We narrowed to 53,288 results for: SON;
-
Plasmid#128378PurposeComplementation in Cryptococcus neoformans. Targets DNA constructs to a Safe Haven Site 4, a small gene-free region. Contains the HYG resistance marker.DepositorTypeEmpty backboneUseUnspecifiedPromoterACT1Available SinceJan. 8, 2020AvailabilityAcademic Institutions and Nonprofits only
-
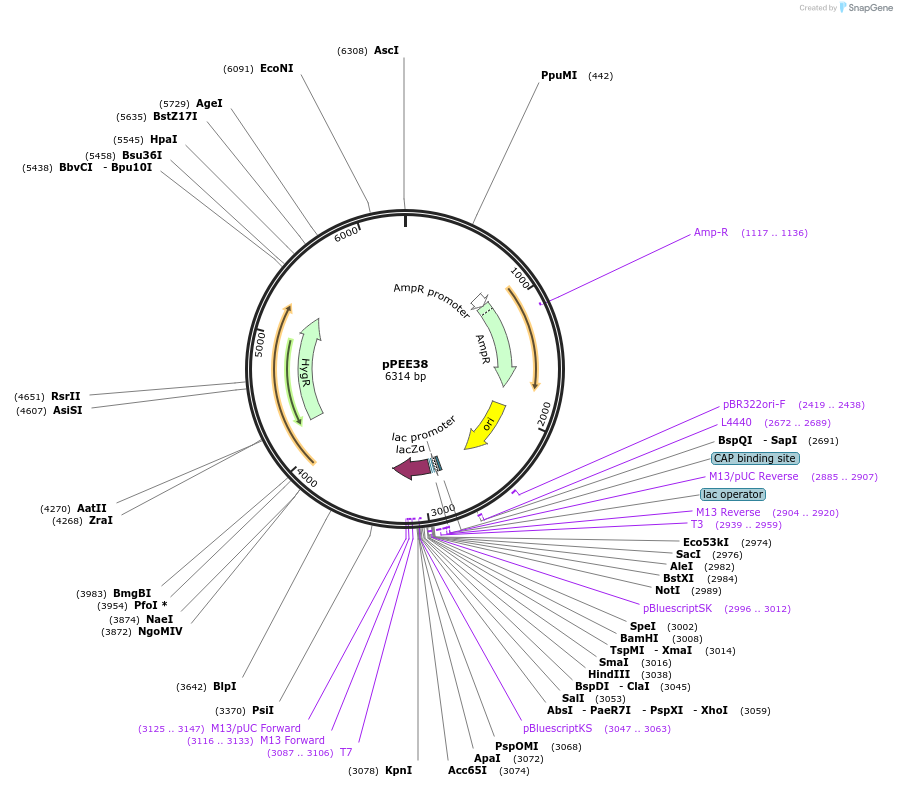
pPEE41
Plasmid#128381PurposeComplementation in Cryptococcus neoformans. Targets DNA constructs to a Safe Haven Site 7, a small gene-free region. Contains the HYG resistance marker.DepositorTypeEmpty backboneUseUnspecifiedPromoterACT1Available SinceAug. 