We narrowed to 3,615 results for: gla
-
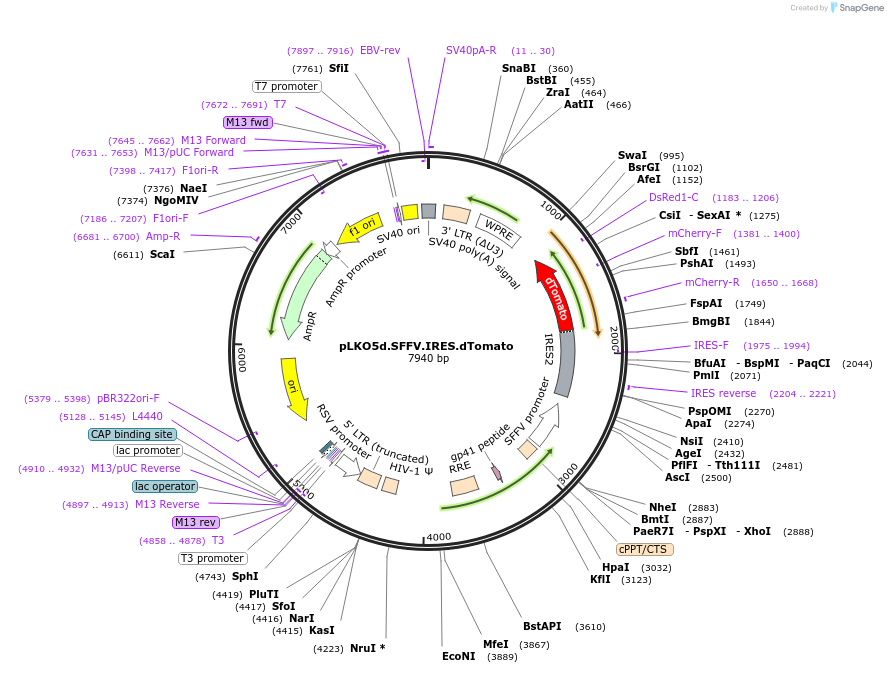
Plasmid#187191PurposeLentiviral gene overexpression (backbone/control)DepositorTypeEmpty backboneUseLentiviralAvailable SinceFeb. 2, 2026AvailabilityAcademic Institutions and Nonprofits only
-
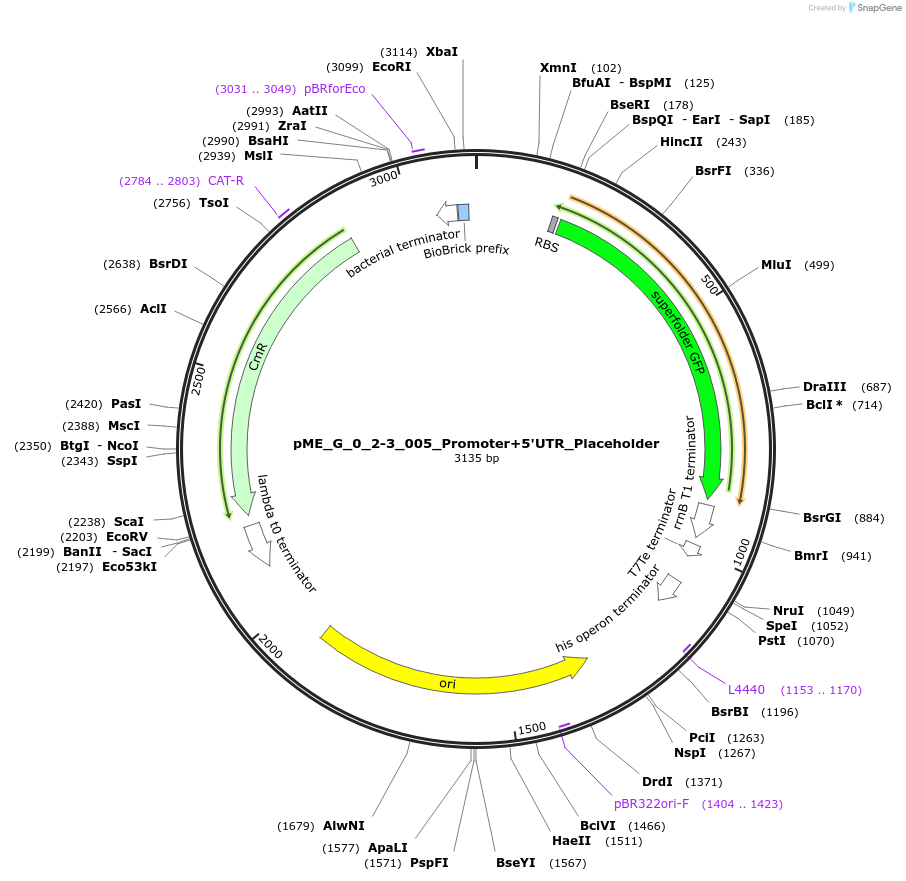
pME_G_0_2-3_005_Promoter+5'UTR_Placeholder
Plasmid#235870PurposeEncodes a sfGFP cassette in the promoter+5'UTR position used for placeholder cloningDepositorInsertPromoter + 5'UTR placeholder
UseSynthetic BiologyMutationnoneAvailable SinceOct. 6, 2025AvailabilityAcademic Institutions and Nonprofits only -
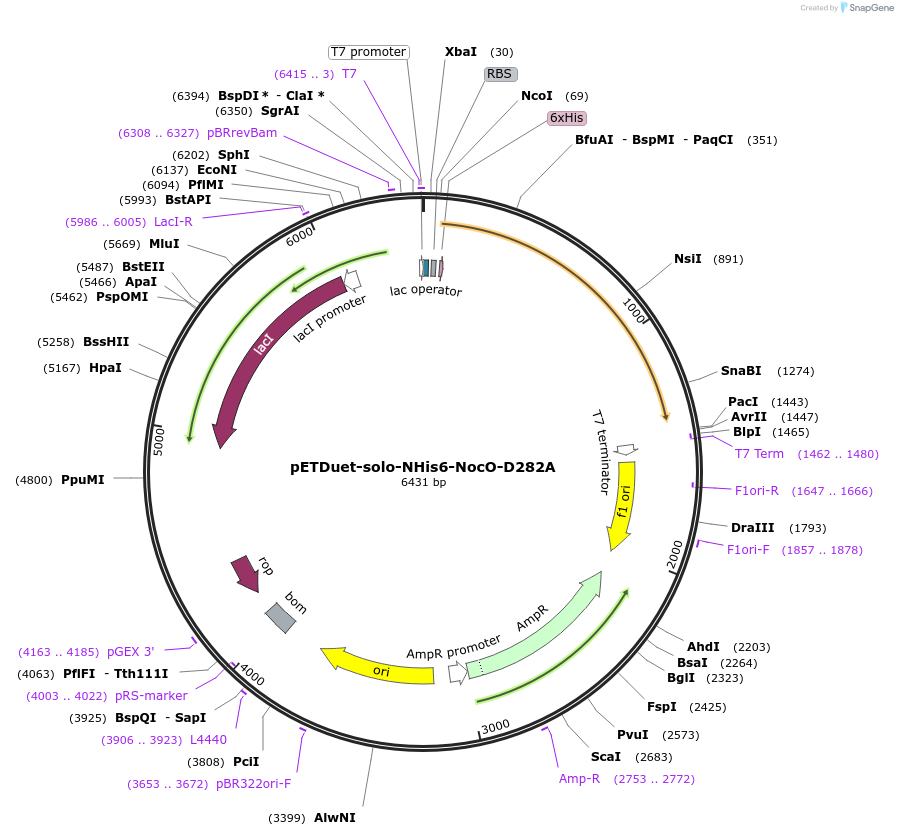
pETDuet-solo-NHis6-NocO-D282A
Plasmid#240407Purposeexpresses N-His tagged NocO D282A (diiron halogenase variant) from Nodularia sp. 06071 (mutation numbering based on amino acid position in native sequence), created from mutagenesis of WT plasmidDepositorInsertnocO
