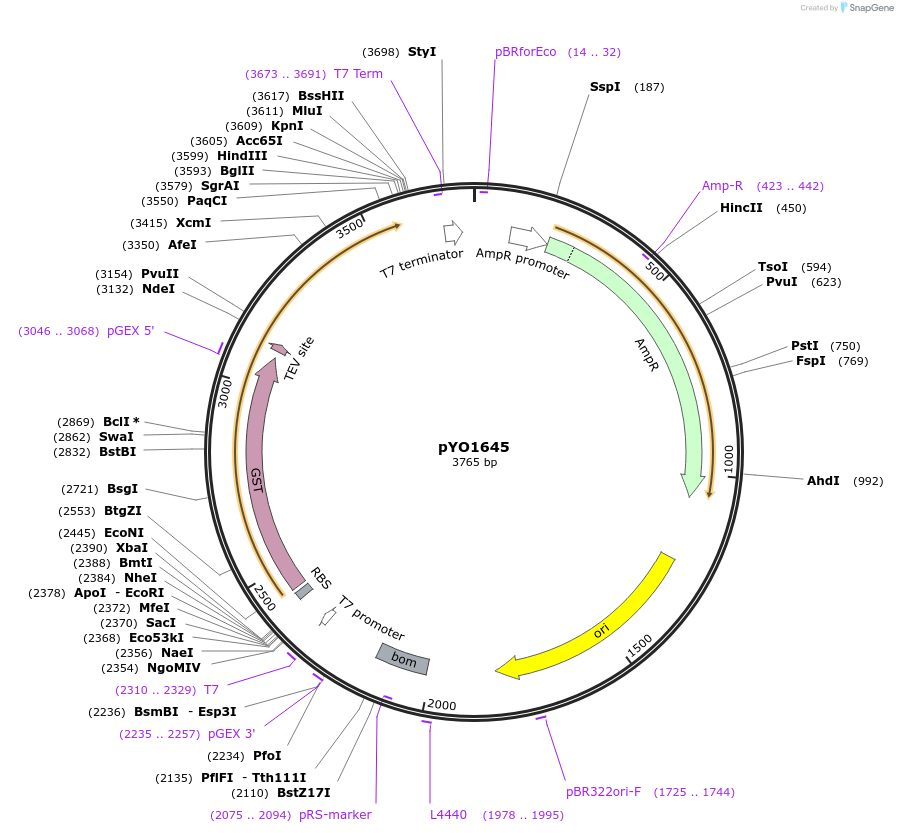
pYO1645
Plasmid#235719PurposeProtein expression of GST-tagged PX domain (Cys at 2 and C85S) in bacteriaDepositorInsertp40phox PX domain (NCF4 Human)
TagsGSTExpressionBacterialMutationCys at 2, C85S, aa 2-149Available SinceApril 16, 2025AvailabilityAcademic Institutions and Nonprofits only -
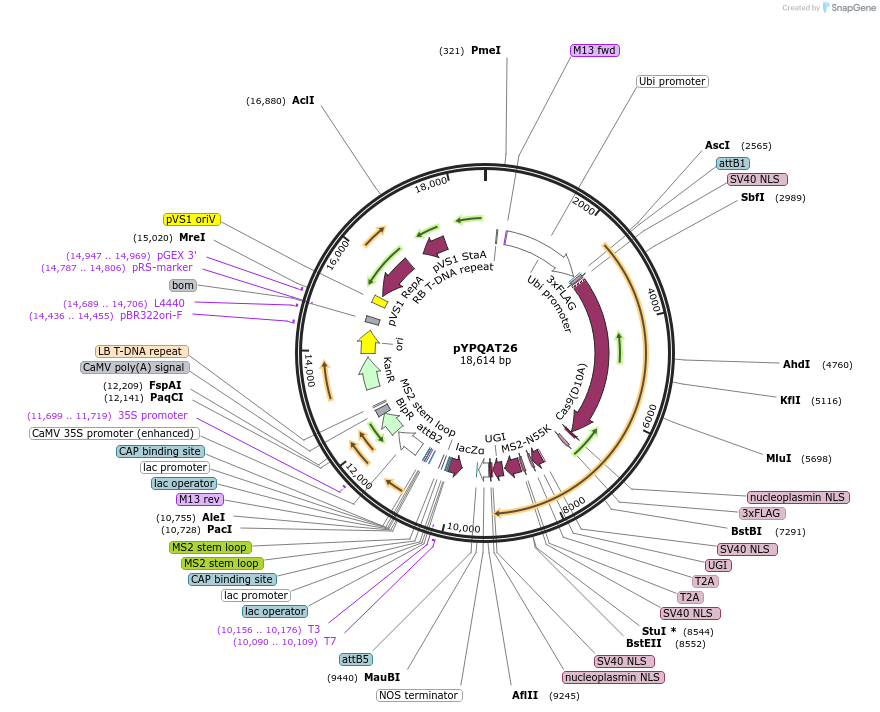
pYPQAT26
Plasmid#223398PurposeT-DNA vector for PmCDA1-CBE_V04 based C-to-T base editing for dicot plants; NGG PAM; 5' editing window; PmCDA1-CBE_V04 was driven by ZmUbi1 and the sgRNA was driven by AtU3; BASTA for plants select.DepositorInsertZmUbi-SpCas9D10A-PmCDA-UGI-T2A-MS2-UGI-AtU3-gRNA scaffold 2.0
UseCRISPRTags3x FLAG and NLSExpressionPlantAvailable SinceApril 10, 2025AvailabilityAcademic Institutions and Nonprofits only -
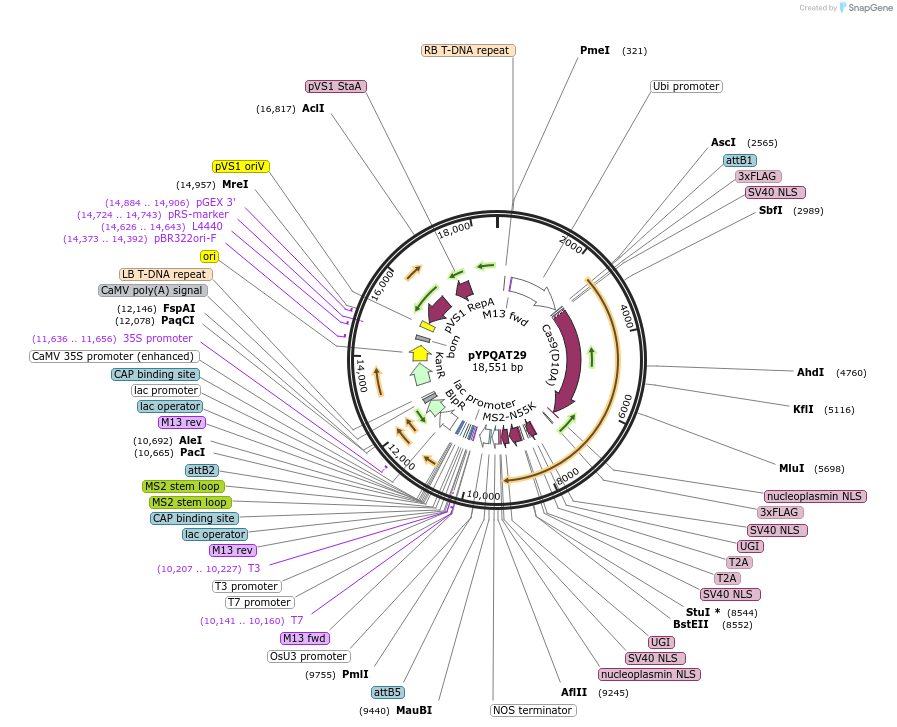
pYPQAT29
Plasmid#223401PurposeT-DNA vector for PmCDA1-CBE_V04 based C-to-T base editing for monocot plants; NGG PAM; 5' editing window; PmCDA1-CBE_V04 was driven by ZmUbi1 and the sgRNA was driven by OsU3; BASTA for plant select.DepositorInsertZmUbi-SpCas9D10A-PmCDA-UGI-T2A-MS2-UGI-OsU3-gRNA scaffold 2.0
UseCRISPRTags3x FLAG and NLSExpressionPlantAvailable SinceApril 10, 2025AvailabilityAcademic Institutions and Nonprofits only -
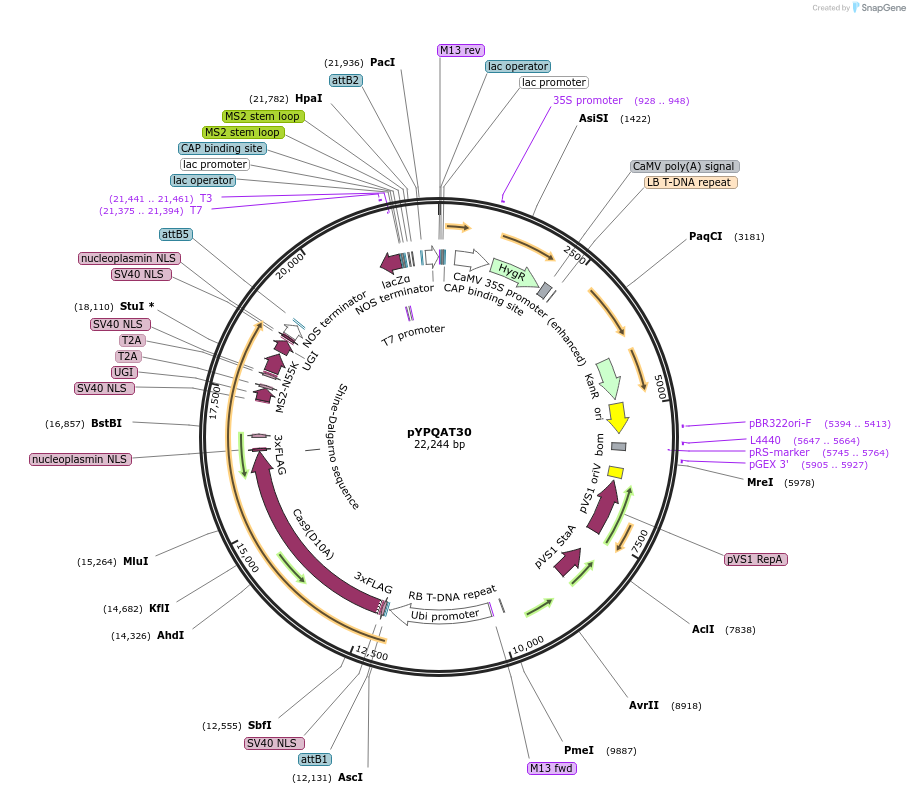
pYPQAT30
Plasmid#223402PurposeT-DNA vector for PmCDA1-CBE_V04 based C-to-T base editing for monocot plants; NGG PAM; 5' editing window; PmCDA1-CBE_V04 and the sgRNA was driven by separate ZmUbi1; Hygromycin for plants selection.DepositorInsertZmUbi-SpCas9D10A-PmCDA-UGI-T2A-MS2-UGI-ZmUbi-gRNA scaffold 2.0-tRNA
UseCRISPRTags3x FLAG and NLSExpressionPlantAvailable SinceApril 10, 2025AvailabilityAcademic Institutions and Nonprofits only -