20, 2019AvailabilityAcademic Institutions and Nonprofits only -
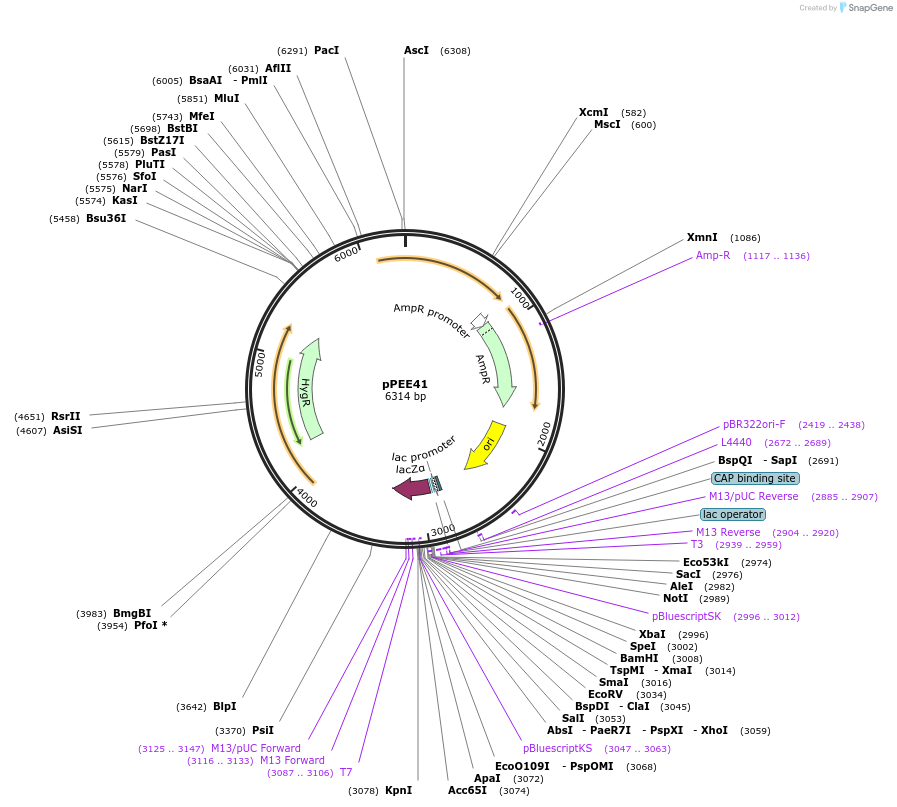
pPEE40
Plasmid#128380PurposeComplementation in Cryptococcus neoformans. Targets DNA constructs to a Safe Haven Site 6, a small gene-free region. Contains the HYG resistance marker.DepositorTypeEmpty backboneUseUnspecifiedPromoterACT1Available SinceAug. 20, 2019AvailabilityAcademic Institutions and Nonprofits only -
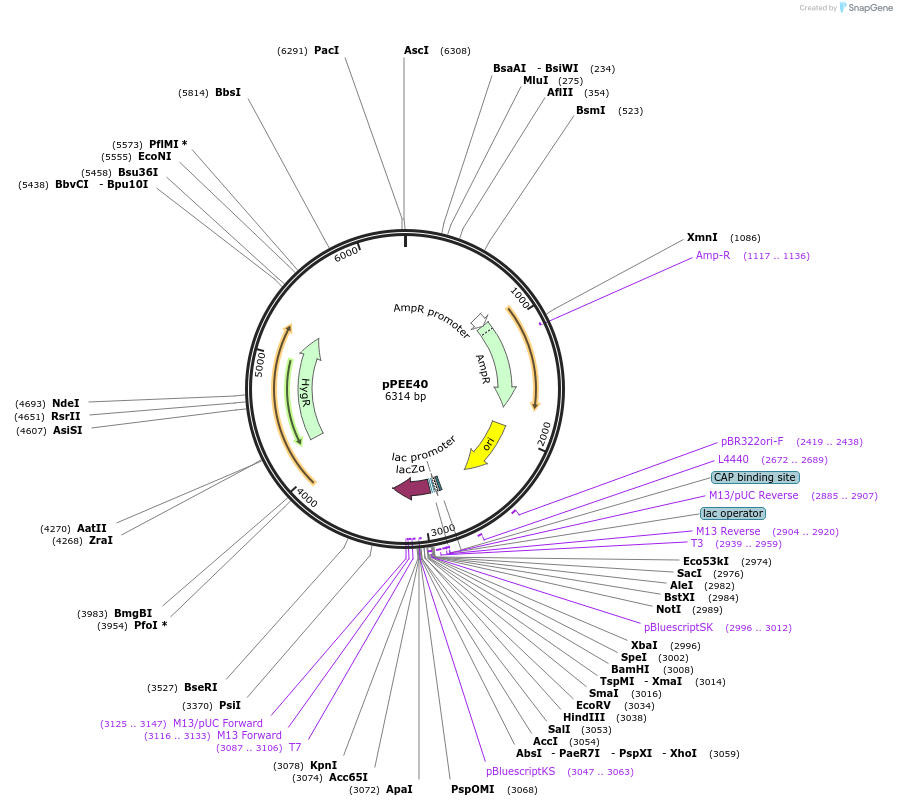
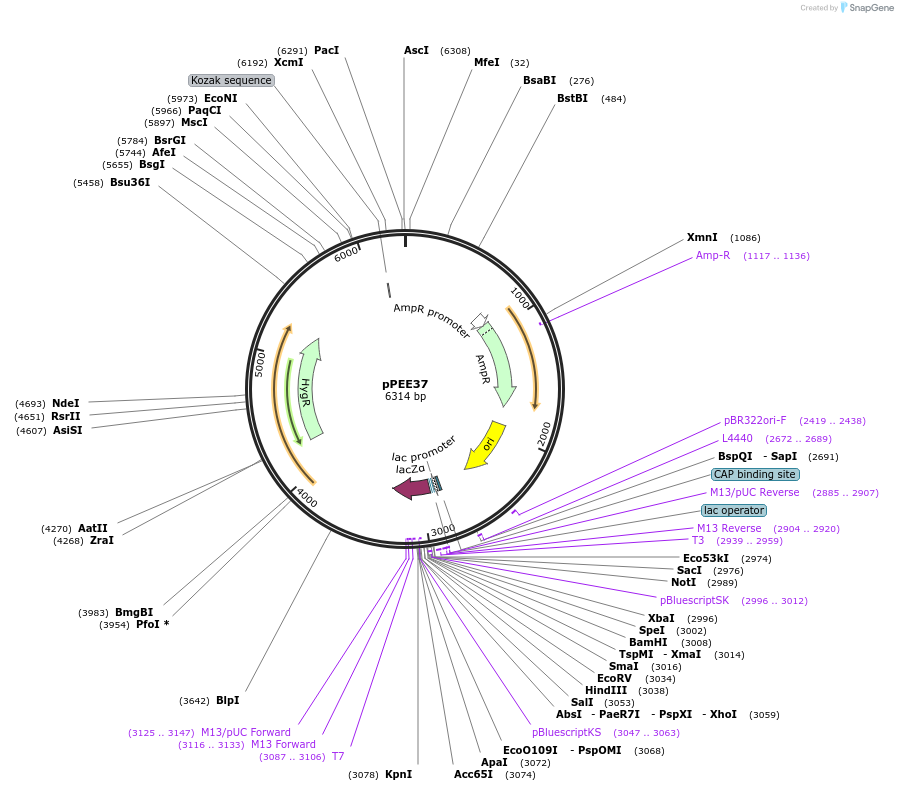
