We narrowed to 52,592 results for: SON
-
-
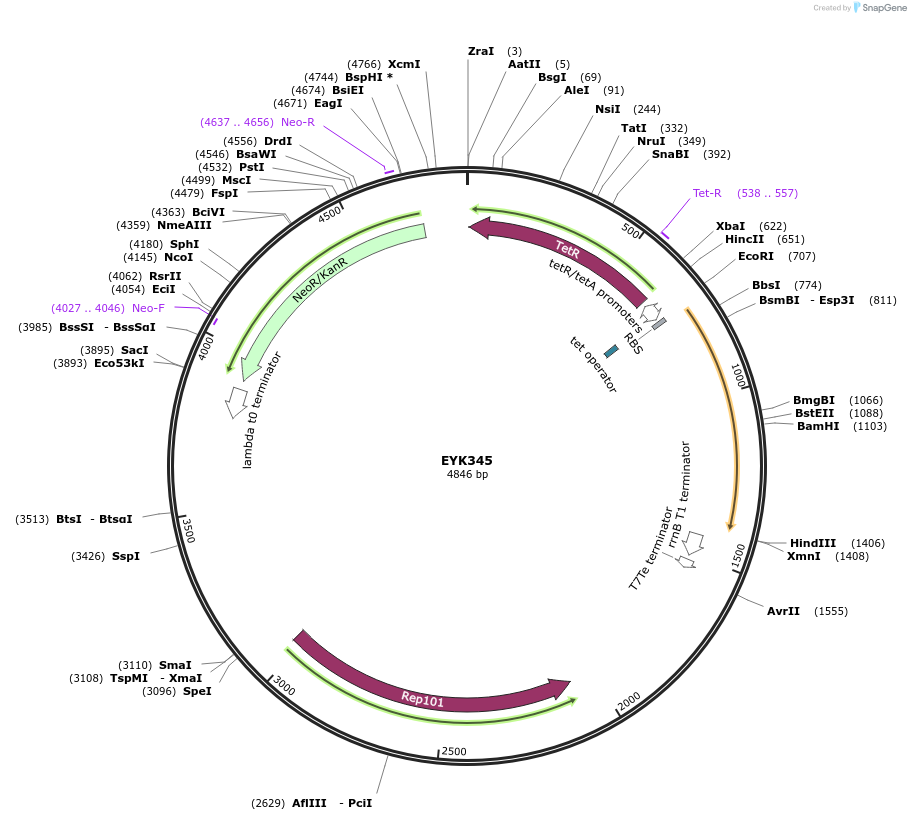
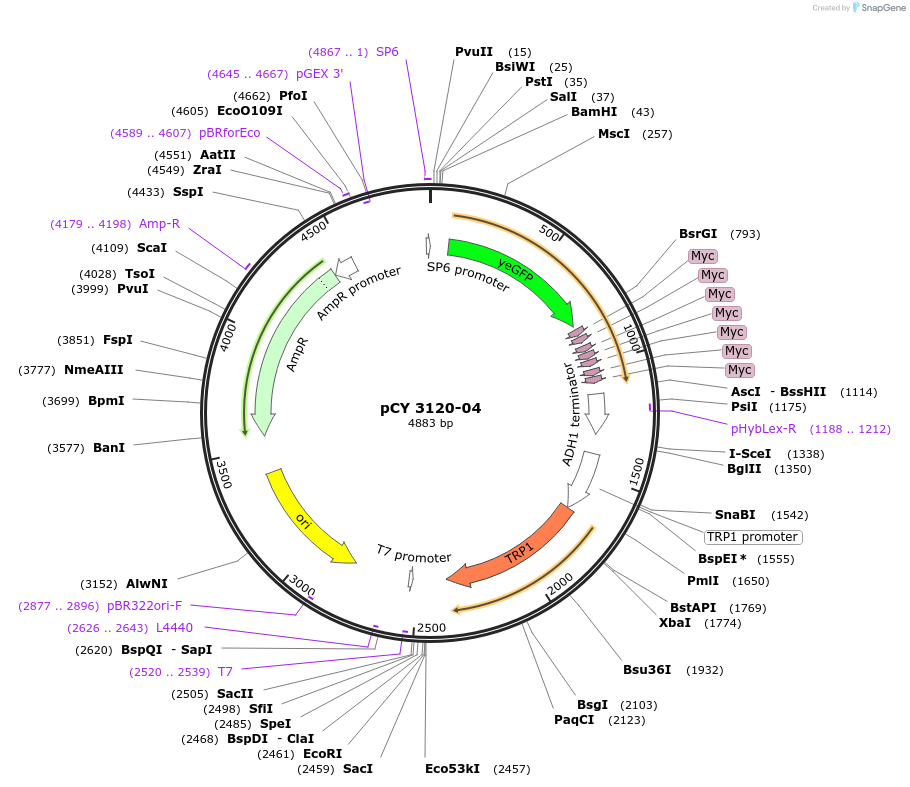
pCY 3120-04
Plasmid#36238DepositorInsertTRP1
UseYeast cassette for pcr and homologous recombinati…TagsyEpolylinker-yECitrine-13Myc-TRP1Available SinceAug. 8, 2012AvailabilityAcademic Institutions and Nonprofits only -
pJD112
Plasmid#23088DepositorInsertyeast Histone H3-2 and Histone H4-2