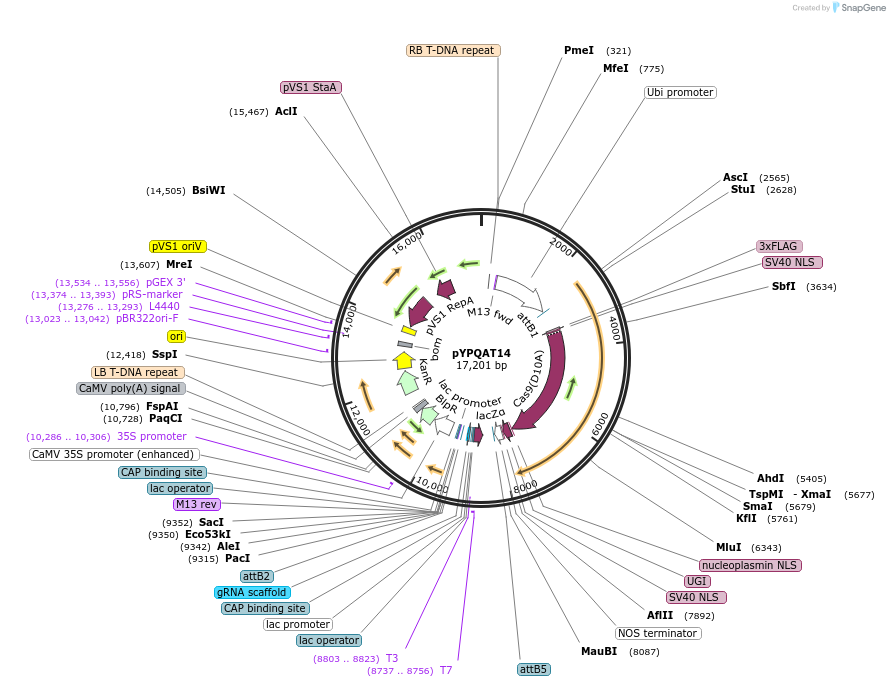
pYPQAT14
Plasmid#223386PurposeT-DNA vector for A3A/Y130F-CBE_V01 based C-to-T base editing for dicot plants; NGG PAM; wide working window; A3A/Y130F-CBE_V01 was driven by ZmUbi1 and sgRNA was by AtU3; BASTA for plant selection.DepositorInsertZmUbi-hA3A-Y130F-SpCas9-D10A-UGI-AtU3-gRNA scaffold
UseCRISPRTags3x FLAG and NLSExpressionPlantAvailable SinceApril 10, 2025AvailabilityAcademic Institutions and Nonprofits only -
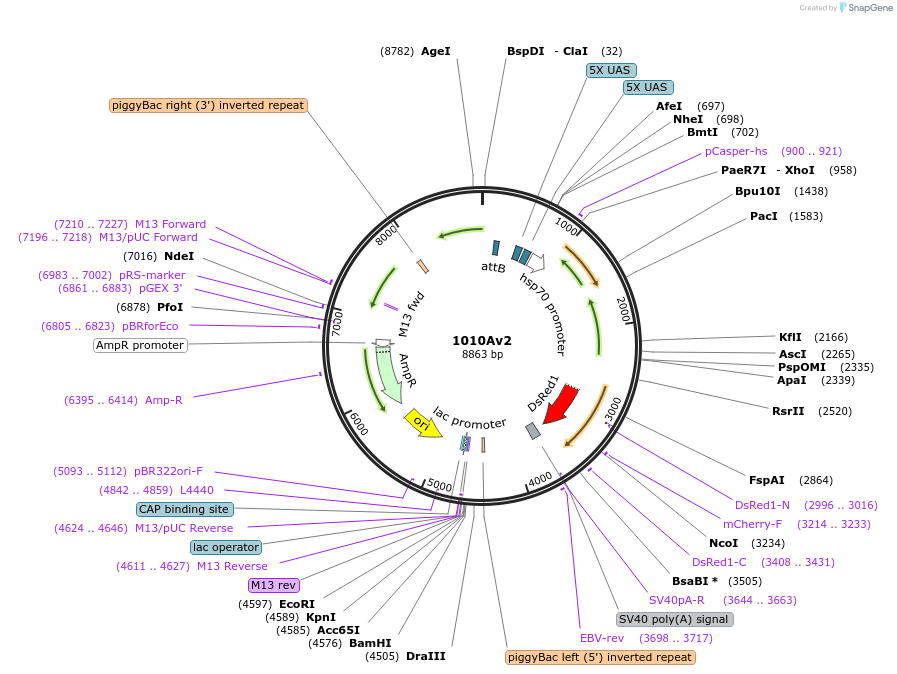
1010Av2
Plasmid#125642PurposeExpresses FAR protein under 10xUAS controlDepositorInsertFAR protein
ExpressionInsectPromoter10xUASAvailable SinceApril 7, 2025AvailabilityAcademic Institutions and Nonprofits only -
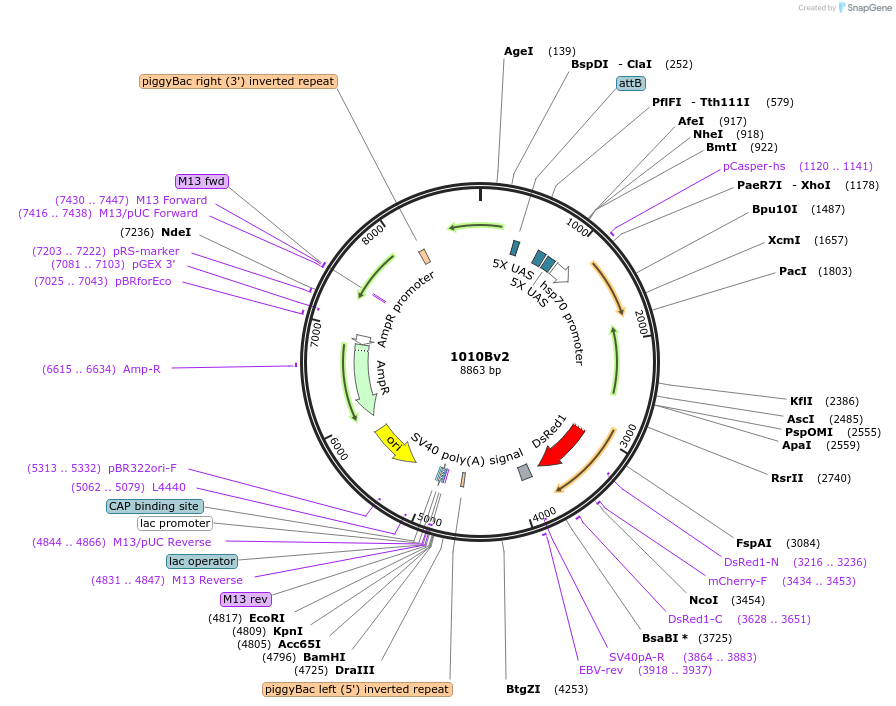
1010Bv2
Plasmid#125643PurposeExpresses FAR protein under 10xUAS controlDepositorInsertFAR protein
ExpressionInsectPromoter10xUASAvailable SinceApril 7, 2025AvailabilityAcademic Institutions and Nonprofits only -
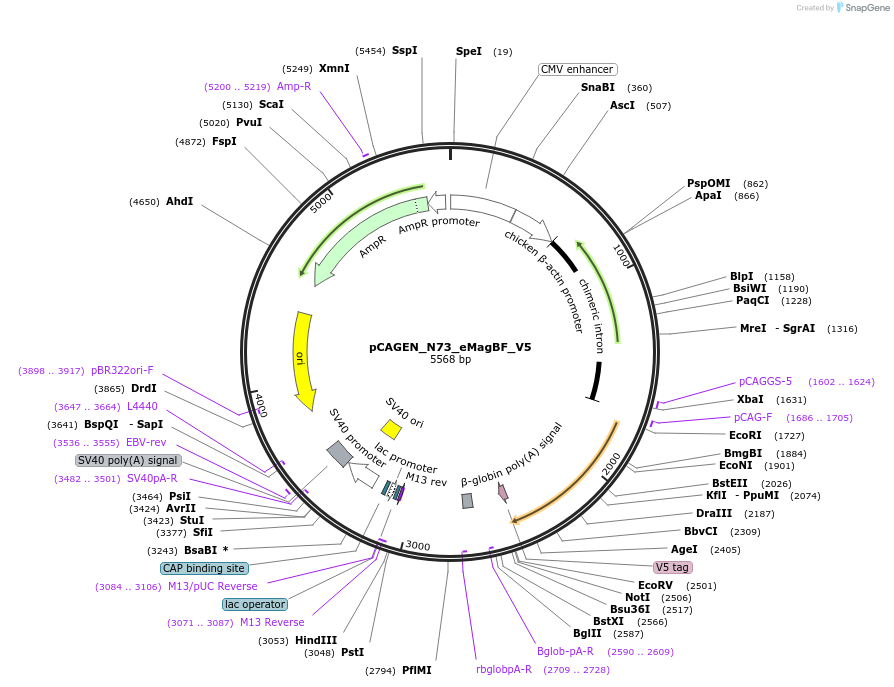
pCAGEN_N73_eMagBF_V5
Plasmid#235642PurposeEncodes ALIBi N-terminal fragment, residues 1-73, fused to eMagBF (enhanced Magnet, basic, fast), V5. No nuclear export sequence to allow nuclear localization.DepositorInsertALIBi N-terminal fragment
TagseMagBF-V5ExpressionMammalianMutationresidues 1-73Available SinceApril 3, 2025AvailabilityAcademic Institutions and Nonprofits only -
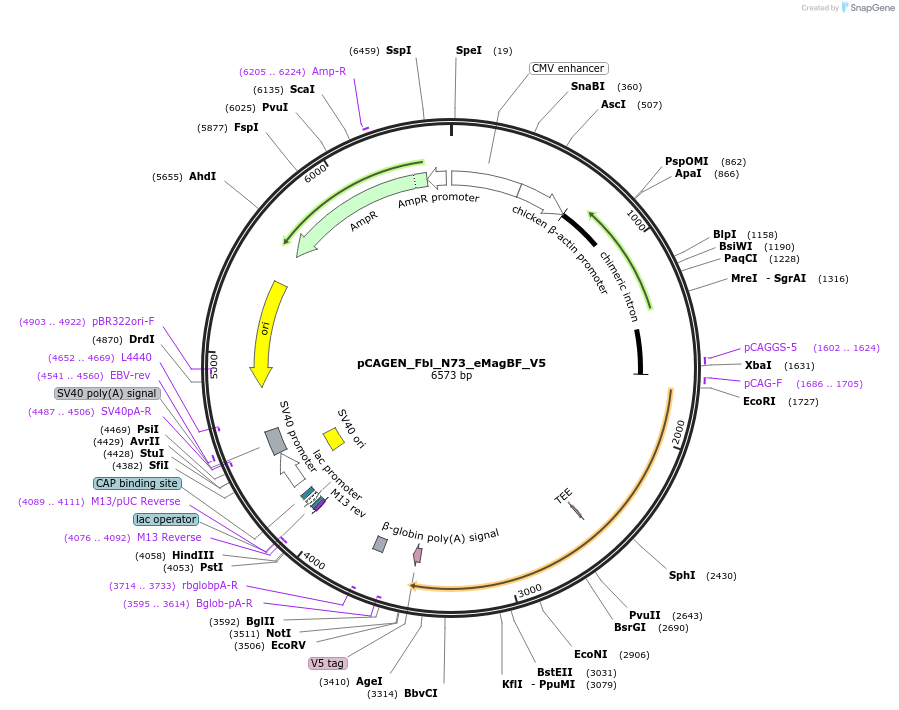
pCAGEN_Fbl_N73_eMagBF_V5
Plasmid#235643PurposeEncodes ALIBi N-terminal fragment, residues 1-73, fused to eMagBF (enhanced Magnet, basic, fast), V5, Fbl for targeting to nucleolus.DepositorInsertALIBi N-terminal fragment
TagsFbl-N73- and eMagBF-V5ExpressionMammalianMutationresidues 1-73Available SinceApril 3, 2025AvailabilityAcademic Institutions and Nonprofits only -