pPEE37
Plasmid#128377PurposeComplementation in Cryptococcus neoformans. Targets DNA constructs to a Safe Haven Site 3, a small gene-free region. Contains the HYG resistance marker.DepositorTypeEmpty backboneUseUnspecifiedPromoterACT1Available SinceAug. 20, 2019AvailabilityAcademic Institutions and Nonprofits only -
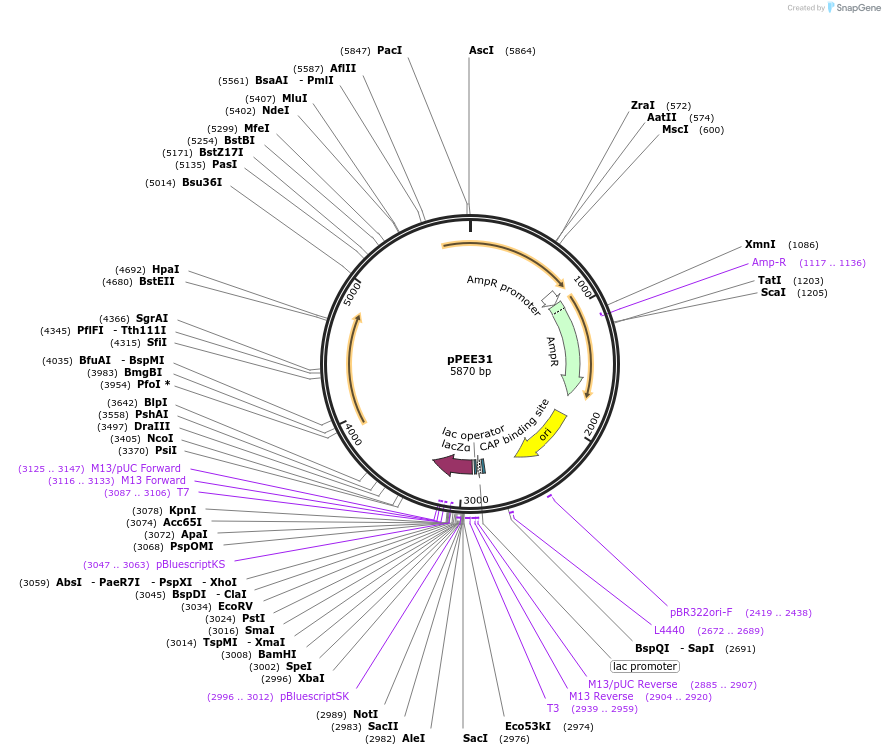
pPEE31
Plasmid#128371PurposeComplementation in Cryptococcus neoformans. Targets DNA constructs to a Safe Haven Site 7, a small gene-free region. Contains the nourseothricin resistance marker (NAT)DepositorTypeEmpty backboneUseUnspecifiedPromoterACT1Available SinceAug. 