ExpressionYeastMutationHistone 3 Changed Serine 10 to Aspartate, H3 S10D.Available SinceFeb. 5, 2010AvailabilityAcademic Institutions and Nonprofits only -
pJD113
Plasmid#23089DepositorInsertyeast Histone H3-2 and Histone H4-2
ExpressionYeastMutationHistone H3 changed Serine 10 to Glutamamte, H3 S1…Available SinceFeb. 5, 2010AvailabilityAcademic Institutions and Nonprofits only -
-
-
-
-
-
-
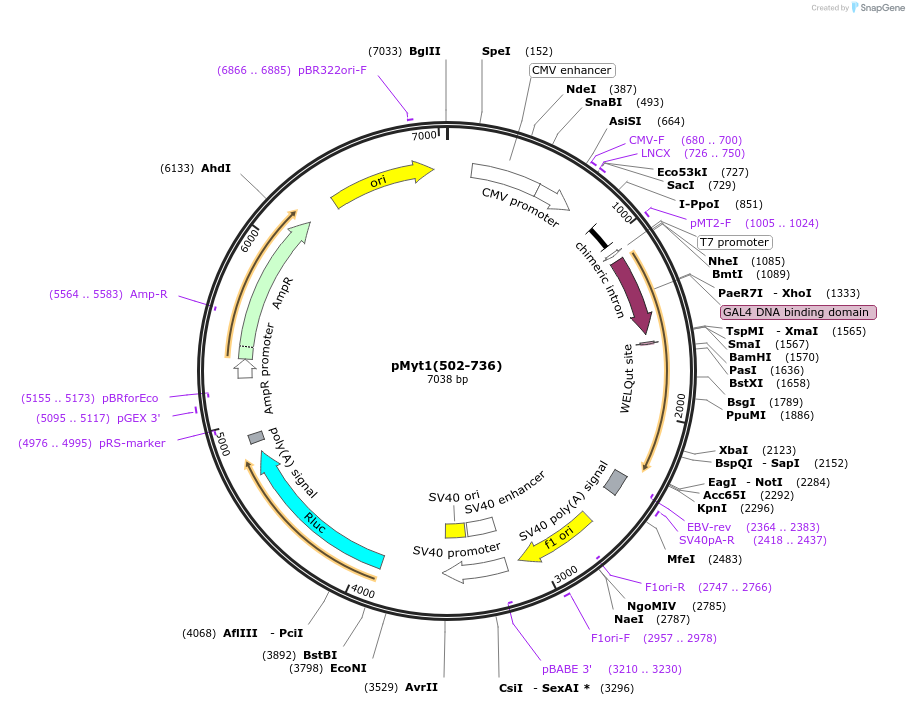
pMyt1(502-736)
Plasmid#22654DepositorInsertMyelin Transcription Factor 1 (Myt1 Mouse)