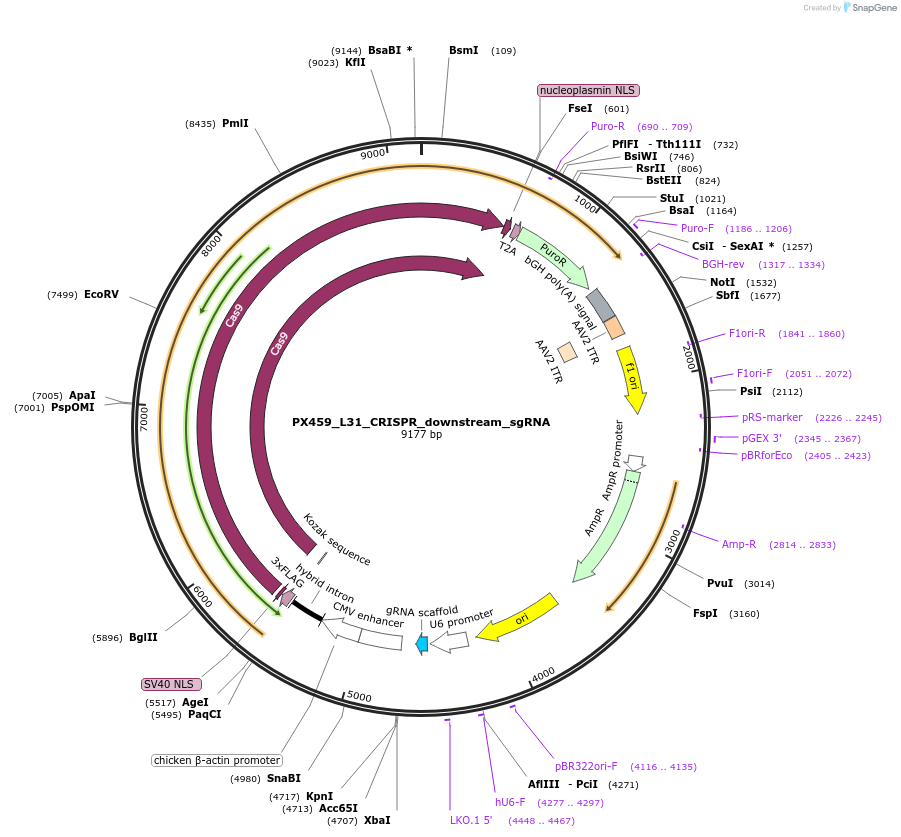
PX459_L31_CRISPR_downstream_sgRNA
Plasmid#235610PurposeEncodes CRISPR gRNA for cleaving downstream of insertion site at C terminus of mouse Rpl31DepositorInsertRpl31 C-terminus downstream sgRNA
UseCRISPRAvailable SinceApril 2, 2025AvailabilityAcademic Institutions and Nonprofits only -
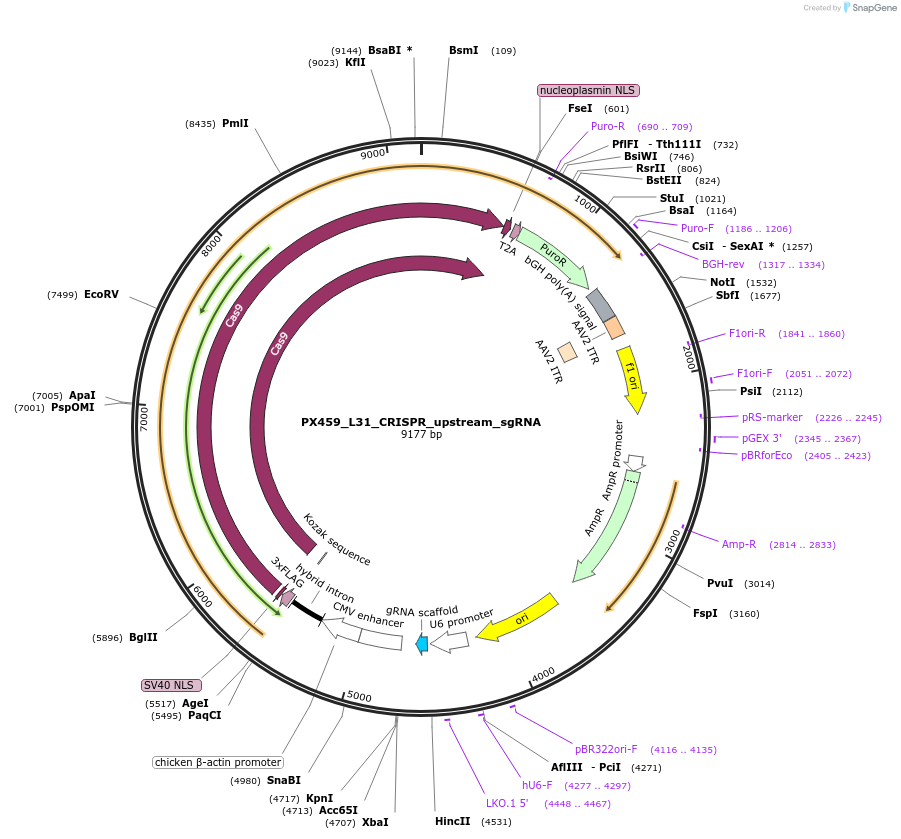
PX459_L31_CRISPR_upstream_sgRNA
Plasmid#235609PurposeEncodes CRISPR gRNA for cleaving upstream of insertion site at C terminus of mouse Rpl31DepositorInsertRpl31 C-terminus upstream sgRNA
UseCRISPRAvailable SinceApril 2, 2025AvailabilityAcademic Institutions and Nonprofits only -
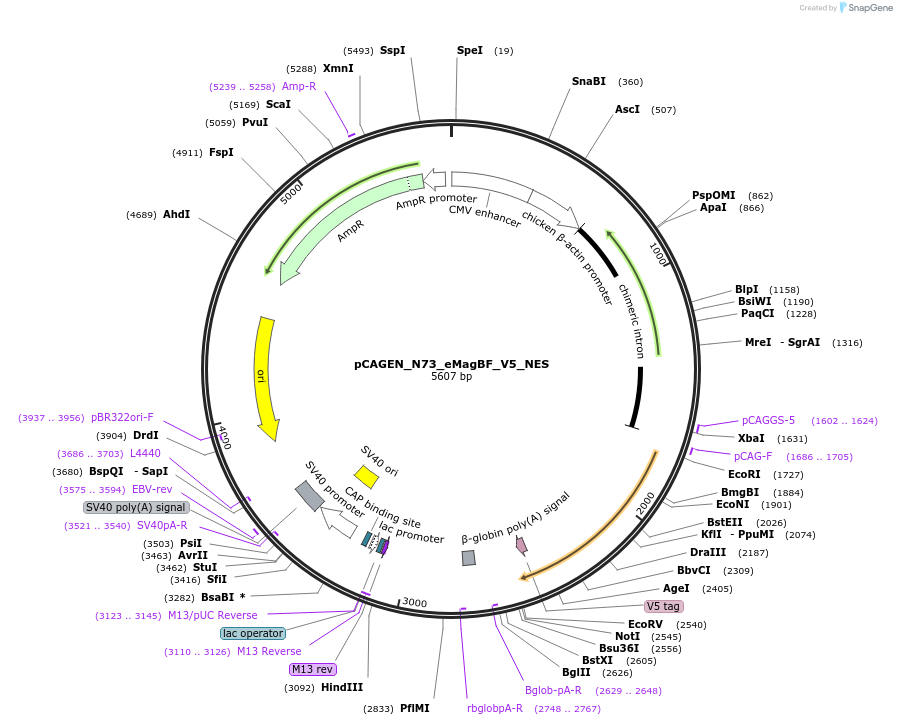
pCAGEN_N73_eMagBF_V5_NES
Plasmid#235614PurposeEncodes ALIBi N-terminal fragment, residues 1-73, fused to eMagBF (enhanced Magnet, basic, fast), V5, and nuclear export sequence.DepositorInsertALIBi N-terminal fragment
TagseMagBF-V5-NESExpressionMammalianMutationresidues 1-73Available SinceApril 2, 2025AvailabilityAcademic Institutions and Nonprofits only -
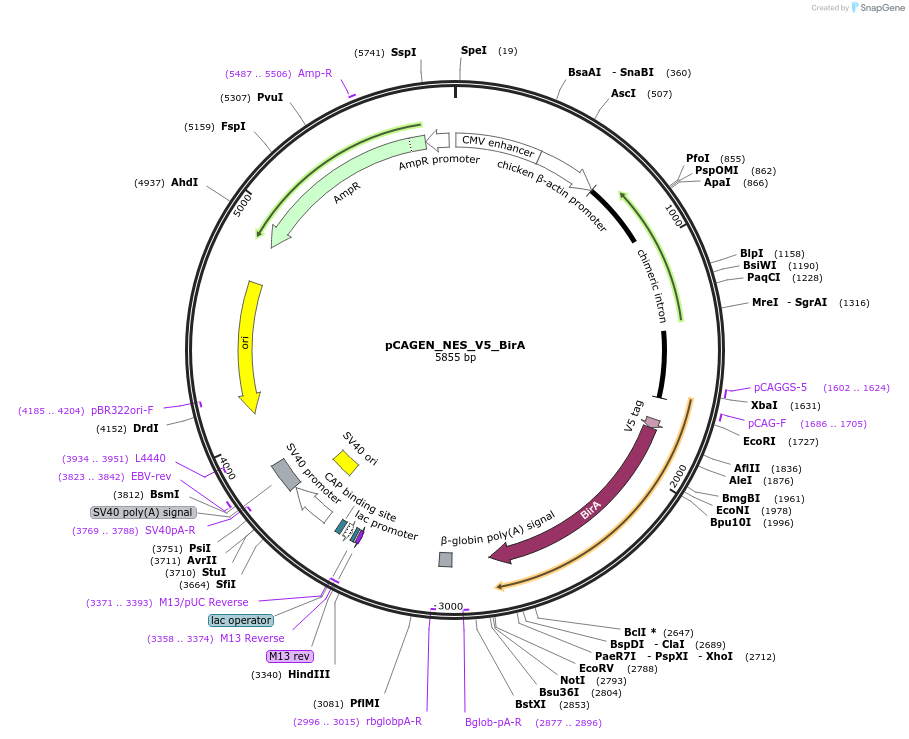
pCAGEN_NES_V5_BirA
Plasmid#235634PurposeEncodes full-length BirA fused to a nuclear export sequence and V5. Used as a positive control in ALIBi experimentsDepositorInsertBirA
TagsNES-V5ExpressionMammalianAvailable SinceApril 2, 2025AvailabilityAcademic Institutions and Nonprofits only -
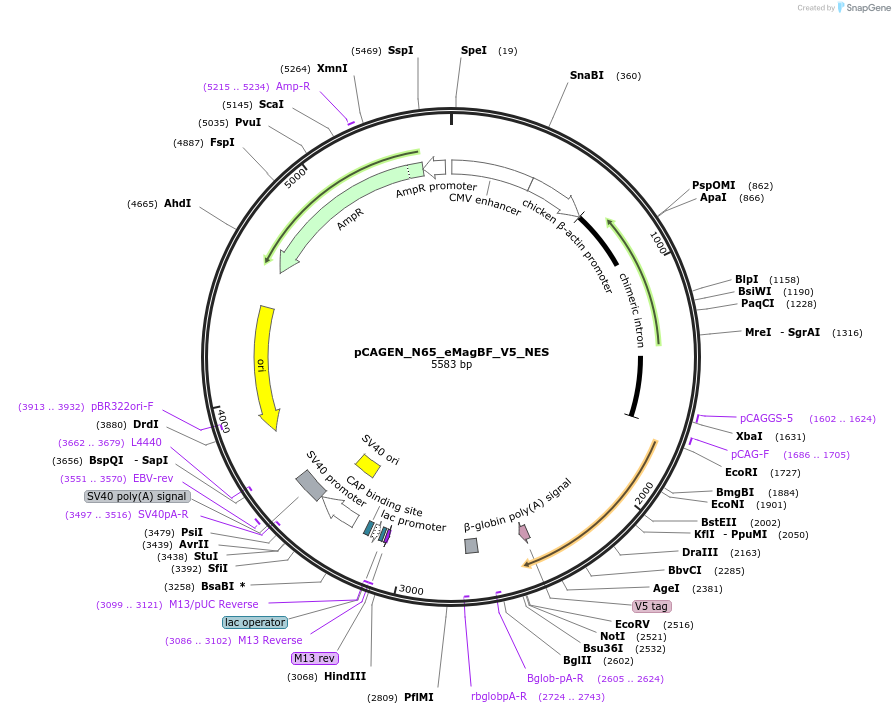
pCAGEN_N65_eMagBF_V5_NES
Plasmid#235612PurposeEncodes ALIBi N-terminal fragment, residues 1-65, fused to eMagBF (enhanced Magnet, basic, fast), V5, and nuclear export sequence.DepositorInsertALIBi N-terminal fragment
TagseMagBF-V5-NESExpressionMammalianMutationresidues 1-65Available SinceApril 2, 2025AvailabilityAcademic Institutions and Nonprofits only -