TagsHisExpressionBacterialMutationM1 of native sequence not included in insert, Asp…PromoterT7Available SinceAug. 28, 2025AvailabilityAcademic Institutions and Nonprofits only -
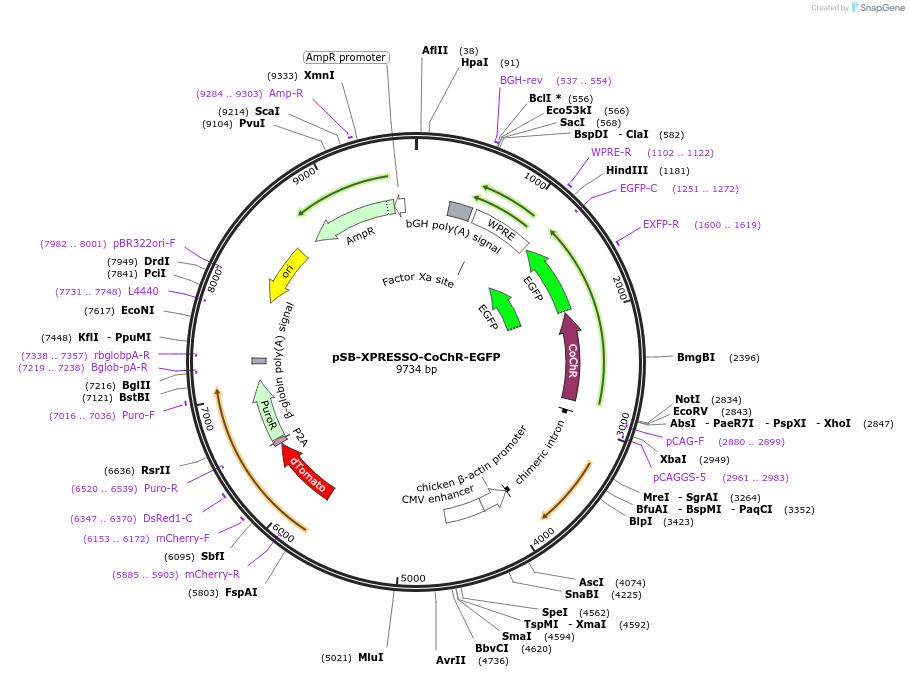
pSB-XPRESSO-CoChR-EGFP
Plasmid#237297PurposeSleeping Beauty XPRESSO vector for optogenetic CoChR-eGFP expressionDepositorInsertCoChR
TagseGFPExpressionMammalianPromoterCAGAvailable SinceAug. 21, 2025AvailabilityAcademic Institutions and Nonprofits only -
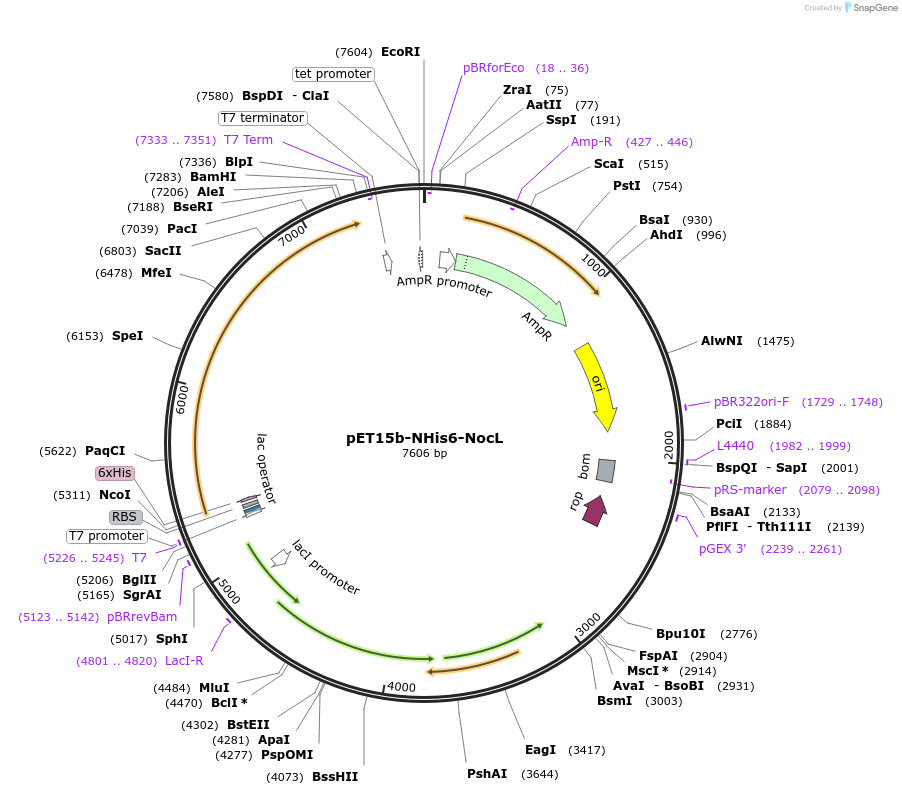
pET15b-NHis6-NocL
Plasmid#240398Purposeexpresses N-His tagged NocL (fatty acid activating ligase) from Nodularia sp. 06071DepositorInsertnocL(fatty acyl-AMP ligase)
TagsHis tagExpressionBacterialPromoterT7Available SinceJuly 17, 2025AvailabilityAcademic Institutions and Nonprofits only -
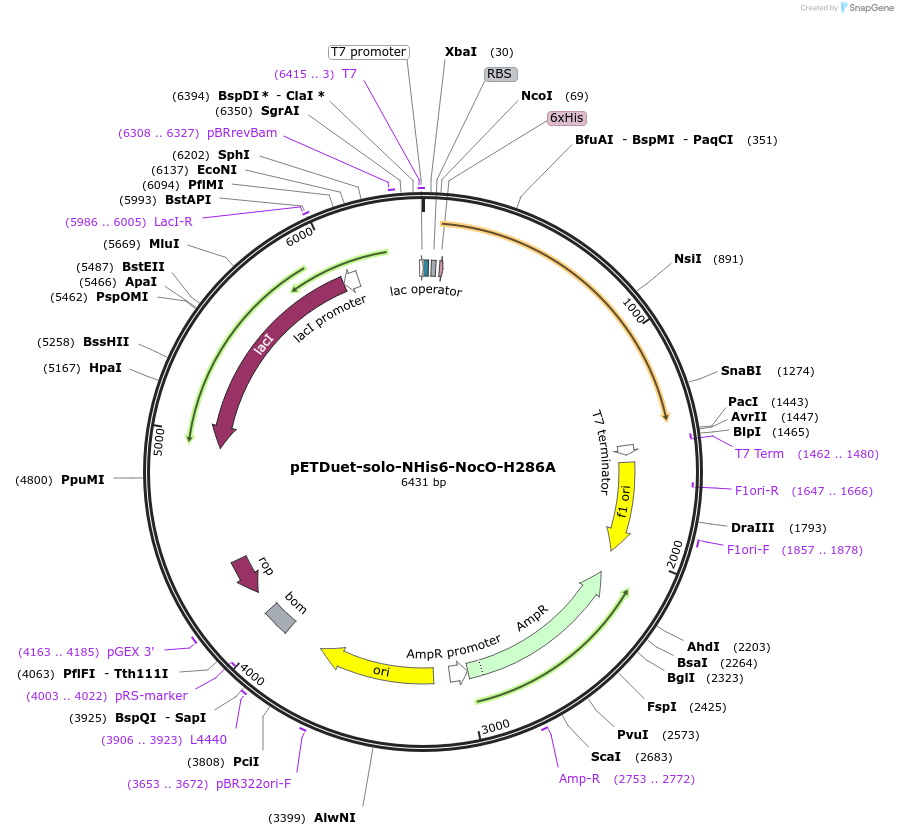
pETDuet-solo-NHis6-NocO-H286A
Plasmid#240409Purposeexpresses N-His tagged NocO H286A (diiron halogenase variant) from Nodularia sp. 06071 (mutation numbering based on amino acid position in native sequence), created from mutagenesis of WT plasmidDepositorInsertnocO