ExpressionMammalianMutationOnly bp 2035-2739 or CD of mouse Myt1Available SinceNov. 24, 2009AvailabilityAcademic Institutions and Nonprofits only -
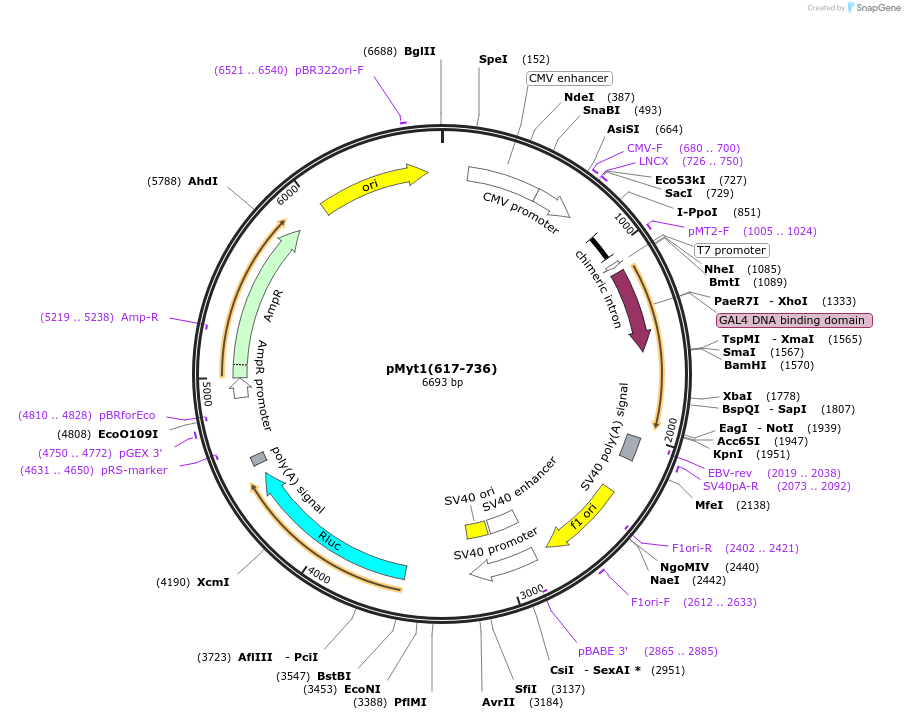
pMyt1(617-736)
Plasmid#22656DepositorInsertmyelin Transcription Factor 1 (Myt1 Mouse)
ExpressionMammalianMutationSecond 1/2 of Myt1 central domain (2380-2739Available SinceNov. 24, 2009AvailabilityAcademic Institutions and Nonprofits only -