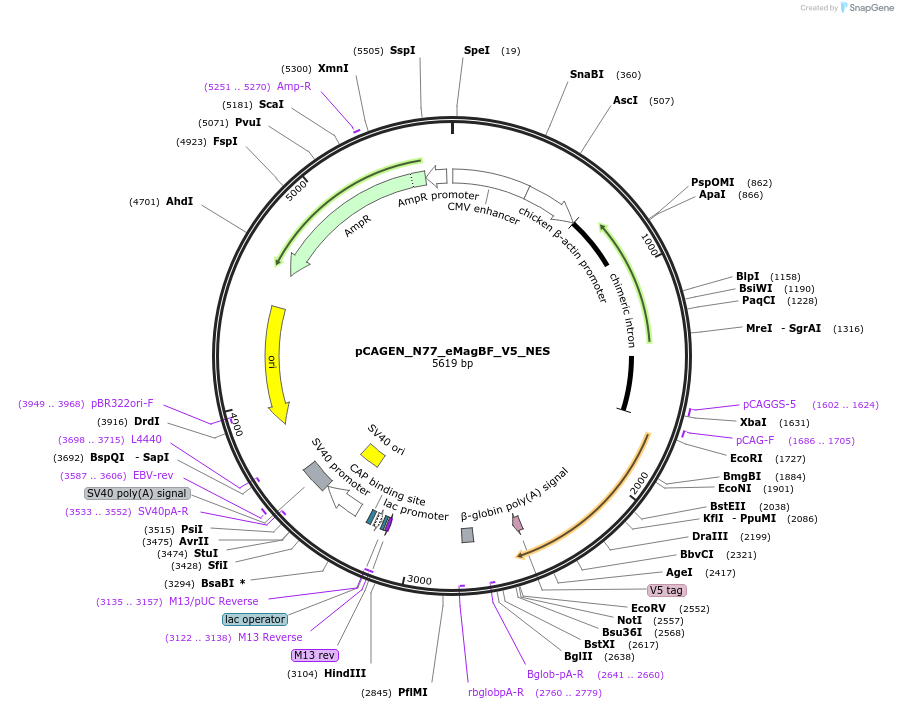
pCAGEN_N77_eMagBF_V5_NES
Plasmid#235616PurposeEncodes ALIBi N-terminal fragment, residues 1-77, fused to eMagBF (enhanced Magnet, basic, fast), V5, and nuclear export sequence.DepositorInsertALIBi N-terminal fragment
TagseMagBF-V5-NESExpressionMammalianMutationresidues 1-77Available SinceApril 2, 2025AvailabilityAcademic Institutions and Nonprofits only -
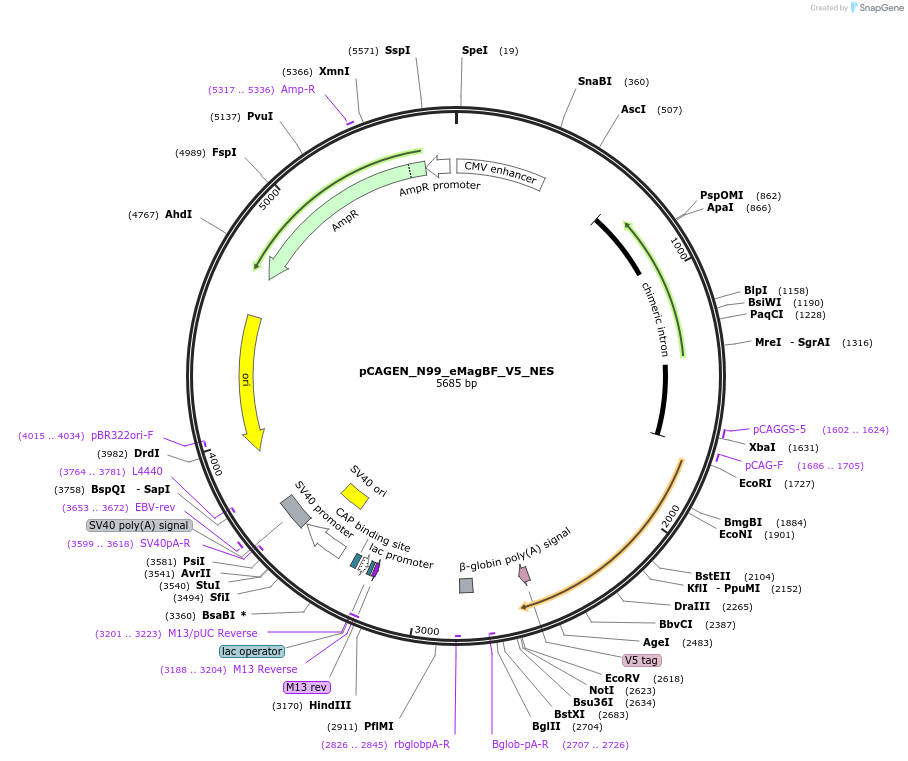
pCAGEN_N99_eMagBF_V5_NES
Plasmid#235618PurposeEncodes ALIBi N-terminal fragment, residues 1-99, fused to eMagBF (enhanced Magnet, basic, fast), V5, and nuclear export sequence.DepositorInsertALIBi N-terminal fragment
TagseMagBF-V5-NESExpressionMammalianMutationresidues 1-99Available SinceApril 2, 2025AvailabilityAcademic Institutions and Nonprofits only -
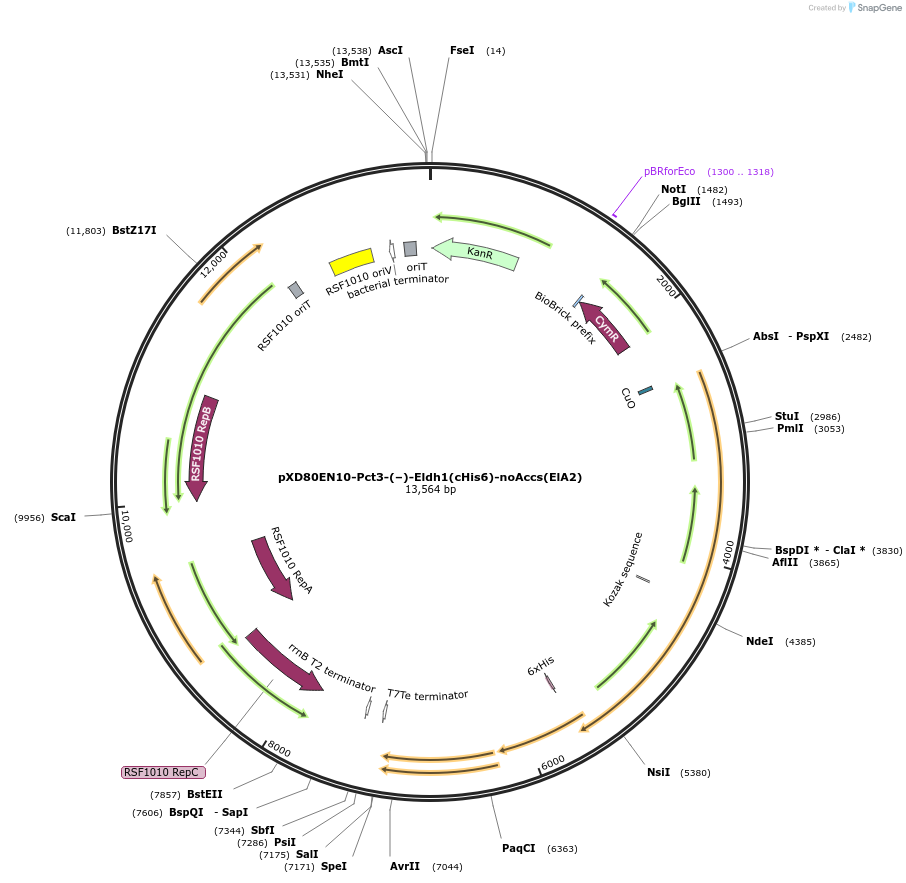
pXD80EN10-Pct3-(–)-Eldh1(cHis6)-noAccs(ElA2)
Plasmid#233880PurposeExpresses His-tagged Eggerthella lenta A2 (–)-Eldh1 with a cumate inducible promoter, without accessory genes (inactive), in Gordonibacter urolithinfaciens DSM 27213DepositorAvailable SinceApril 2, 2025AvailabilityAcademic Institutions and Nonprofits only -
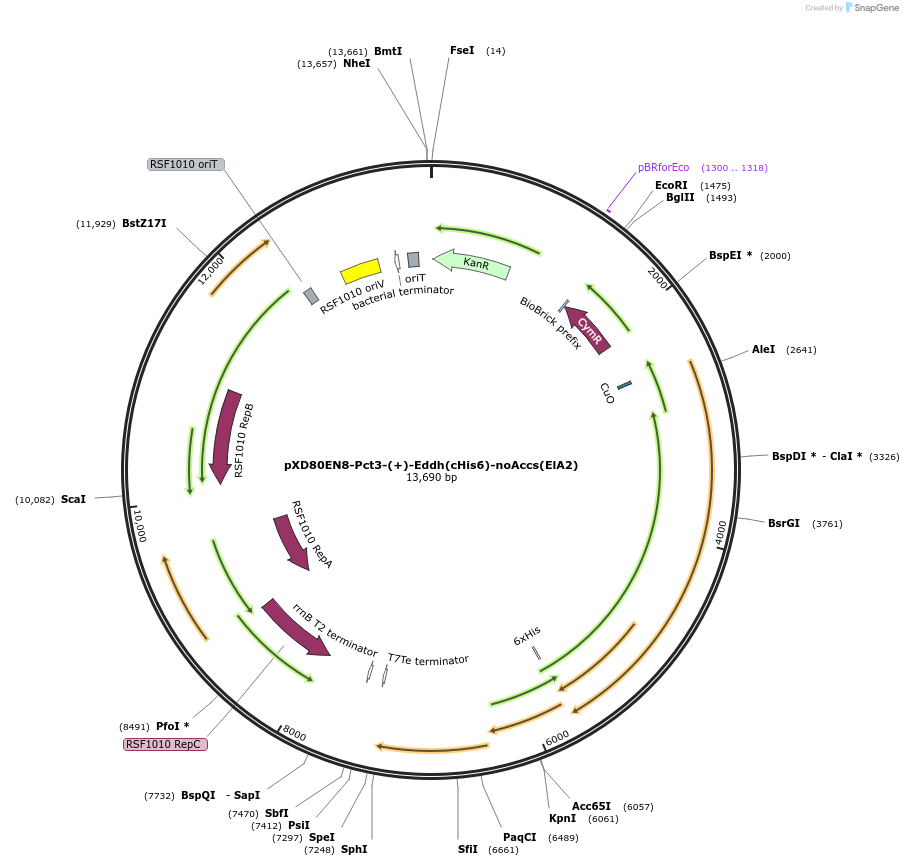
pXD80EN8-Pct3-(+)-Eddh(cHis6)-noAccs(ElA2)
Plasmid#233879PurposeExpresses His-tagged Eggerthella lenta A2 (+)-Eddh with a cumate inducible promoter, without accessory genes (inactive), in Gordonibacter urolithinfaciens DSM 27213DepositorAvailable SinceApril 2, 2025AvailabilityAcademic Institutions and Nonprofits only -
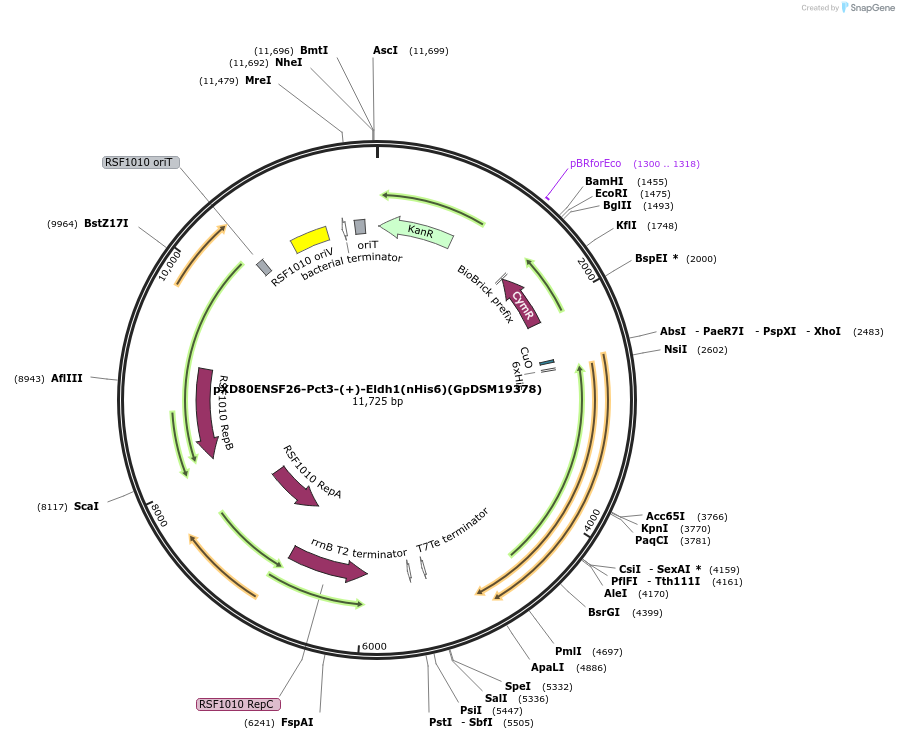
pXD80ENSF26-Pct3-(+)-Eldh1(nHis6)(GpDSM19378)