20, 2019AvailabilityAcademic Institutions and Nonprofits only -
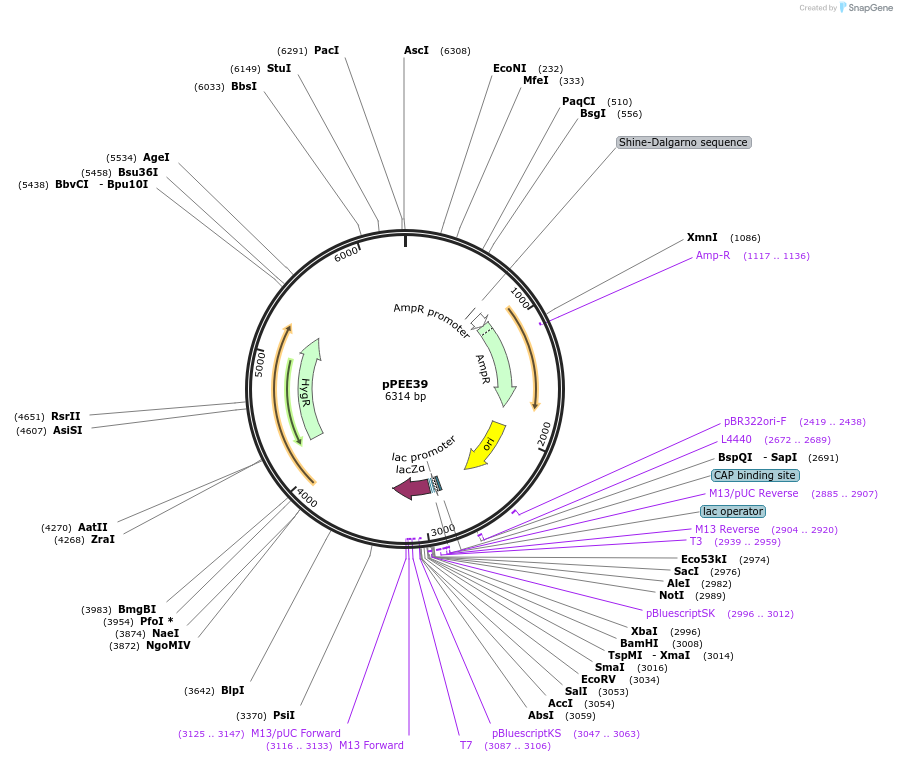
pPEE39
Plasmid#128379PurposeComplementation in Cryptococcus neoformans. Targets DNA constructs to a Safe Haven Site 5, a small gene-free region. Contains the HYG resistance marker.DepositorTypeEmpty backboneUseUnspecifiedPromoterACT1Available SinceAug. 20, 2019AvailabilityAcademic Institutions and Nonprofits only -
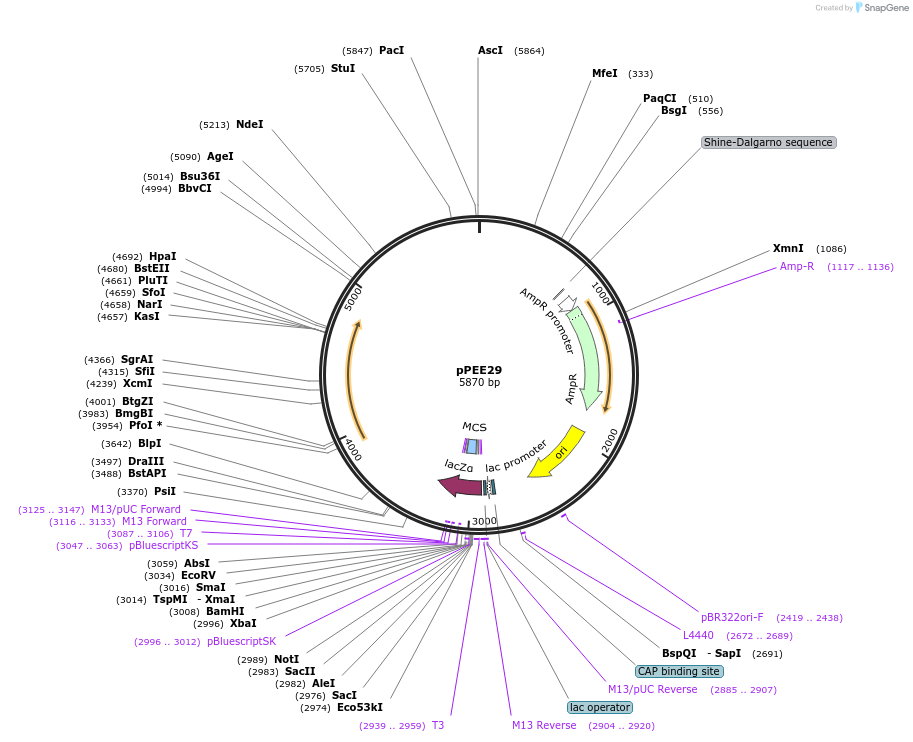
pPEE29