RMC sup35 promotor integr
Plasmid#1175DepositorAvailable SinceJan. 21, 2005AvailabilityAcademic Institutions and Nonprofits only -
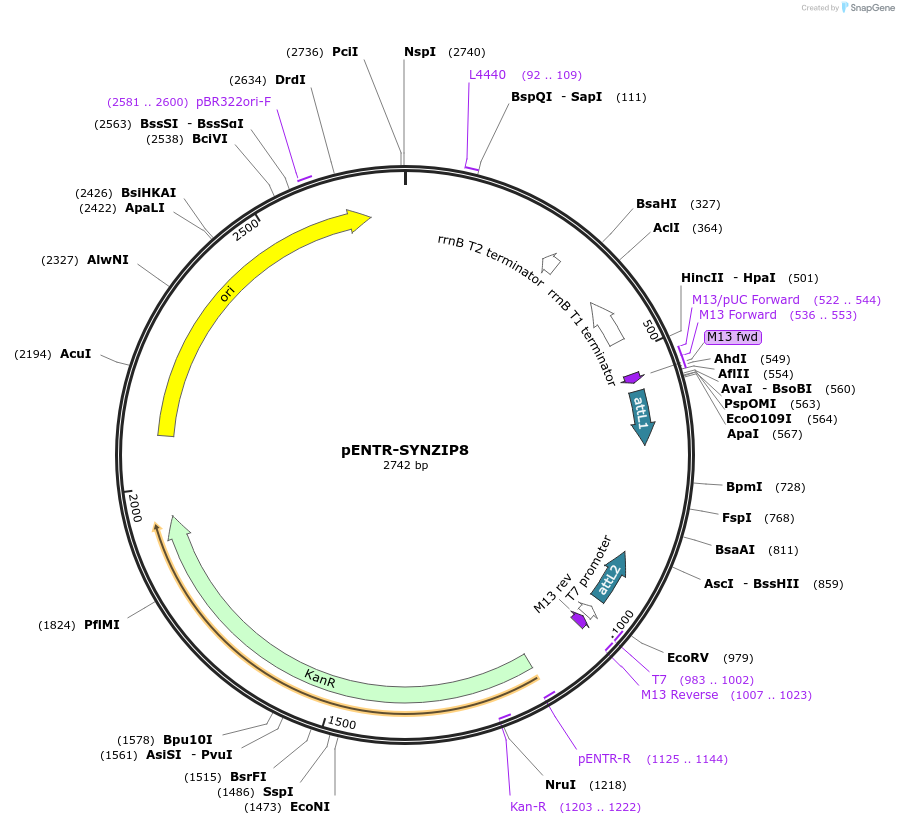
pENTR-SYNZIP8
Plasmid#80664PurposeEncodes the synthetic coiled coil peptide SYNZIP8DepositorInsertSYNZIP8
UseEntry vectorAvailabilityAcademic Institutions and Nonprofits only -
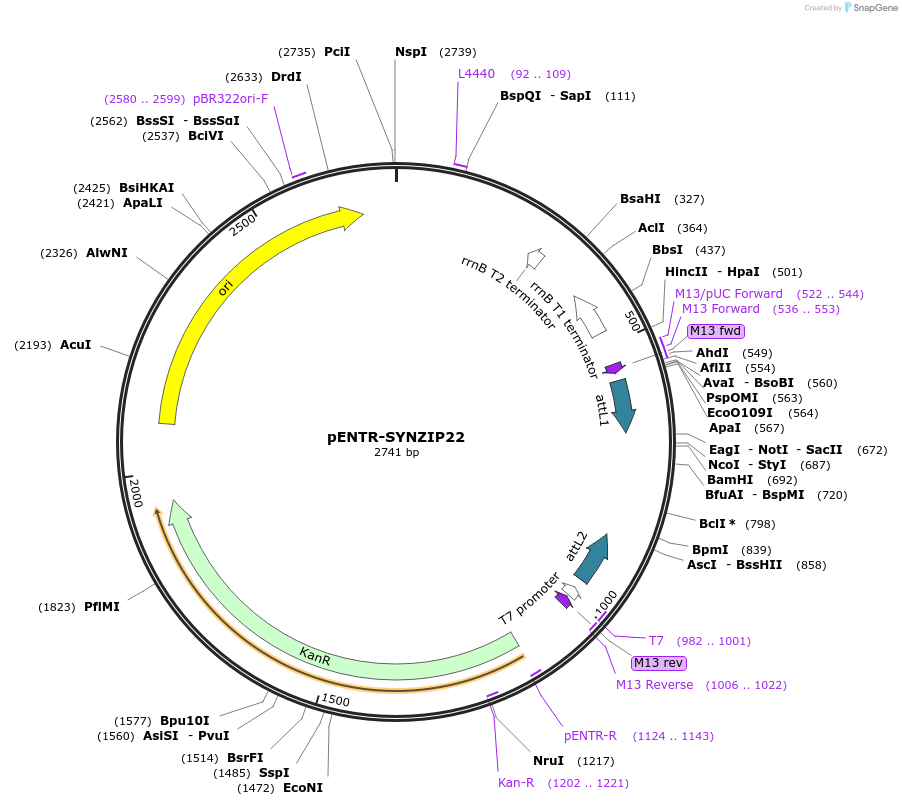
pENTR-SYNZIP22
Plasmid#80676PurposeEncodes the synthetic coiled coil peptide SYNZIP22DepositorInsertSYNZIP22