TagsHisExpressionBacterialMutationM1 of native sequence not included in insert, His…PromoterT7Available SinceJuly 10, 2025AvailabilityAcademic Institutions and Nonprofits only -
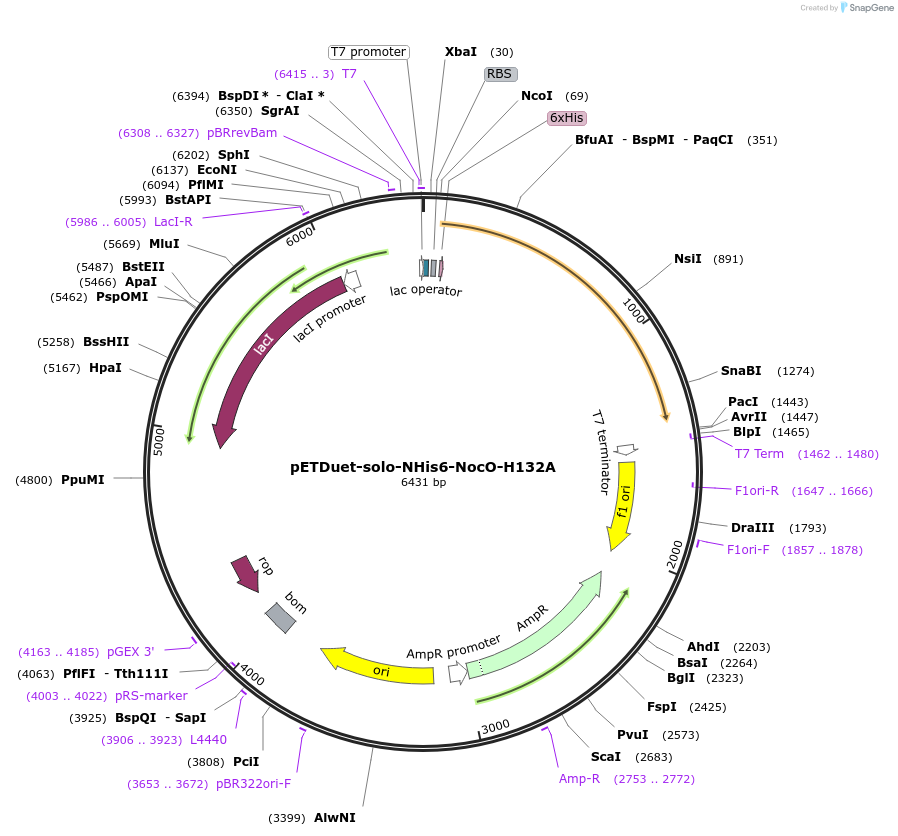
pETDuet-solo-NHis6-NocO-H132A
Plasmid#240404Purposeexpresses N-His tagged NocO H132A (diiron halogenase variant) from Nodularia sp. 06071 (mutation numbering based on amino acid position in native sequence), created from mutagenesis of WT plasmidDepositorInsertnocO
TagsHisExpressionBacterialMutationM1 of native sequence not included in insert, His…PromoterT7Available SinceJuly 10, 2025AvailabilityAcademic Institutions and Nonprofits only -
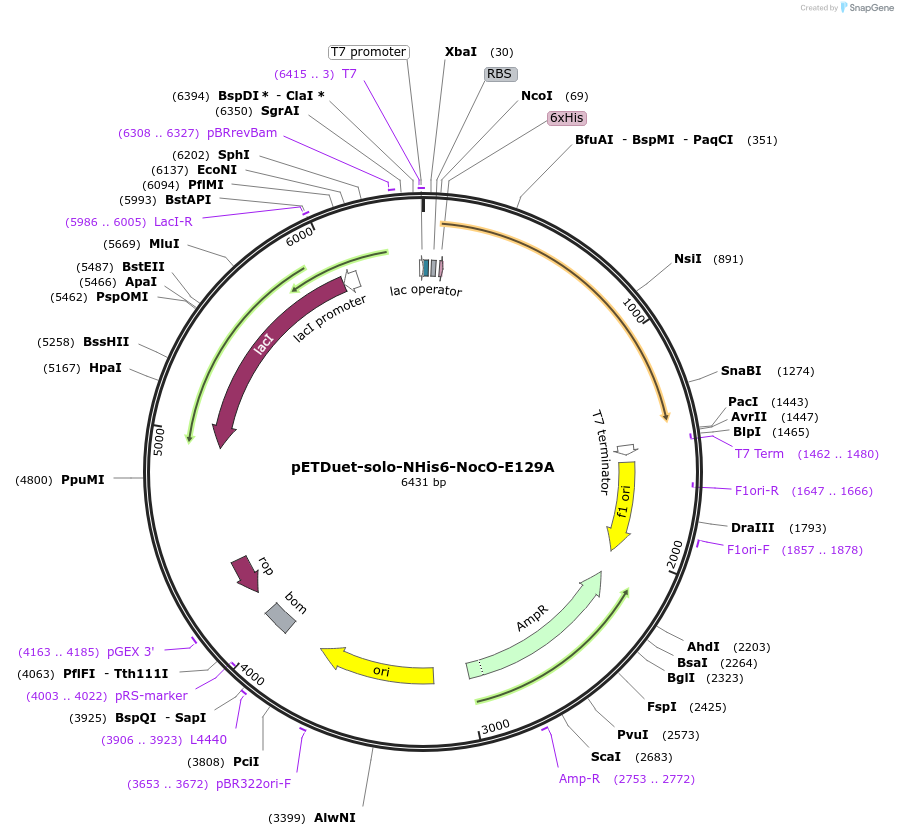
pETDuet-solo-NHis6-NocO-E129A
Plasmid#240403Purposeexpresses N-His tagged NocO E129A (diiron halogenase variant) from Nodularia sp. 06071 (mutation numbering based on amino acid position in native sequence), created from mutagenesis of WT plasmidDepositorInsertnocO