Plasmid#128369PurposeComplementation in Cryptococcus neoformans. Targets DNA constructs to a Safe Haven Site 5, a small gene-free region. Contains the nourseothricin resistance marker (NAT)DepositorTypeEmpty backboneUseUnspecifiedPromoterACT1Available SinceAug. 13, 2019AvailabilityAcademic Institutions and Nonprofits only -
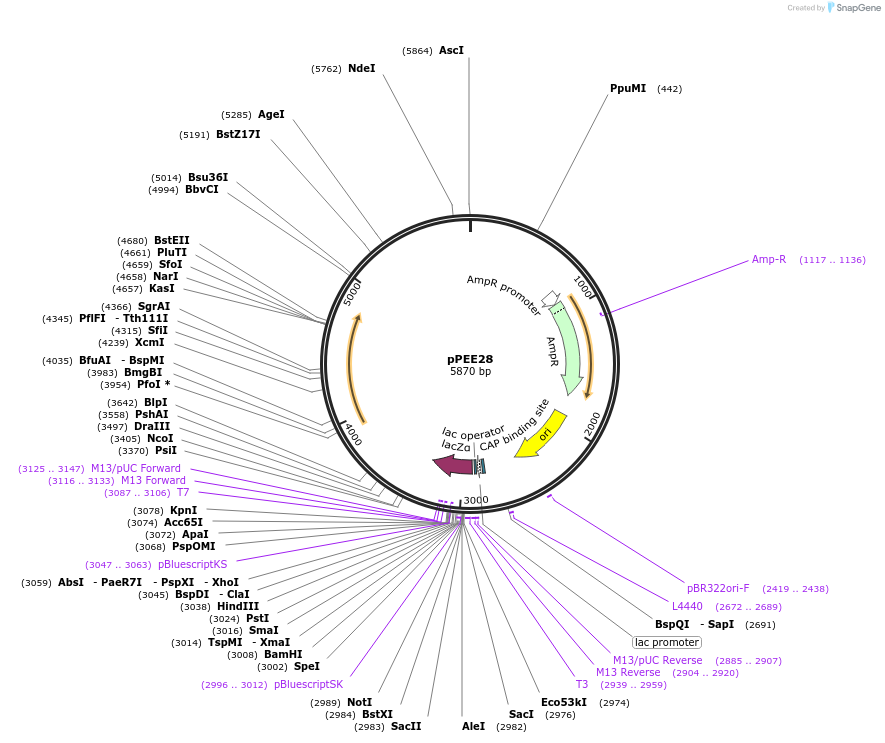
pPEE28
Plasmid#128368PurposeComplementation in Cryptococcus neoformans. Targets DNA constructs to a Safe Haven Site 4, a small gene-free region. Contains the nourseothricin resistance marker (NAT)DepositorTypeEmpty backboneUseUnspecifiedPromoterACT1Available SinceAug. 