UseEntry vectorAvailabilityAcademic Institutions and Nonprofits only -
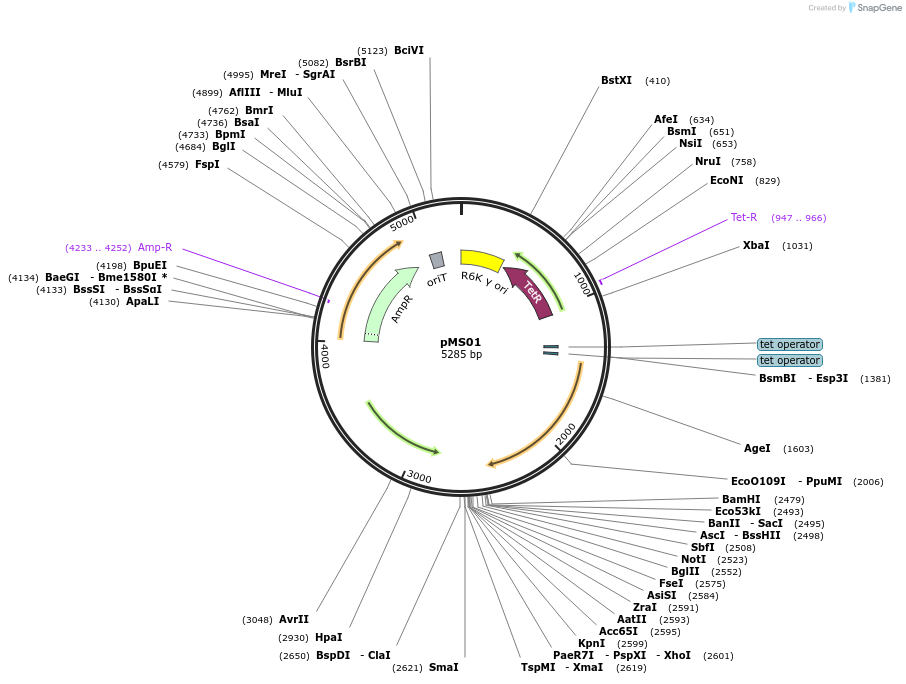
pMS01
Plasmid#230064PurposeFor allelic exchange into Bacteroides fragilis (pLGB13-based); ermC, aTC-inducible counterselection with ssbfe1; R6K ori, RP4 oriTDepositorTypeEmpty backboneUseSynthetic Biology; Allelic exchangeExpressionBacterialPromoterNoneAvailable SinceJune 23, 2025AvailabilityAcademic Institutions and Nonprofits only -