TagsHisExpressionBacterialMutationM1 of native sequence not included in insert, Glu…PromoterT7Available SinceJuly 10, 2025AvailabilityAcademic Institutions and Nonprofits only -
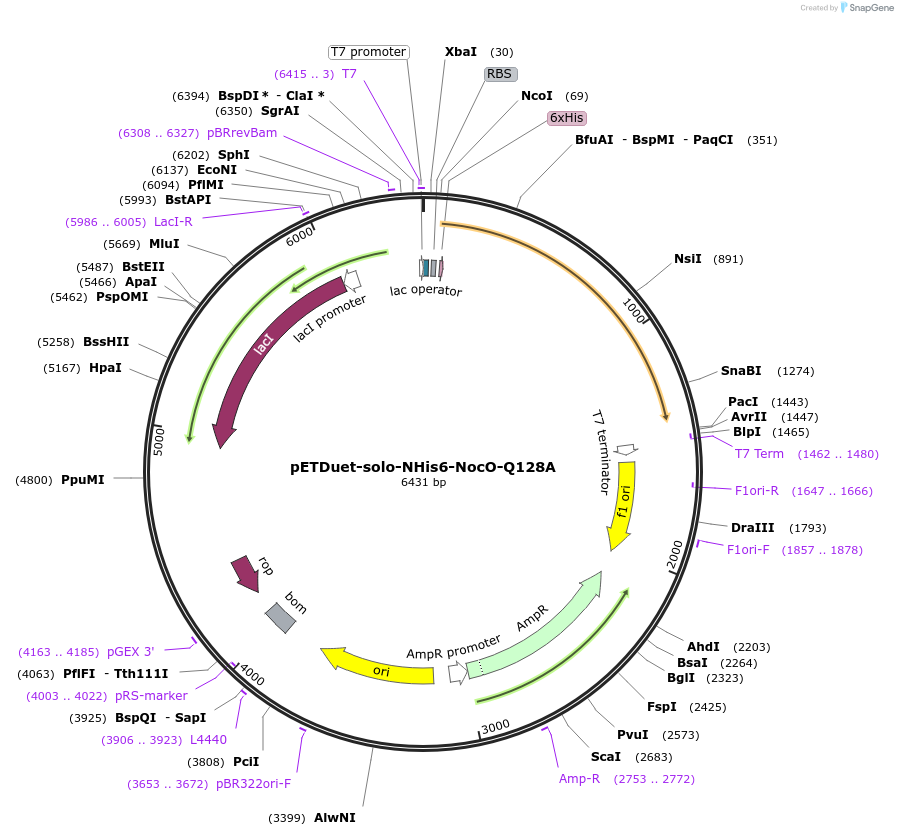
pETDuet-solo-NHis6-NocO-Q128A
Plasmid#240402Purposeexpresses N-His tagged NocO Q128A (diiron halogenase variant) from Nodularia sp. 06071 (mutation numbering based on amino acid position in native sequence), created from mutagenesis of WT plasmidDepositorInsertnocO
TagsHisExpressionBacterialMutationM1 of native sequence not included in insert, Glu…PromoterT7Available SinceJuly 10, 2025AvailabilityAcademic Institutions and Nonprofits only -
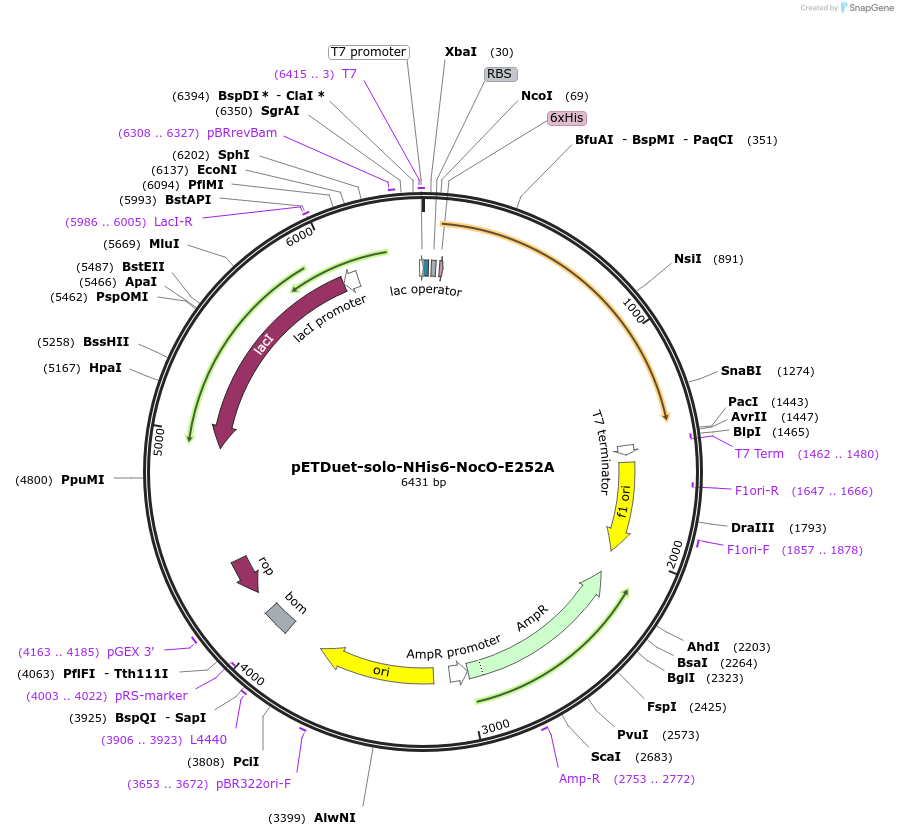
pETDuet-solo-NHis6-NocO-E252A
Plasmid#240405Purposeexpresses N-His tagged NocO E252A (diiron halogenase variant) from Nodularia sp. 06071 (mutation numbering based on amino acid position in native sequence), created from mutagenesis of WT plasmidDepositorInsertnocO