Plasmid#233874PurposeExpresses His-tagged Gordonibacter pamelaeae DSM 19378 (+)-Eldh1 with a cumate inducible promoter in Gordonibacter urolithinfaciens DSM 27213 for protein purificationDepositorAvailable SinceApril 2, 2025AvailabilityAcademic Institutions and Nonprofits only -
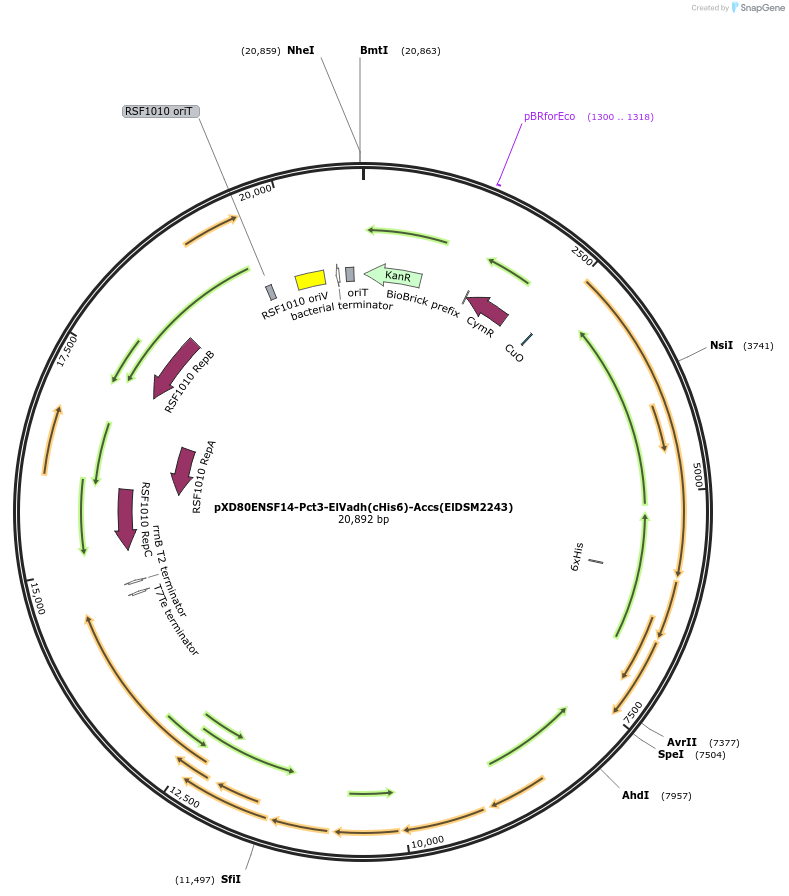
pXD80ENSF14-Pct3-ElVadh(cHis6)-Accs(ElDSM2243)
Plasmid#233871PurposeExpresses His-tagged Eggerthella lenta DSM 2243 El Vadh with a cumate inducible promoter in Gordonibacter urolithinfaciens DSM 27213 for protein purificationDepositorAvailable SinceApril 2, 2025AvailabilityAcademic Institutions and Nonprofits only -
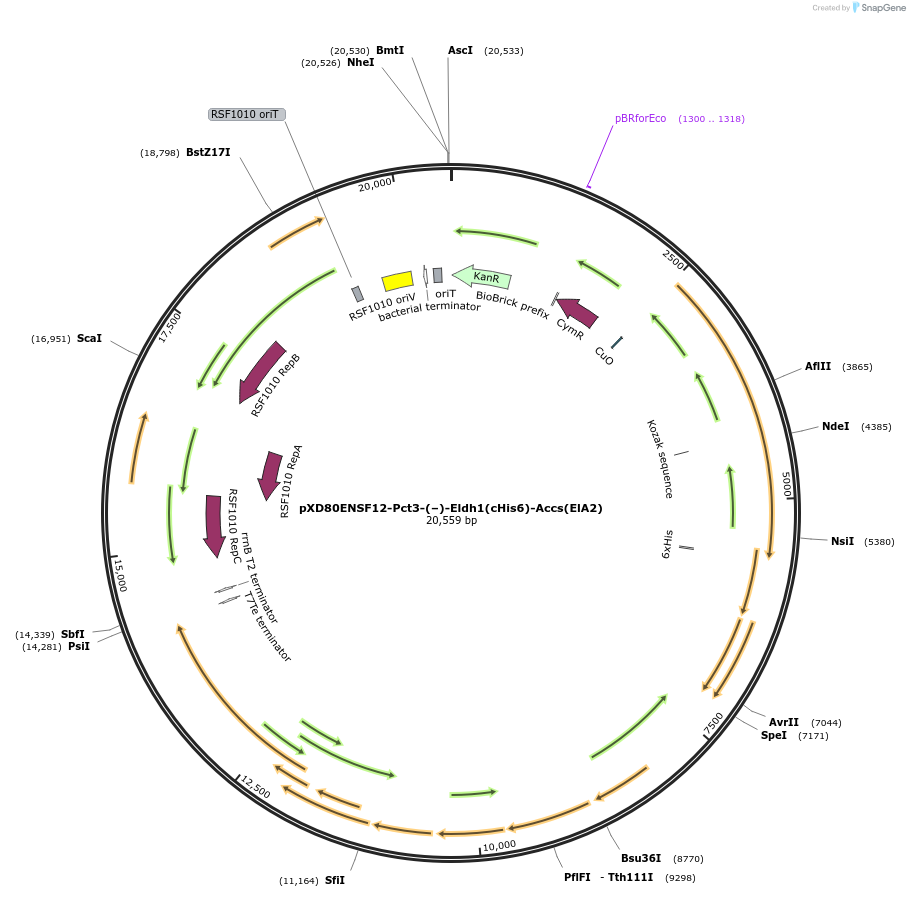
pXD80ENSF12-Pct3-(–)-Eldh1(cHis6)-Accs(ElA2)
Plasmid#233870PurposeExpresses His-tagged Eggerthella lenta A2 (–)-Eldh1 with a cumate inducible promoter in Gordonibacter urolithinfaciens DSM 27213 for protein purificationDepositorAvailable SinceApril 2, 2025AvailabilityAcademic Institutions and Nonprofits only -
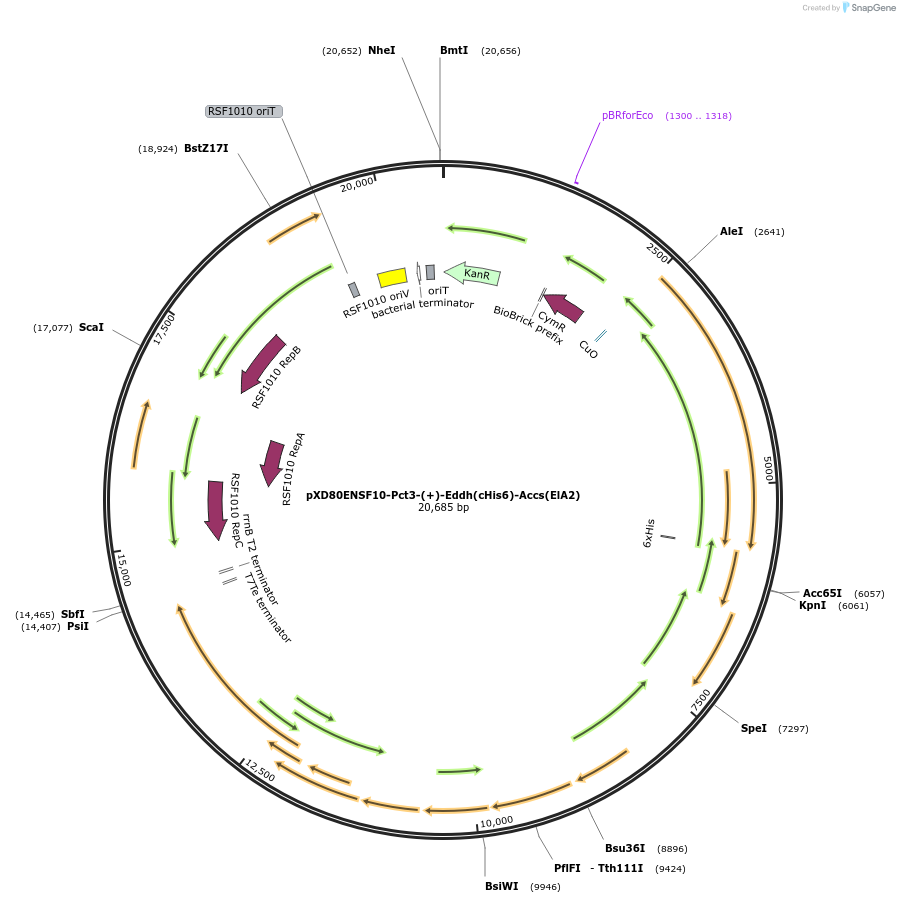
pXD80ENSF10-Pct3-(+)-Eddh(cHis6)-Accs(ElA2)
Plasmid#233869PurposeExpresses His-tagged Eggerthella lenta A2 (+)-Eddh with a cumate inducible promoter in Gordonibacter urolithinfaciens DSM 27213 for protein purificationDepositorAvailable SinceApril 2, 2025AvailabilityAcademic Institutions and Nonprofits only -
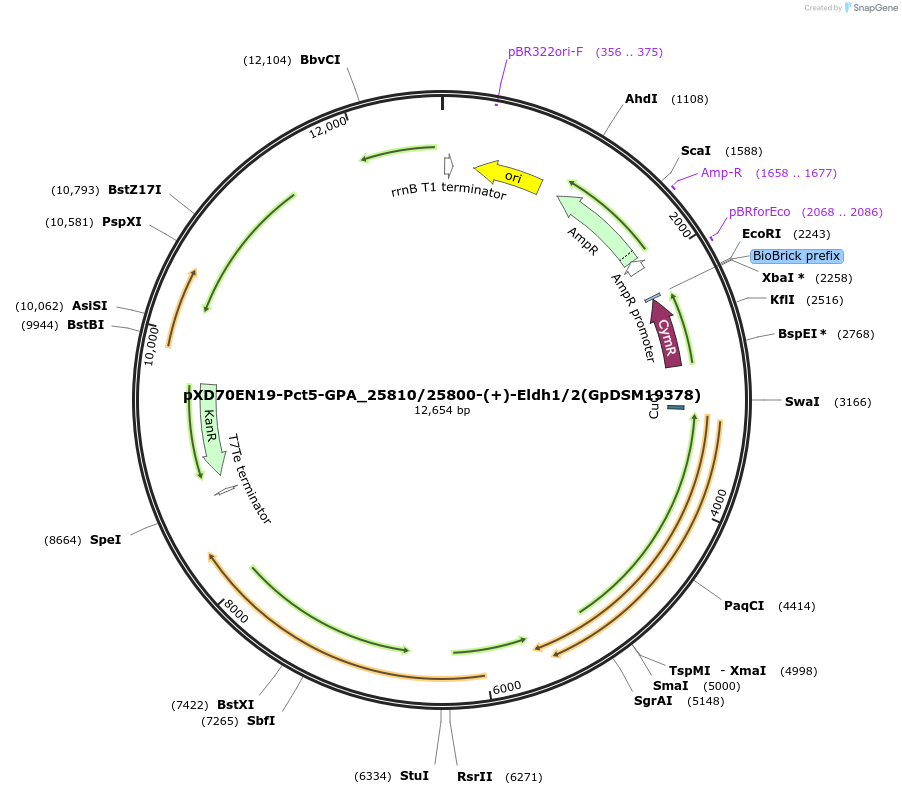
pXD70EN19-Pct5-GPA_25810/25800-(+)-Eldh1/2(GpDSM19378)
Plasmid#225305PurposeExpresses Gordonibacter pamelaeae DSM 19378 (+)-Eldh1 and (+)-Eldh2 in Eggerthella lenta DSM 2243 with a cumate inducible promoterDepositorInsert(+)-Eldh1, (+)-Eldh2
ExpressionBacterialAvailable SinceMarch 20, 2025AvailabilityAcademic Institutions and Nonprofits only -
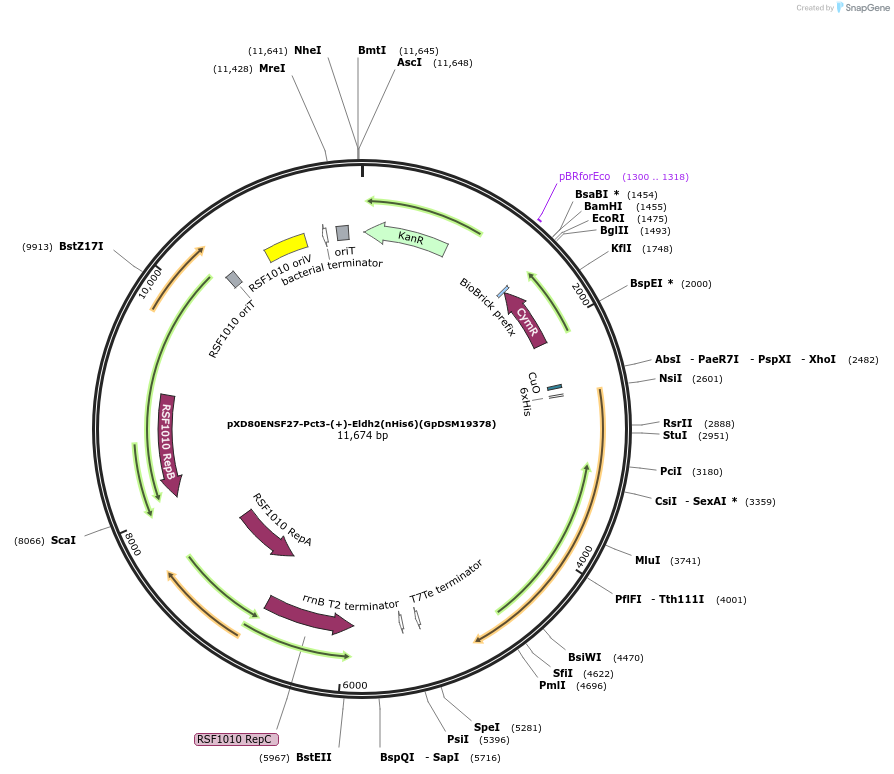
pXD80ENSF27-Pct3-(+)-Eldh2(nHis6)(GpDSM19378)