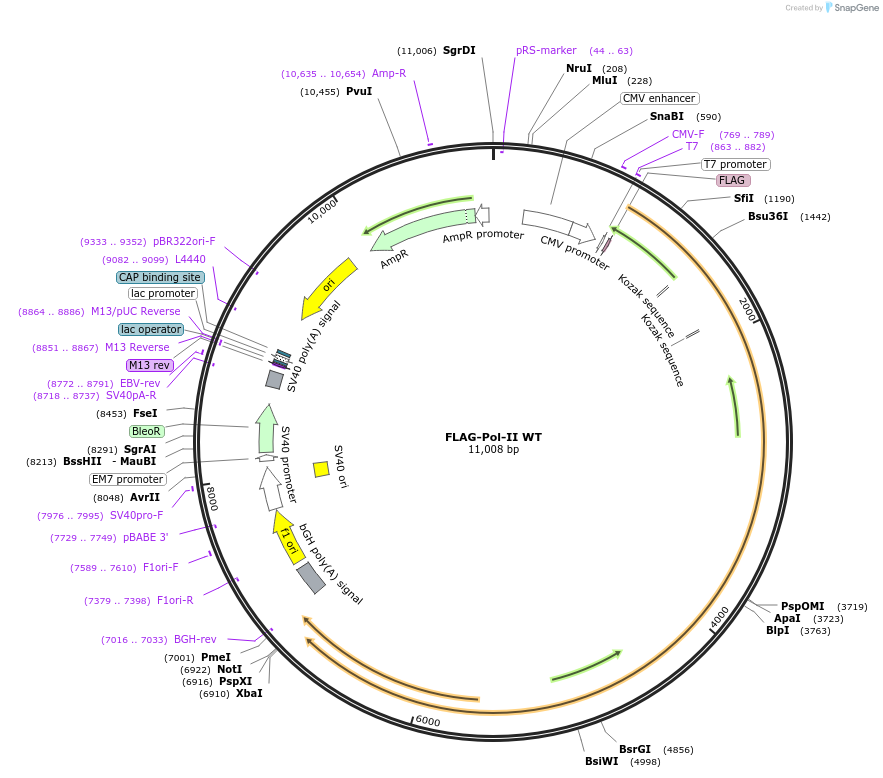
FLAG-Pol-II WT
Plasmid#35175DepositorAvailable SinceMarch 30, 2012AvailabilityAcademic Institutions and Nonprofits only -
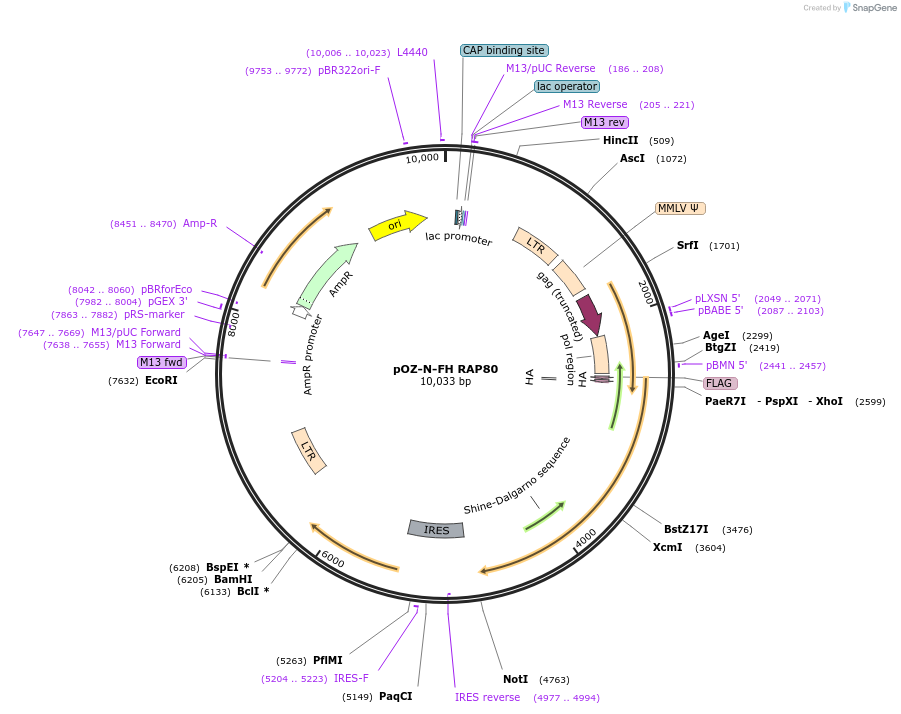
pOZ-N-FH RAP80
Plasmid#27501DepositorAvailable SinceMarch 1, 2011AvailabilityAcademic Institutions and Nonprofits only -
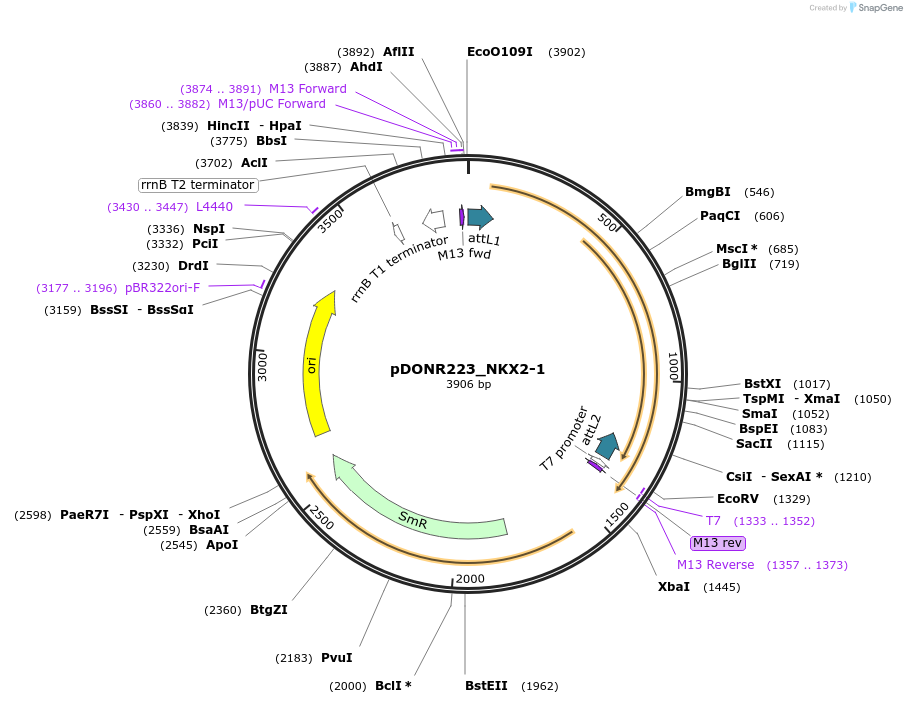
pDONR223_NKX2-1