13, 2019AvailabilityAcademic Institutions and Nonprofits only -
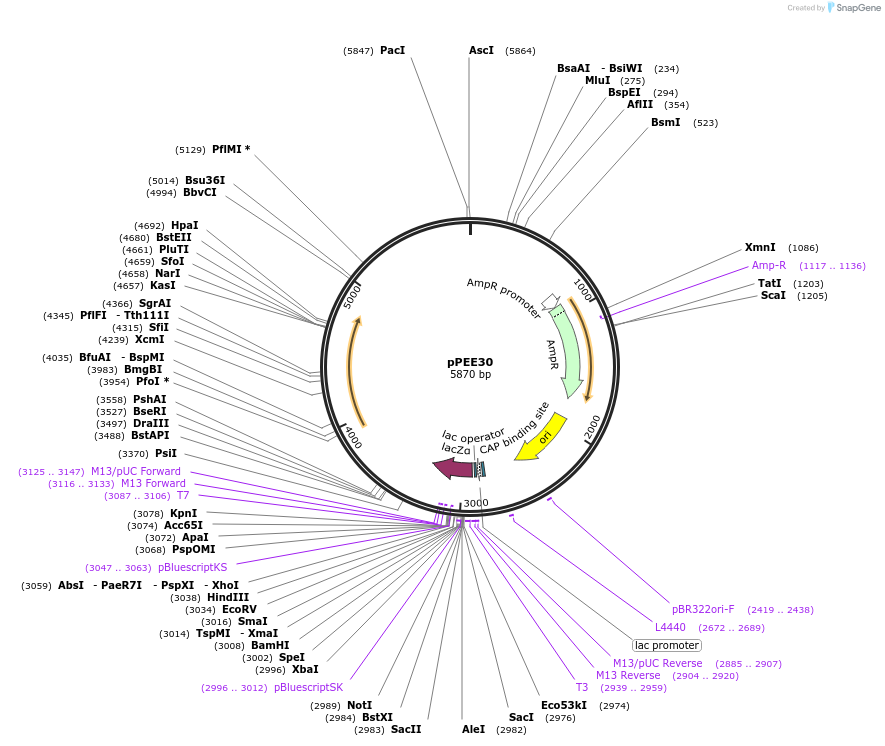
pPEE30
Plasmid#128370PurposeComplementation in Cryptococcus neoformans. Targets DNA constructs to a Safe Haven Site 6, a small gene-free region. Contains the nourseothricin resistance marker (NAT)DepositorTypeEmpty backboneUseUnspecifiedPromoterACT1Available SinceAug. 13, 2019AvailabilityAcademic Institutions and Nonprofits only -
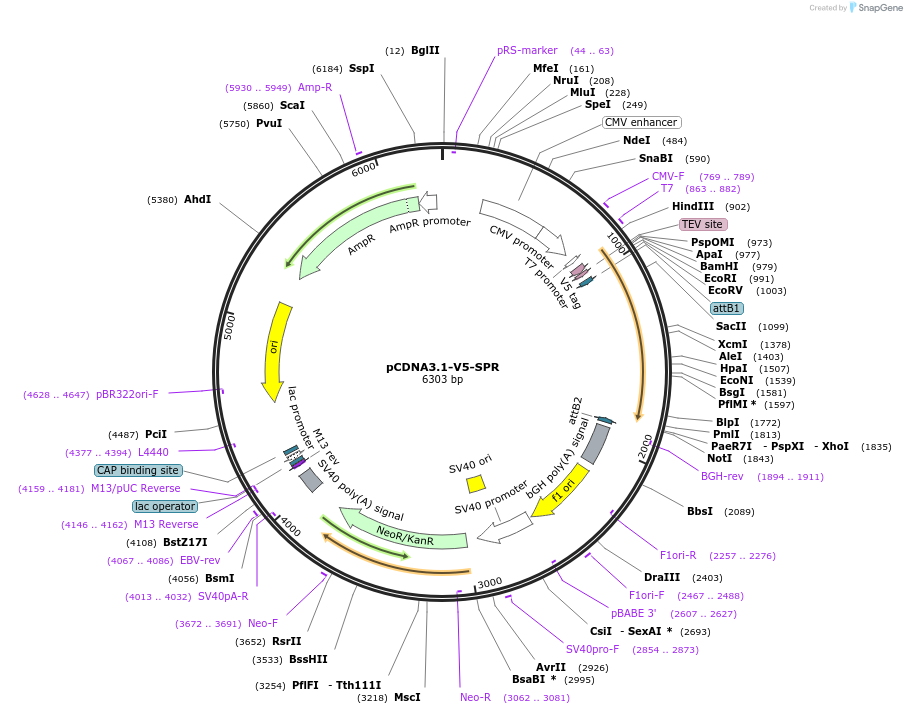
pCDNA3.1-V5-SPR