TagsHisExpressionBacterialMutationM1 of native sequence not included in insert, Glu…PromoterT7Available SinceJuly 10, 2025AvailabilityAcademic Institutions and Nonprofits only -
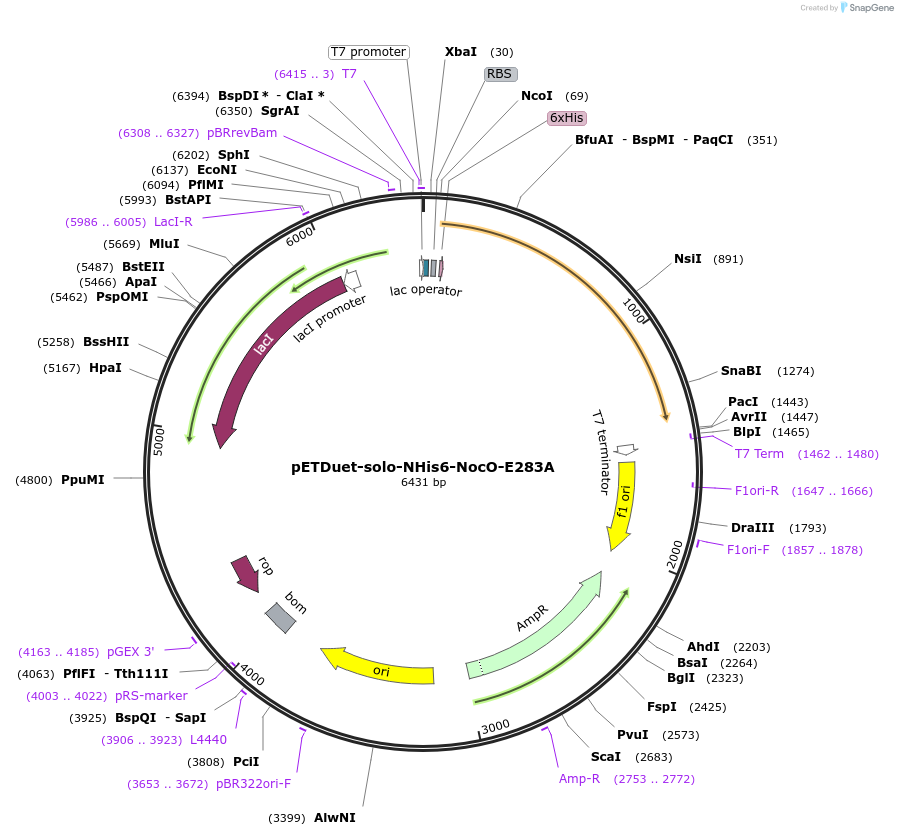
pETDuet-solo-NHis6-NocO-E283A
Plasmid#240408Purposeexpresses N-His tagged NocO E283A (diiron halogenase variant) from Nodularia sp. 06071 (mutation numbering based on amino acid position in native sequence), created from mutagenesis of WT plasmidDepositorInsertnocO
TagsHisExpressionBacterialMutationM1 of native sequence not included in insert, Glu…PromoterT7Available SinceJuly 10, 2025AvailabilityAcademic Institutions and Nonprofits only -
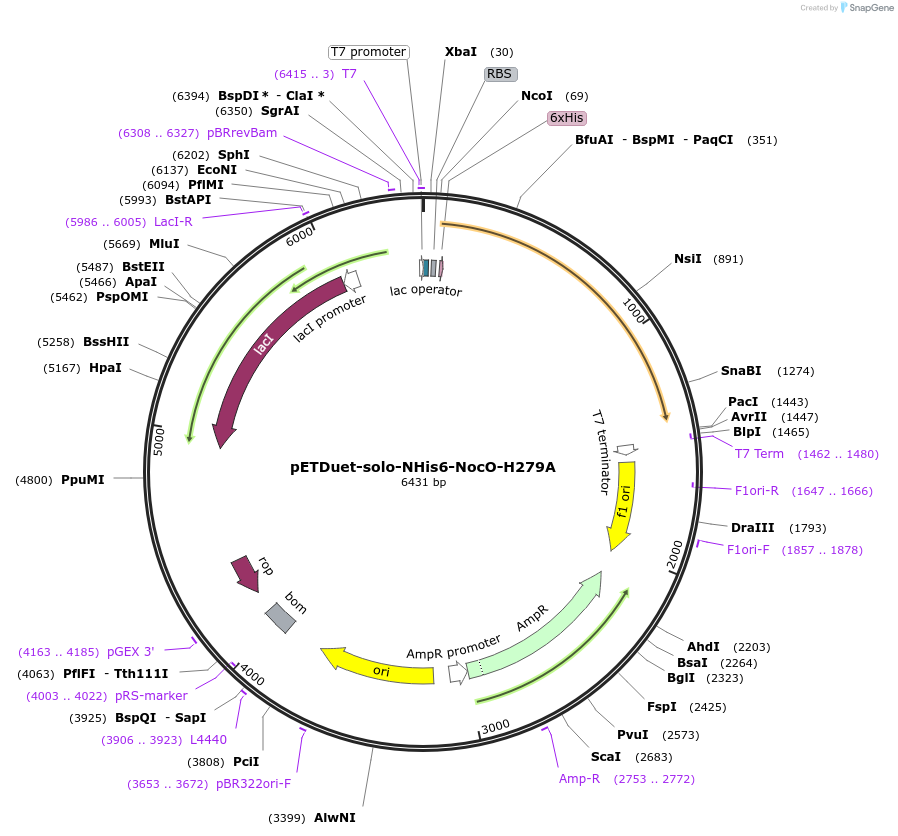
pETDuet-solo-NHis6-NocO-H279A
Plasmid#240406Purposeexpresses N-His tagged NocO H279A (diiron halogenase variant) from Nodularia sp. 06071 (mutation numbering based on amino acid position in native sequence), created from mutagenesis of WT plasmidDepositorInsertnocO
TagsHisExpressionBacterialMutationM1 of native sequence not included in insert, His…PromoterT7Available SinceJuly 10, 2025AvailabilityAcademic Institutions and Nonprofits only -