Plasmid#221472PurposeGateway cloning of NKX2-1 short isoform overexpressionDepositorInsertNKX2-1 (NKX2-1 Human)
UseGateway donor vectorAvailable SinceJuly 2, 2024AvailabilityAcademic Institutions and Nonprofits only -
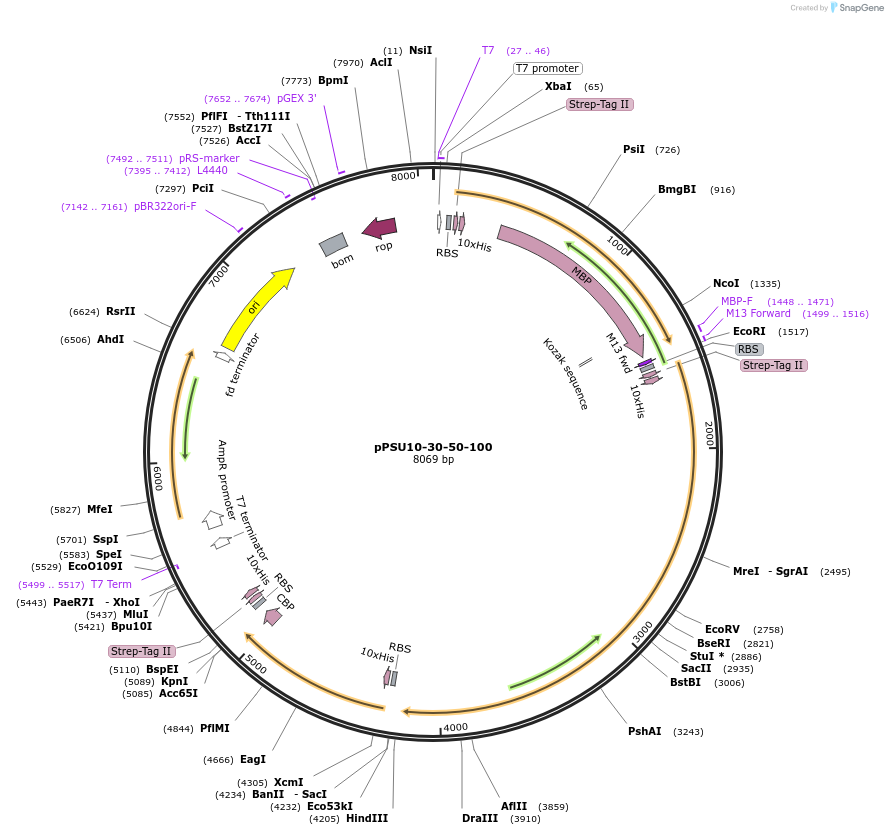
pPSU10-30-50-100
Plasmid#173838PurposeCoexpresses 10, 30, 50 and 100 kD Penn State ladder proteinsDepositorInsertSTRHSTPAB
ExpressionBacterialPromoterT7Available SinceAug. 17, 2021AvailabilityAcademic Institutions and Nonprofits only