Plasmid#127236PurposeEukaryotic expression of SPRDepositorAvailable SinceJune 17, 2019AvailabilityAcademic Institutions and Nonprofits only -
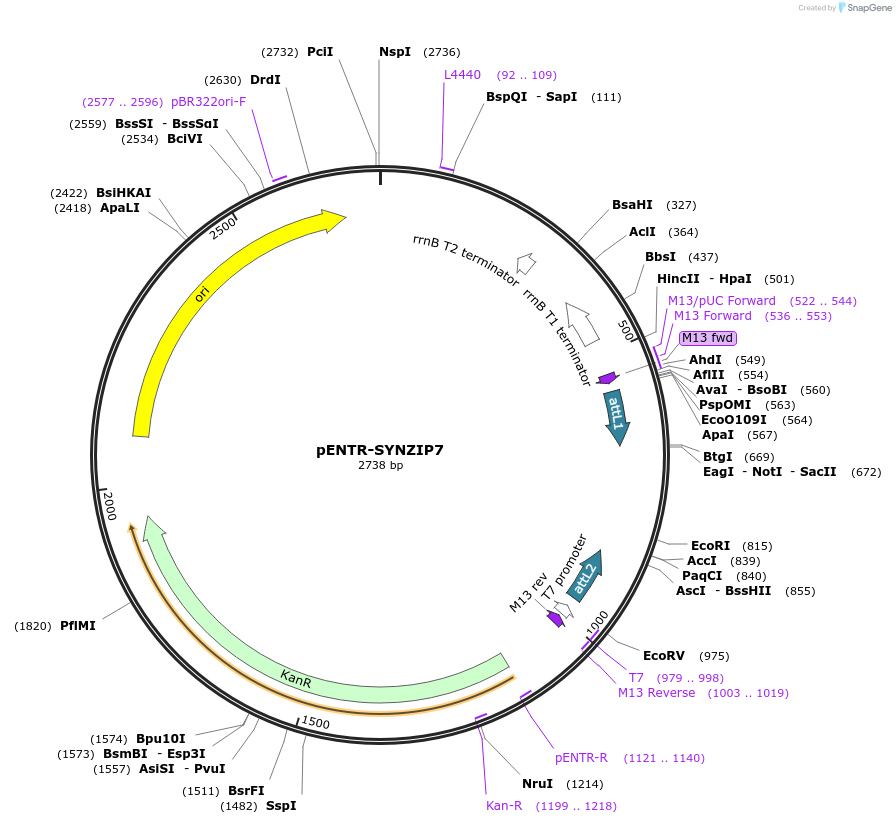
pENTR-SYNZIP7
Plasmid#80663PurposeEncodes the synthetic coiled coil peptide SYNZIP7DepositorInsertSYNZIP7
UseGateway CloningAvailable SinceFeb. 8, 2017AvailabilityAcademic Institutions and Nonprofits only -
-
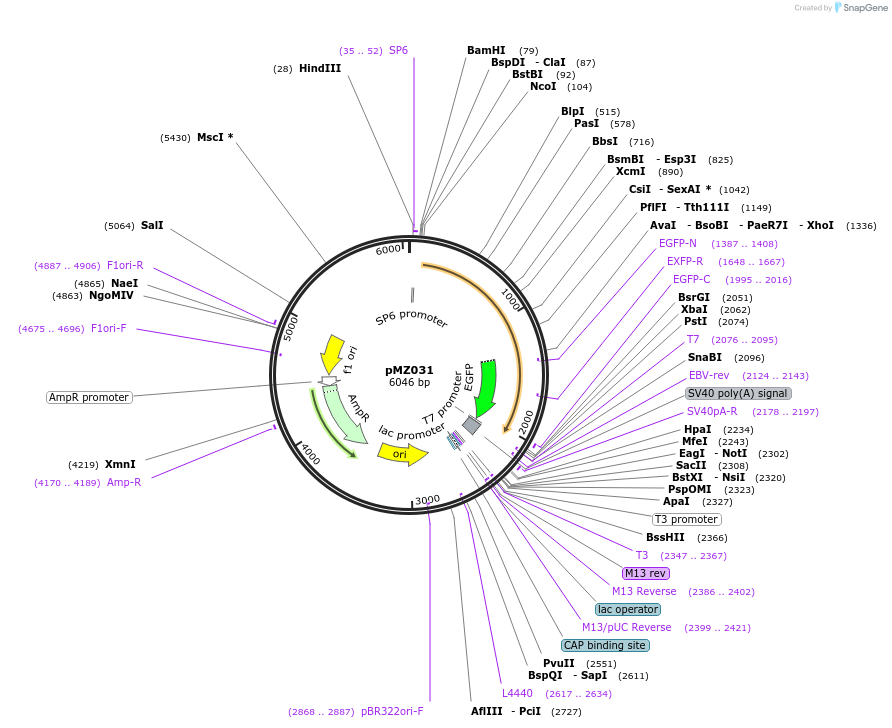
pMZ031
Plasmid#66971PurposeExpresses zebrafish Chk1 mutant with C-term GFP fluorescent tagDepositorInsertzebrafish Chk1
TagsGFPMutationmutated S256E, S280E, T292E, and S301EAvailable SinceDec. 1, 2015AvailabilityAcademic Institutions and Nonprofits only -
-
pJD112
Plasmid#23088DepositorInsertyeast Histone H3-2 and Histone H4-2
ExpressionYeastMutationHistone 3 Changed Serine 10 to Aspartate, H3 S10D.Available SinceFeb. 5, 2010AvailabilityAcademic Institutions and Nonprofits only -
pJD113
Plasmid#23089DepositorInsertyeast Histone H3-2 and Histone H4-2
ExpressionYeastMutationHistone H3 changed Serine 10 to Glutamamte, H3 S1…Available SinceFeb. 5, 2010AvailabilityAcademic Institutions and Nonprofits only -
-
-
-