Plasmid#233875PurposeExpresses His-tagged Gordonibacter pamelaeae DSM 19378 (+)-Eldh2 with a cumate inducible promoter in Gordonibacter urolithinfaciens DSM 27213 for protein purificationDepositorAvailable SinceMarch 17, 2025AvailabilityAcademic Institutions and Nonprofits only -
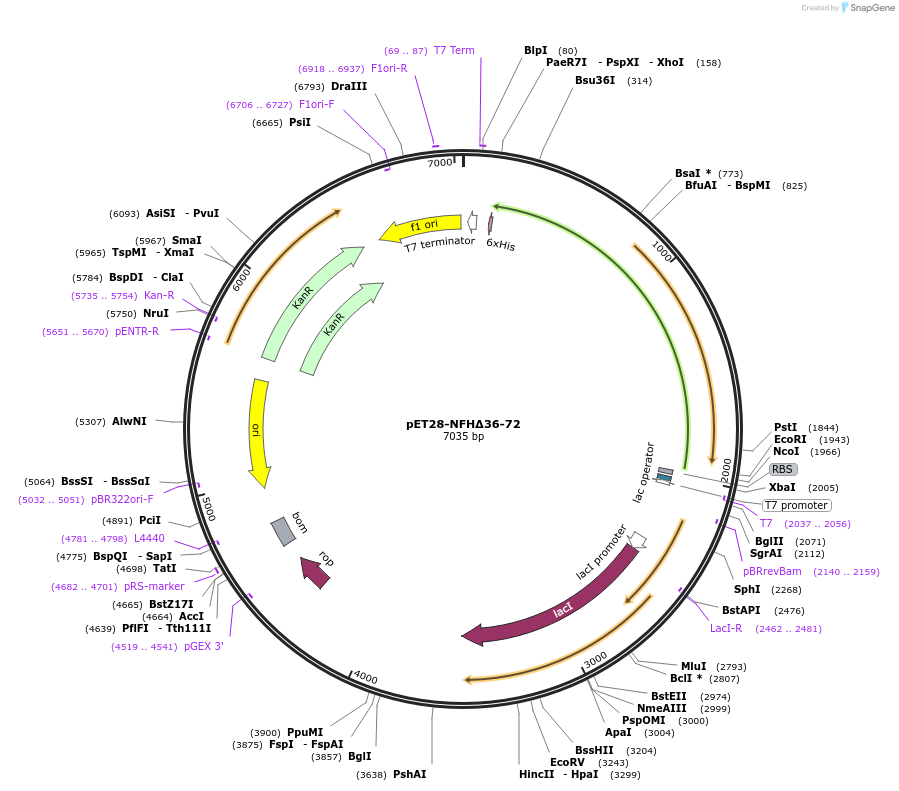
pET28-NFHΔ36-72
Plasmid#232060PurposeExpresses construct for surface tethering: R. norvegicus NFH tail domain lacking residues 36-72DepositorInsertNeurofilament-Heavy tail domain lacking residues 36-72
Tags6xHisExpressionBacterialMutationadded N-terminal tryptophan and tyrosine, and cys…Available SinceMarch 5, 2025AvailabilityAcademic Institutions and Nonprofits only -
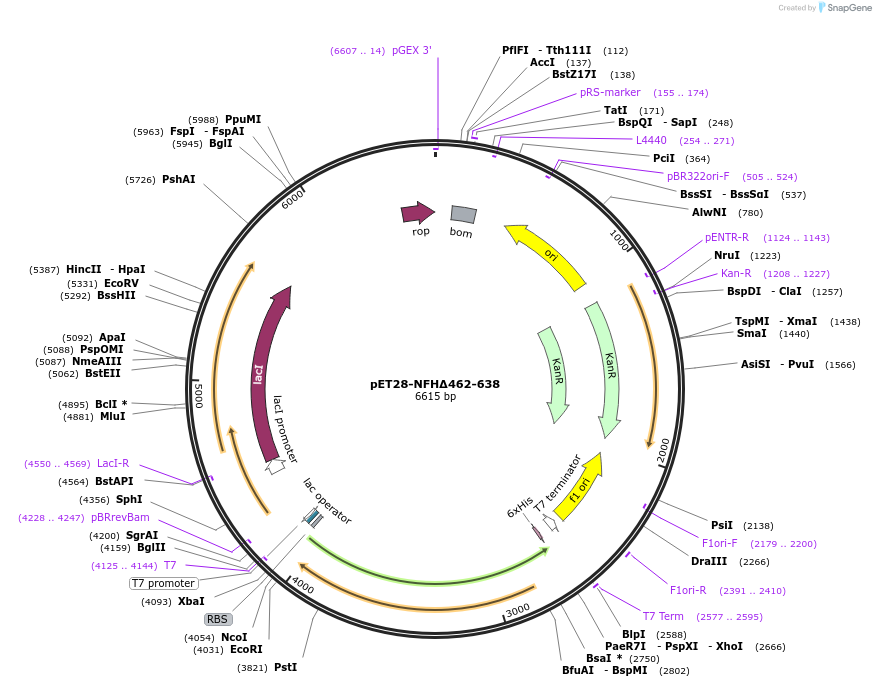
pET28-NFHΔ462-638
Plasmid#232061PurposeExpresses construct for surface tethering: R. norvegicus NFH tail domain lacking residues 462-638DepositorInsertNeurofilament-Heavy tail domain lacking residues 462-638
Tags6xHisExpressionBacterialMutationadded N-terminal tryptophan and tyrosine, and cys…Available SinceMarch 5, 2025AvailabilityAcademic Institutions and Nonprofits only -
pET28-NFLtail
Plasmid#232055PurposeExpresses construct for surface tethering: R. norvegicus NFL tail domainDepositorInsertNeurofilament-Light tail domain
Tags6xHisExpressionBacterialMutationcDNA codon-optimized for E. coli; added N-termina…PromoterT7Available SinceMarch 5, 2025AvailabilityAcademic Institutions and Nonprofits only -
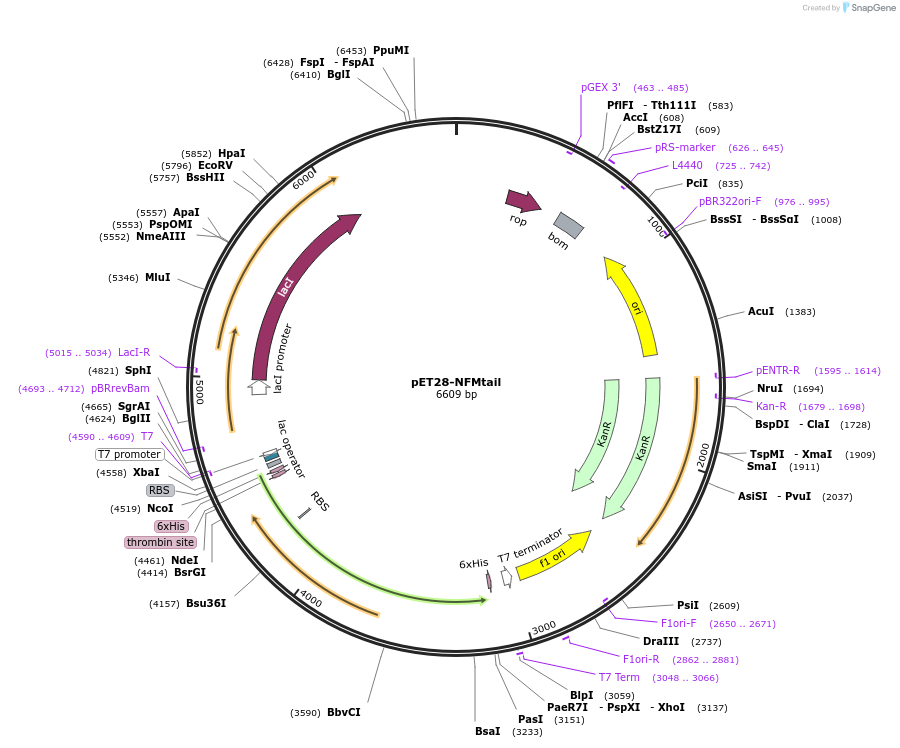
pET28-NFMtail
Plasmid#232056PurposeExpresses construct for surface tethering: R. norvegicus NFM tail domainDepositorInsertNeurofilament-Medium tail domain
Tags6xHis, thrombinExpressionBacterialMutationcDNA codon-optimized for E. coli; added N-termina…PromoterT7Available SinceMarch 5, 2025AvailabilityAcademic Institutions and Nonprofits only -
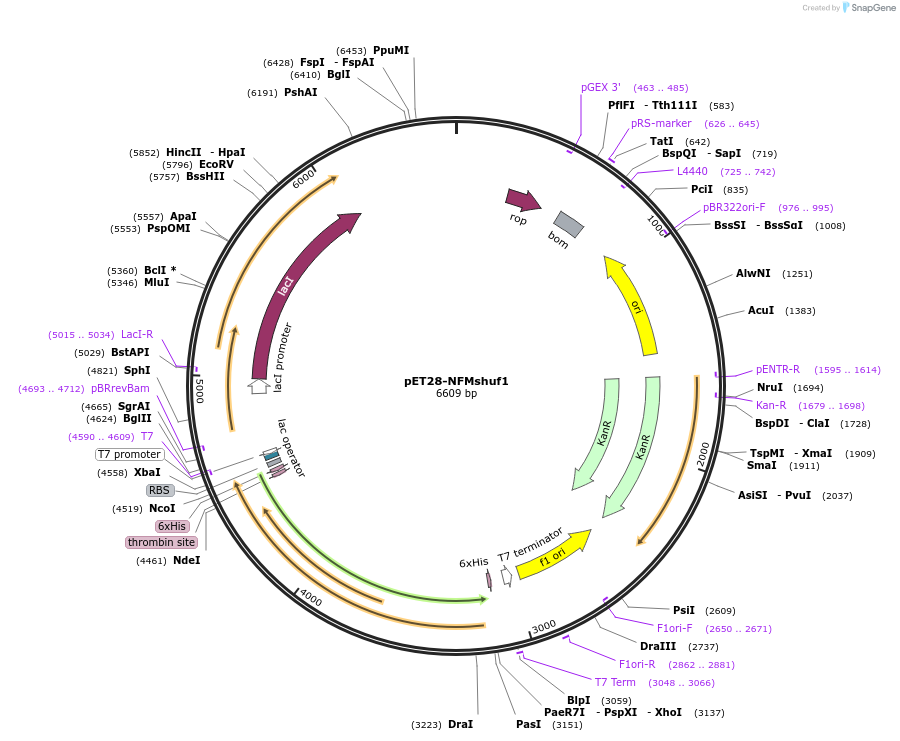