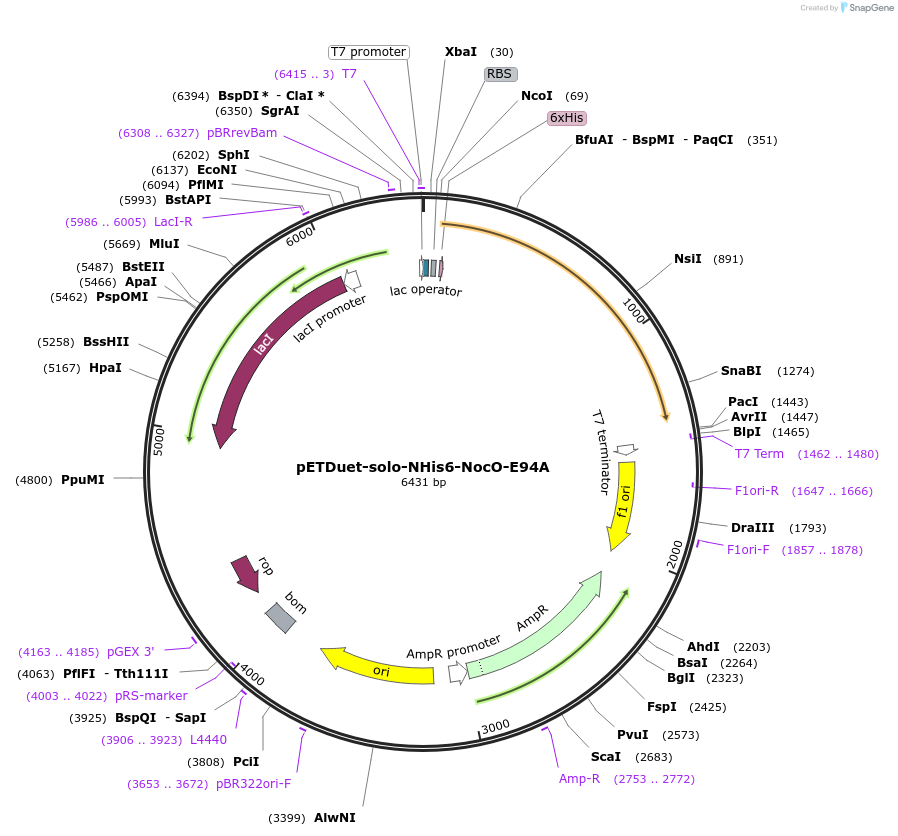
pETDuet-solo-NHis6-NocO-E94A
Plasmid#240401Purposeexpresses N-His tagged NocO E94A (diiron halogenase variant) from Nodularia sp. 06071 (mutation numbering based on amino acid position in native sequence), created from mutagenesis of WT plasmidDepositorInsertnocO
TagsHisExpressionBacterialMutationM1 of native sequence not included in insert, Glu…PromoterT7Available SinceJuly 10, 2025AvailabilityAcademic Institutions and Nonprofits only -
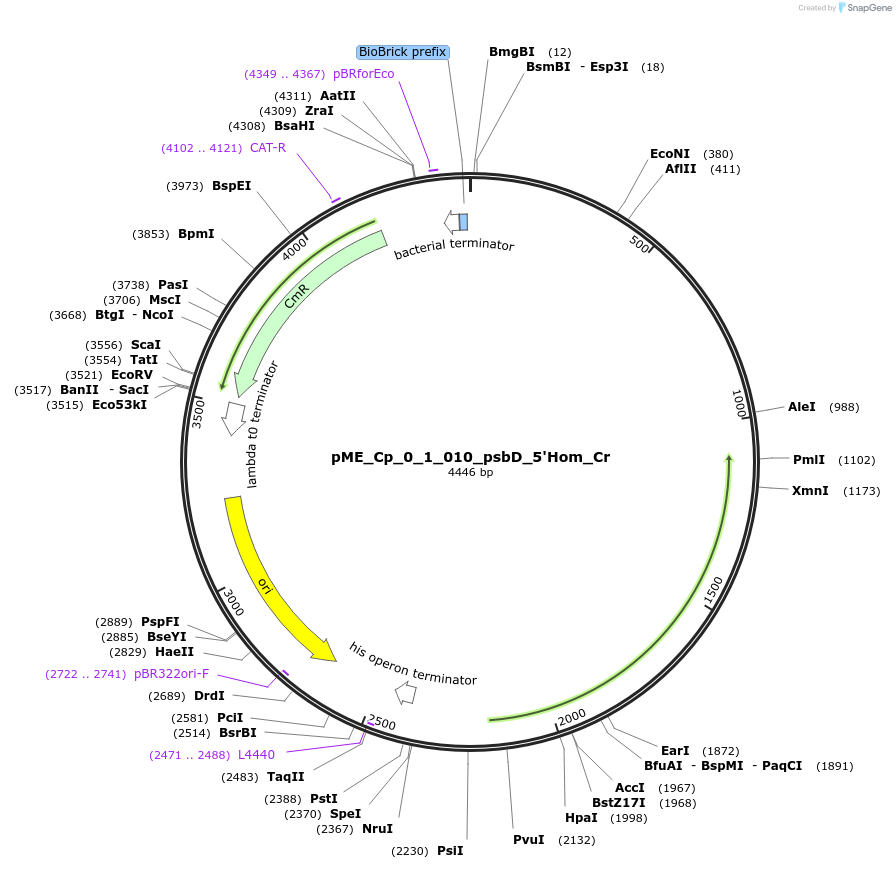
pME_Cp_0_1_010_psbD_5'Hom_Cr
Plasmid#235788Purpose5' Homology psbD in the M1 position used for genome integration at the psbD locusDepositorInsert5' Homology region of the psbD locus of Chlamydomonas reinhardtii chloroplast CC-125
UseSynthetic BiologyMutationnoneAvailable SinceJune 12, 2025AvailabilityAcademic Institutions and Nonprofits only -
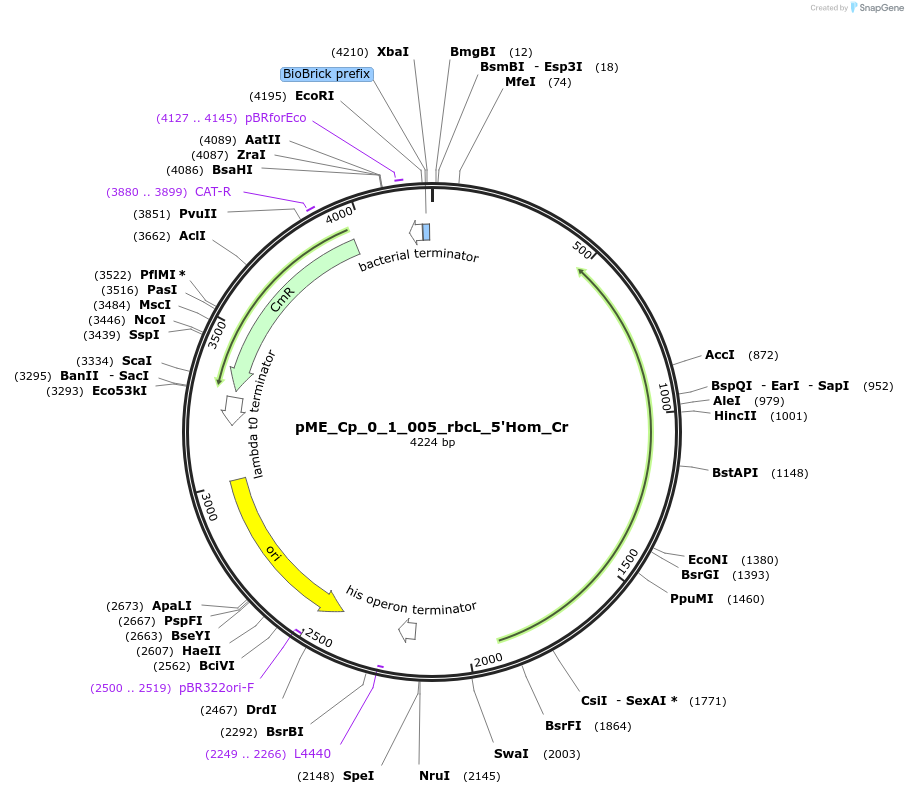
pME_Cp_0_1_005_rbcL_5'Hom_Cr