pET28-NFMshuf1
Plasmid#232058PurposeExpresses construct for surface tethering: R. norvegicus NFM tail domain with more evenly distributed charged residues, sequence 1DepositorInsertNeurofilament-Medium tail domain, charge shuffle 1
Tags6xHis, thrombinExpressionBacterialAvailable SinceMarch 5, 2025AvailabilityAcademic Institutions and Nonprofits only -
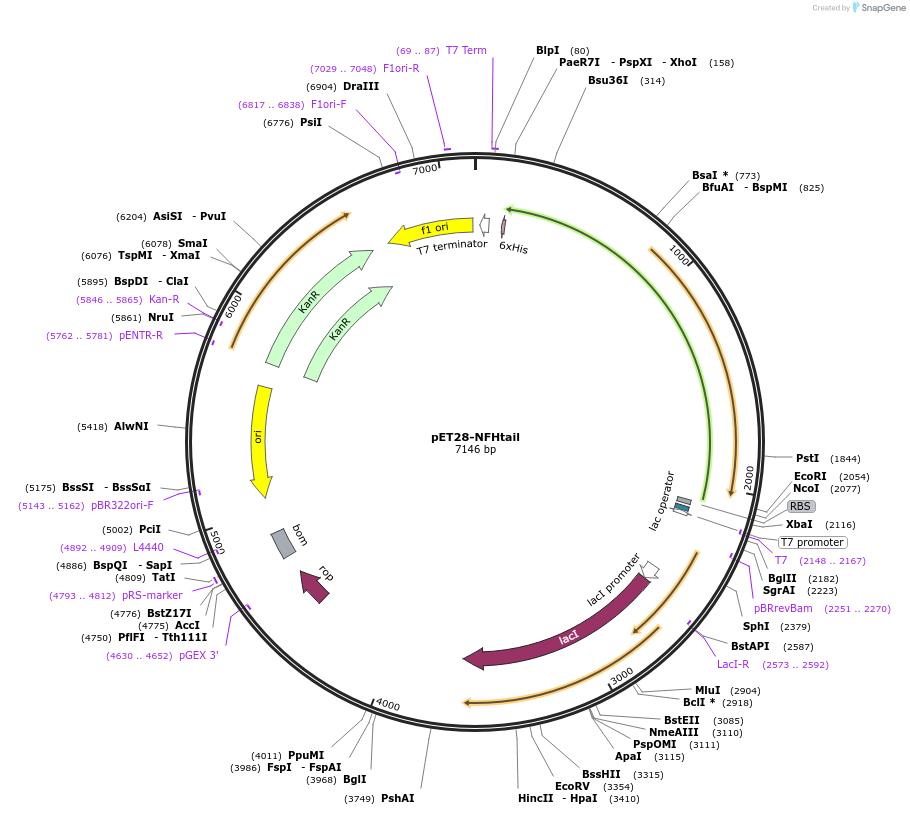
pET28-NFHtail
Plasmid#232057PurposeExpresses construct for surface tethering: R. norvegicus NFH tail domainDepositorInsertNeurofilament-Heavy tail domain
Tags6xHisExpressionBacterialMutationadded N-terminal tryptophan and tyrosine, and cys…Available SinceMarch 5, 2025AvailabilityAcademic Institutions and Nonprofits only -
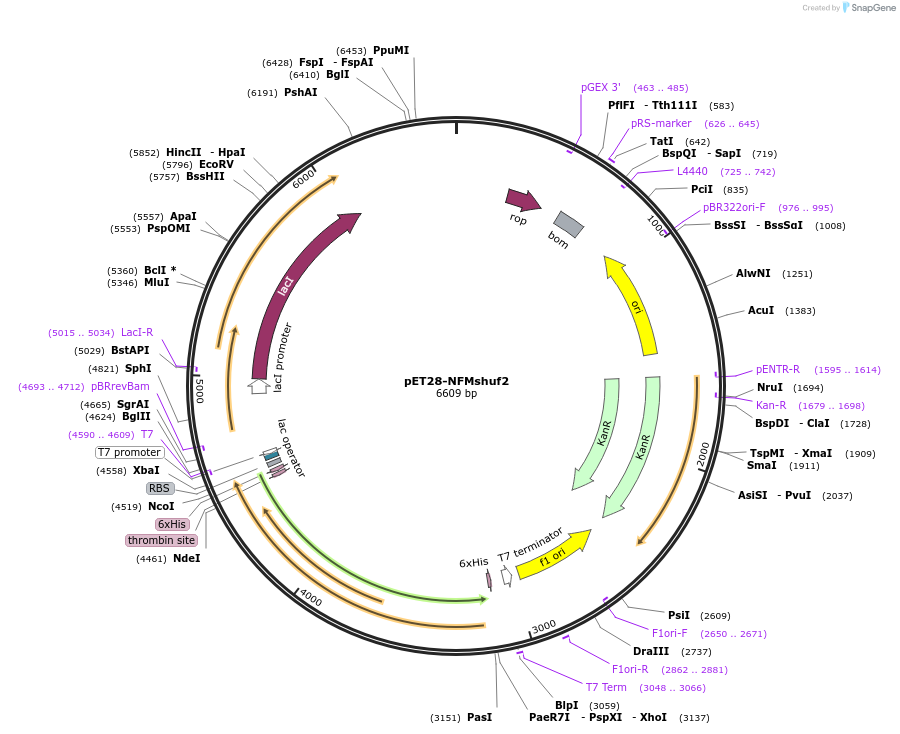
pET28-NFMshuf2
Plasmid#232059PurposeExpresses construct for surface tethering: R. norvegicus NFM tail domain with more evenly distributed charged residues, sequence 2DepositorInsertNeurofilament-Medium tail domain, charge shuffle 2
Tags6xHis, thrombinExpressionBacterialAvailable SinceMarch 5, 2025AvailabilityAcademic Institutions and Nonprofits only -
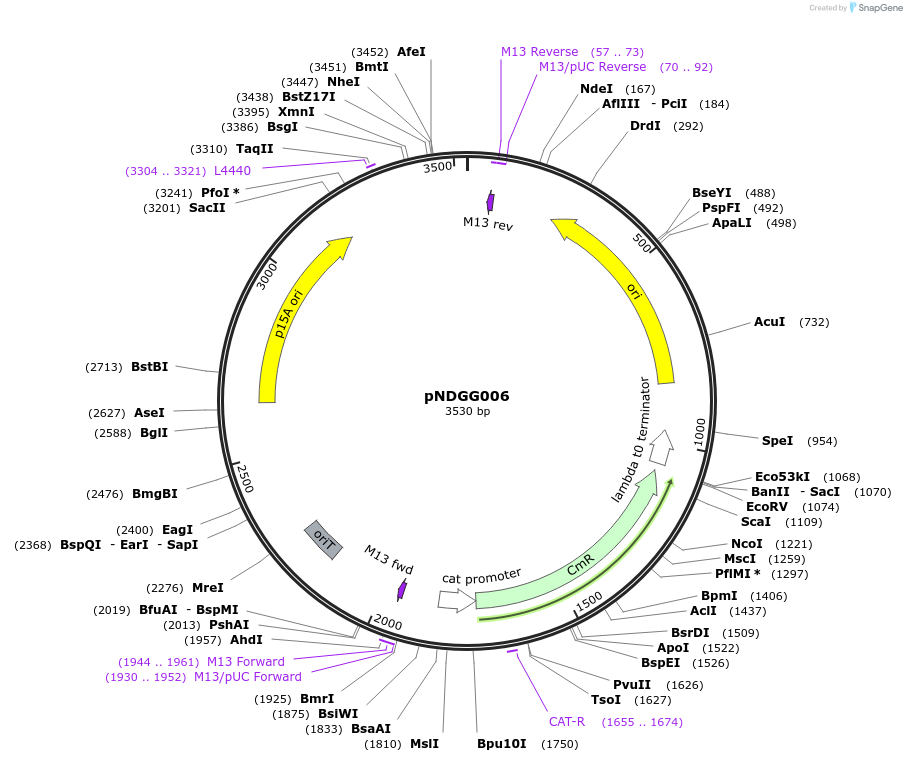
pNDGG006
Plasmid#231310PurposeVector part for BEVA; BEVA2.0 Position 3 p15A narrow host range origin of replication with oriT, CmR.DepositorInsertBEVA2.0 Position 3 p15A narrow host range origin of replication with oriT, CmR.
ExpressionBacterialAvailable SinceMarch 4, 2025AvailabilityAcademic Institutions and Nonprofits only -
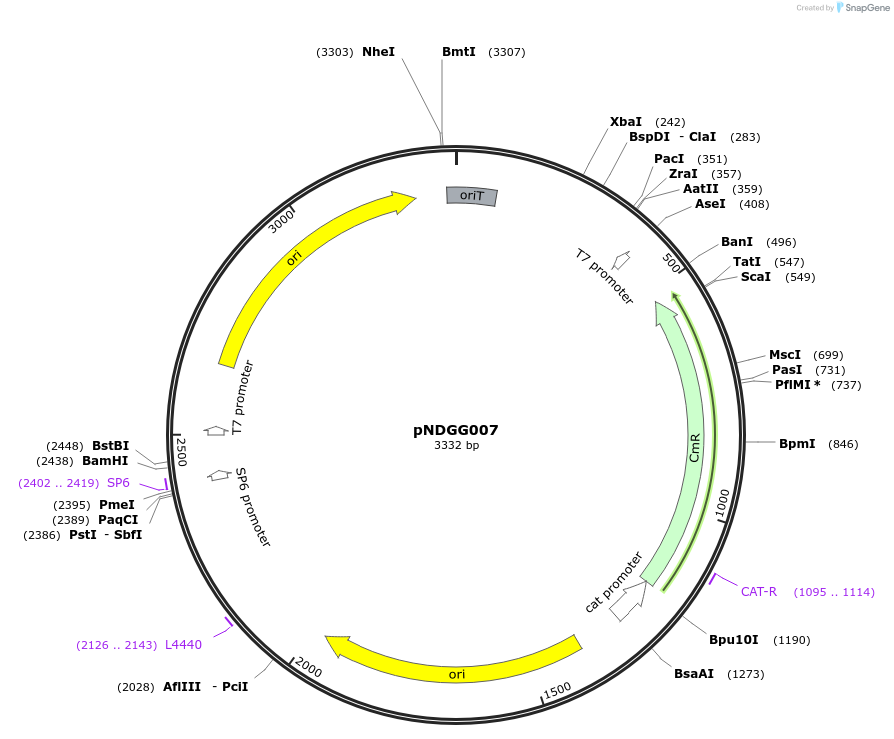
pNDGG007
Plasmid#231311PurposeVector part for BEVA; BEVA2.0 Position 3 pMB1 narrow broad host range origin of replication with oriT, CmR.DepositorInsertBEVA2.0 Position 3 pMB1 narrow broad host range origin of replication with oriT, CmR.