Plasmid#235786Purpose5' Homology rbcL in the M1 position used for genome integration at the rbcL locusDepositorInsert5' Homology region of the rbcL locus of Chlamydomonas reinhardtii chloroplast CC-125
UseSynthetic BiologyMutationnoneAvailable SinceJune 12, 2025AvailabilityAcademic Institutions and Nonprofits only -
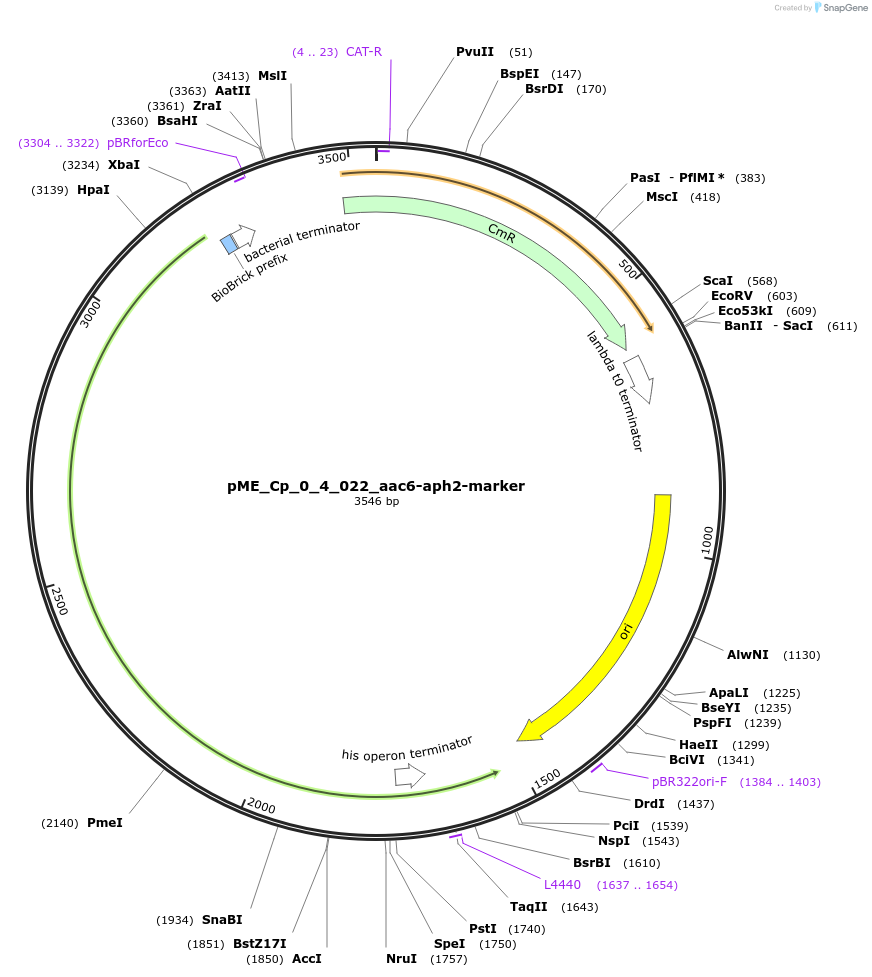
pME_Cp_0_4_022_aac6-aph2-marker
Plasmid#235898Purposeaac6-aph2 tobramycin resistance gene, codon optimized for Chlamydomonas reinhardtiiDepositorInsertaac6-aph2-marker
UseSynthetic BiologyAvailable SinceMay 5, 2025AvailabilityAcademic Institutions and Nonprofits only -
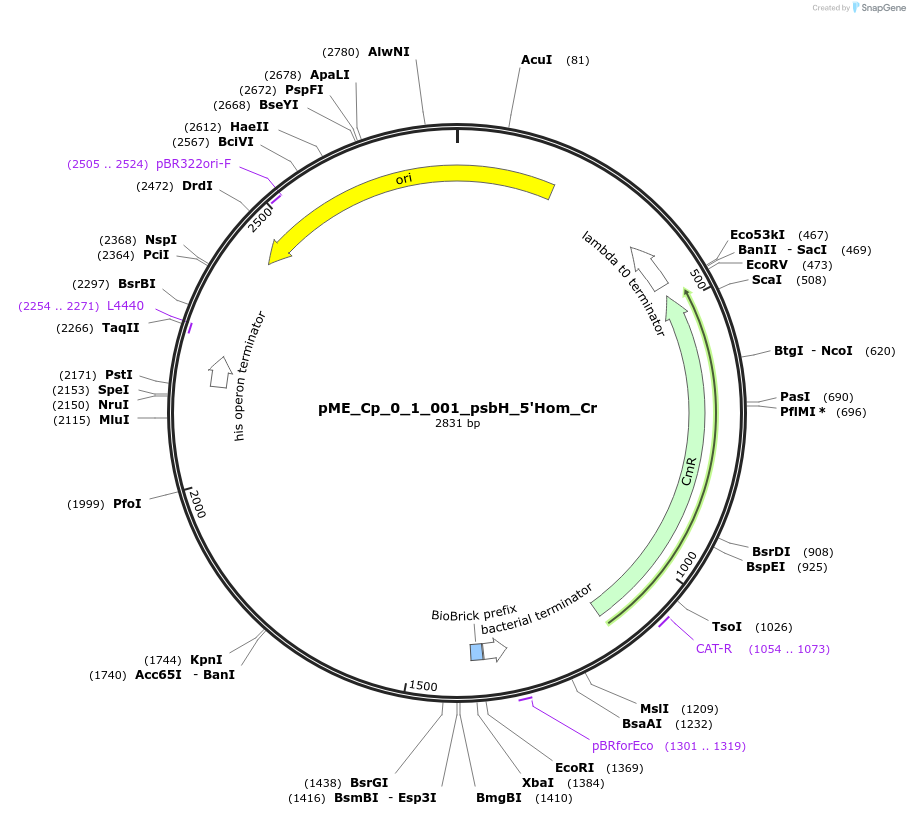
pME_Cp_0_1_001_psbH_5'Hom_Cr
Plasmid#235784Purpose5' Homology psbH in the M1 position used for genome integration at the psbH locusDepositorInsert5' Homology region of the psbH locus of Chlamydomonas reinhardtii chloroplast CC-125
UseSynthetic BiologyMutationnoneAvailable SinceMay 5, 2025AvailabilityAcademic Institutions and Nonprofits only -
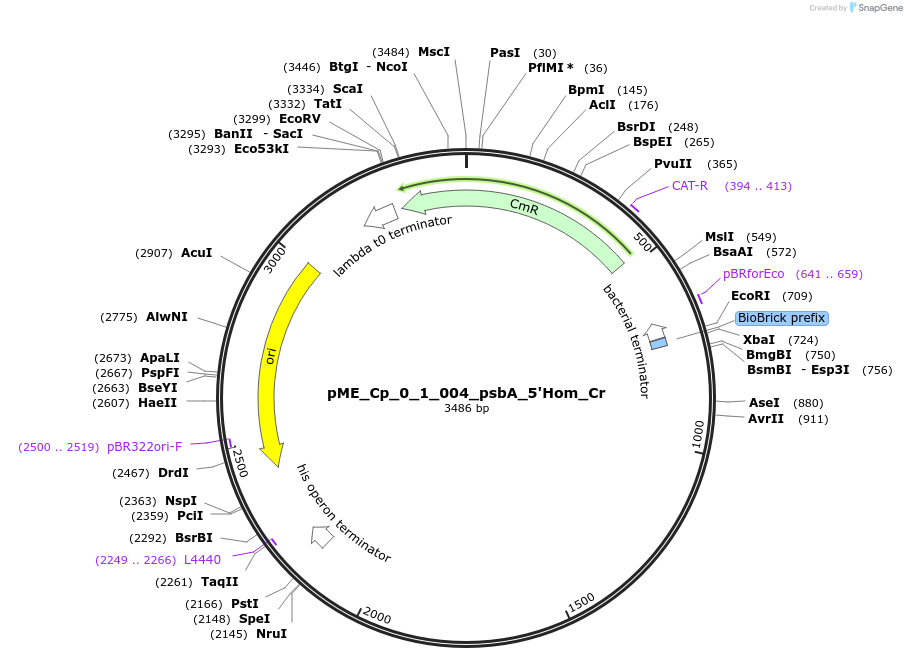
pME_Cp_0_1_004_psbA_5'Hom_Cr
Plasmid#235785Purpose5' Homology psbA in the M1 position used for genome integration at the psbA locusDepositorInsert5' Homology region of the psbA locus of Chlamydomonas reinhardtii chloroplast CC-125
UseSynthetic BiologyMutationnoneAvailable SinceMay 5, 2025AvailabilityAcademic Institutions and Nonprofits only -
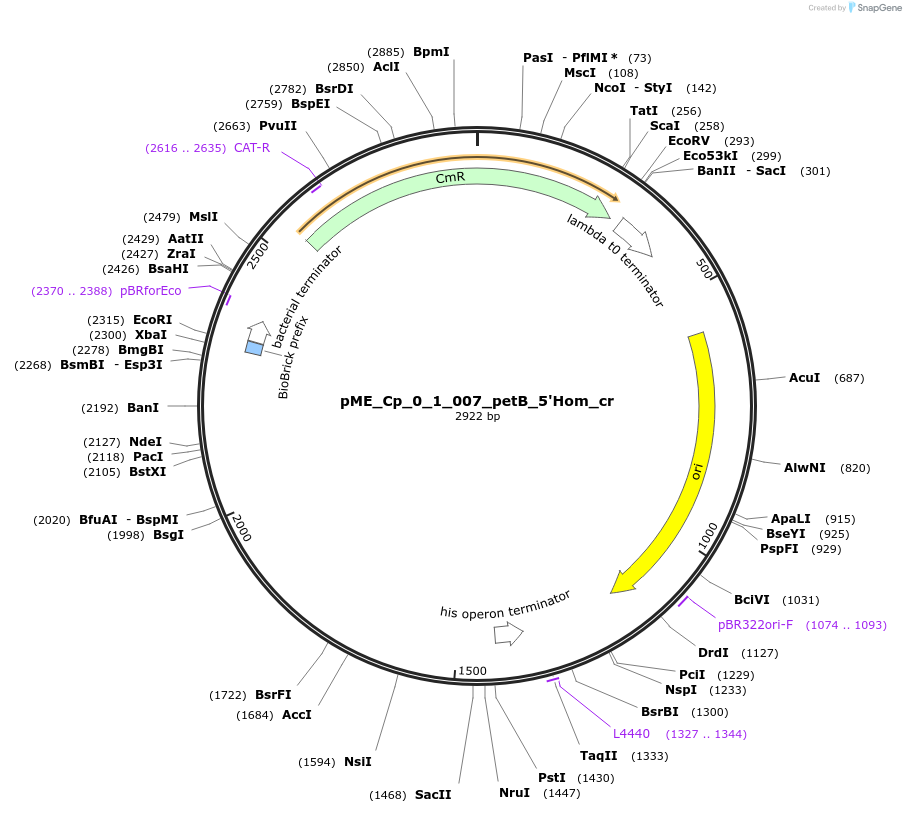
pME_Cp_0_1_007_petB_5'Hom_cr
Plasmid#235787Purpose5' Homology petB in the M1 position used for genome integration at the petB locusDepositorInsert5' Homology region of the petB locus of Chlamydomonas reinhardtii chloroplast CC-125
UseSynthetic BiologyMutationnoneAvailable SinceMay 5, 2025AvailabilityAcademic Institutions and Nonprofits only -
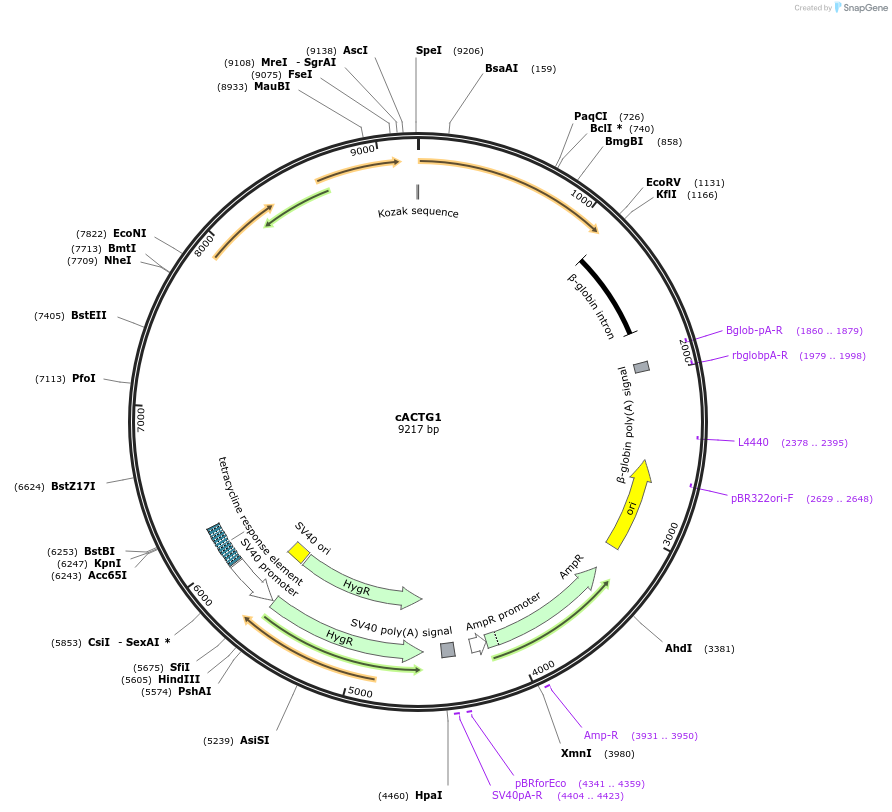
cACTG1
Plasmid#236543PurposeFor expression of ACTG1 in mammalian cellsDepositorInsertACTG1
ExpressionMammalianMutationWTPromoterhpACTBAvailable SinceApril 23, 2025AvailabilityAcademic Institutions and Nonprofits only