ExpressionBacterialAvailable SinceMarch 4, 2025AvailabilityAcademic Institutions and Nonprofits only -
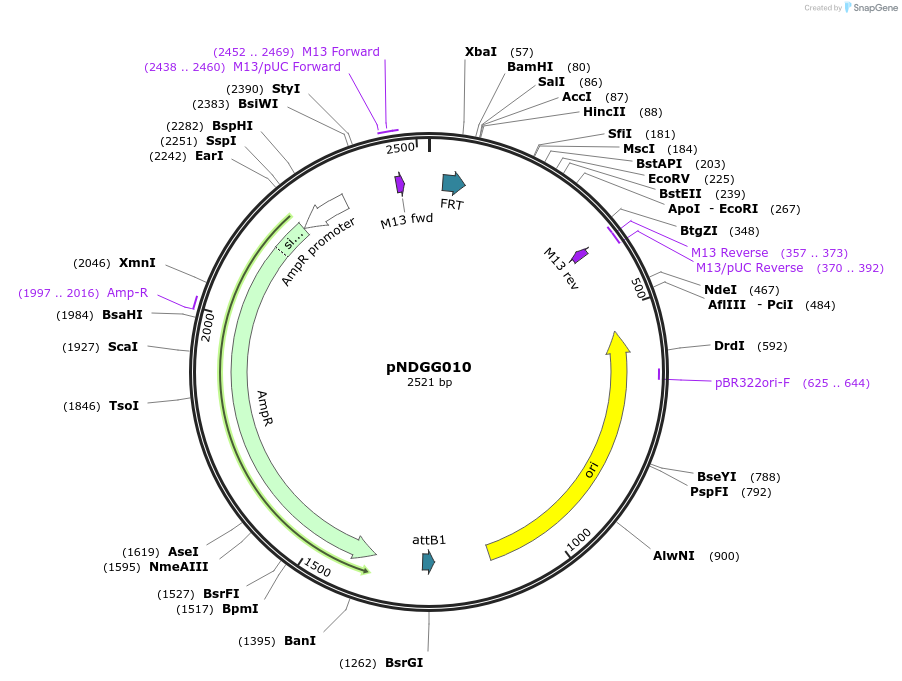
pNDGG010
Plasmid#231315PurposeVector part for BEVA; BEVA2.0 Position 4 attB recombinase site, AmpR. Also includes FRT EF module for L1 GG cloning.DepositorInsertBEVA2.0 Position 4 attB recombinase site, AmpR. Also includes FRT EF module for L1 GG cloning.
ExpressionBacterialAvailable SinceFeb. 27, 2025AvailabilityAcademic Institutions and Nonprofits only -
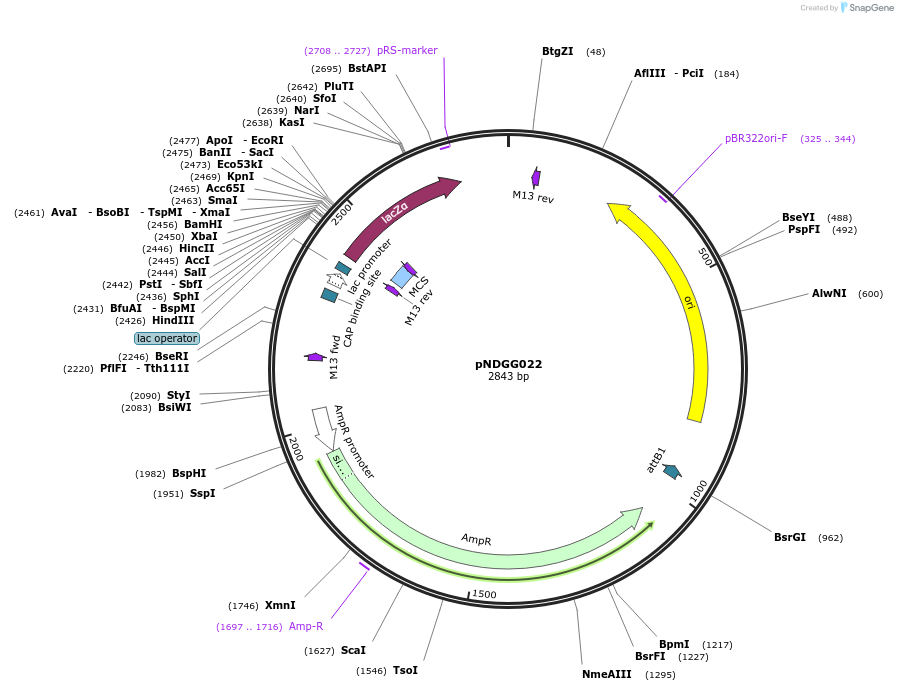
pNDGG022
Plasmid#231306PurposeVector part for BEVA; BEVA2.0 Position 1 Level 1 BsaI cloning site with alternate CAGA 3' fusion site, without T0, AmpRDepositorInsertBEVA2.0 Position 1 Level 1 BsaI cloning site with alternate CAGA 3' fusion site, without T0, AmpR
ExpressionBacterialAvailable SinceFeb. 27, 2025AvailabilityAcademic Institutions and Nonprofits only -
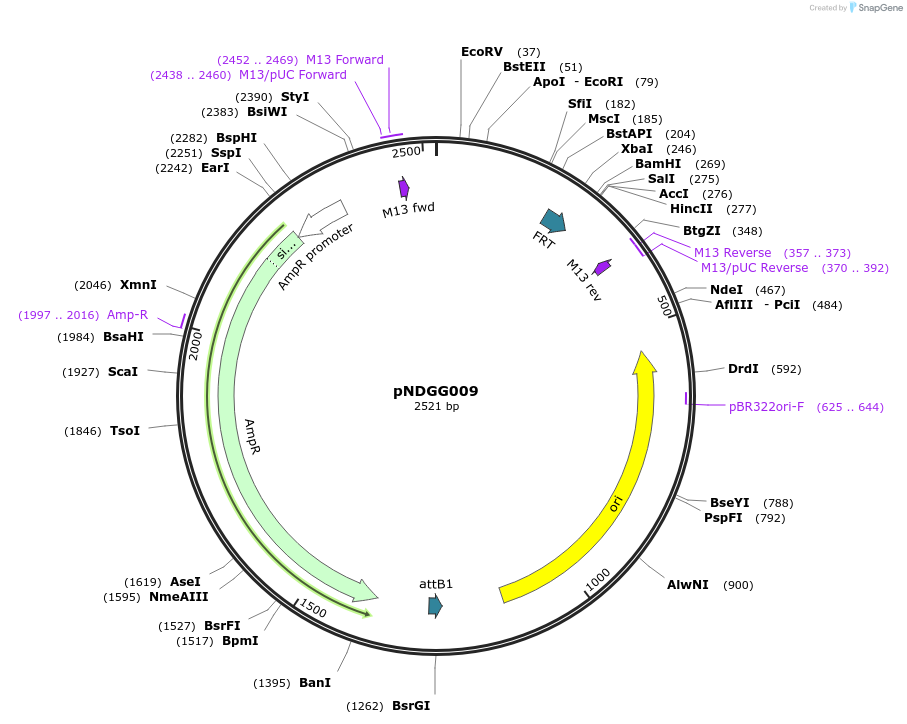
pNDGG009
Plasmid#231314PurposeVector part for BEVA; BEVA2.0 Position 4 FRT recombinase site, AmpR. Also includes attB EF module for L1 GG cloning.DepositorInsertBEVA2.0 Position 4 FRT recombinase site, AmpR. Also includes attB EF module for L1 GG cloning.
ExpressionBacterialAvailable SinceFeb. 27, 2025AvailabilityAcademic Institutions and Nonprofits only -
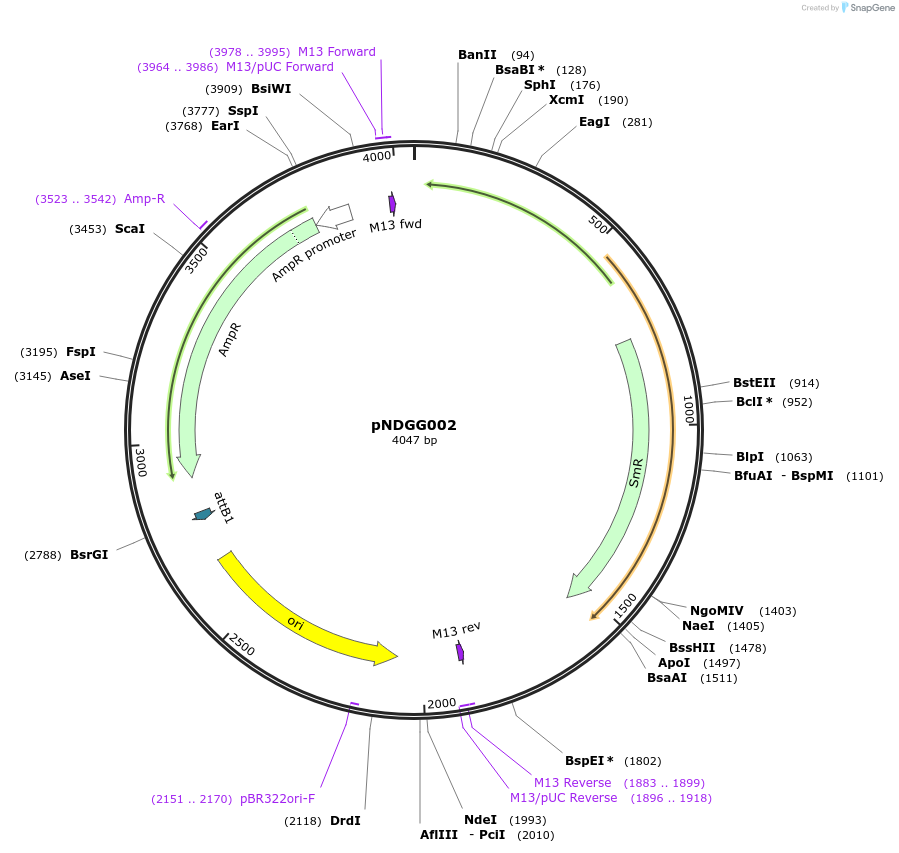
pNDGG002
Plasmid#231308PurposeVector part for BEVA; BEVA2.0 Position 2 Sp antibiotic resistance via aadA, AmpR/SpRDepositorInsertBEVA2.0 Position 2 Sp antibiotic resistance via aadA, AmpR/SpR
ExpressionBacterialAvailable SinceFeb. 20, 2025AvailabilityAcademic Institutions and Nonprofits only -
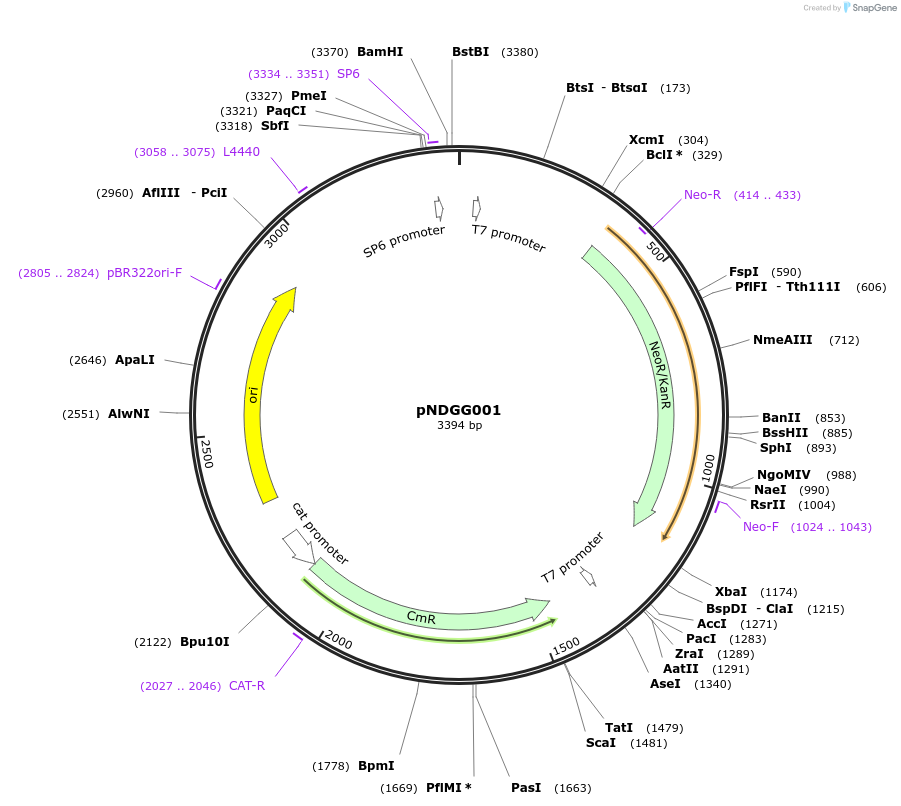
pNDGG001
Plasmid#231307PurposeVector part for BEVA; BEVA2.0 Position 2 Km/Nm antibiotic resistance via nptII, CmR/KmR/NmR.DepositorInsertBEVA2.0 Position 2 Km/Nm antibiotic resistance via nptII, CmR/KmR/NmR.
ExpressionBacterialAvailable SinceFeb. 20, 2025AvailabilityAcademic Institutions and Nonprofits only