We narrowed to 3,157 results for: bad
-
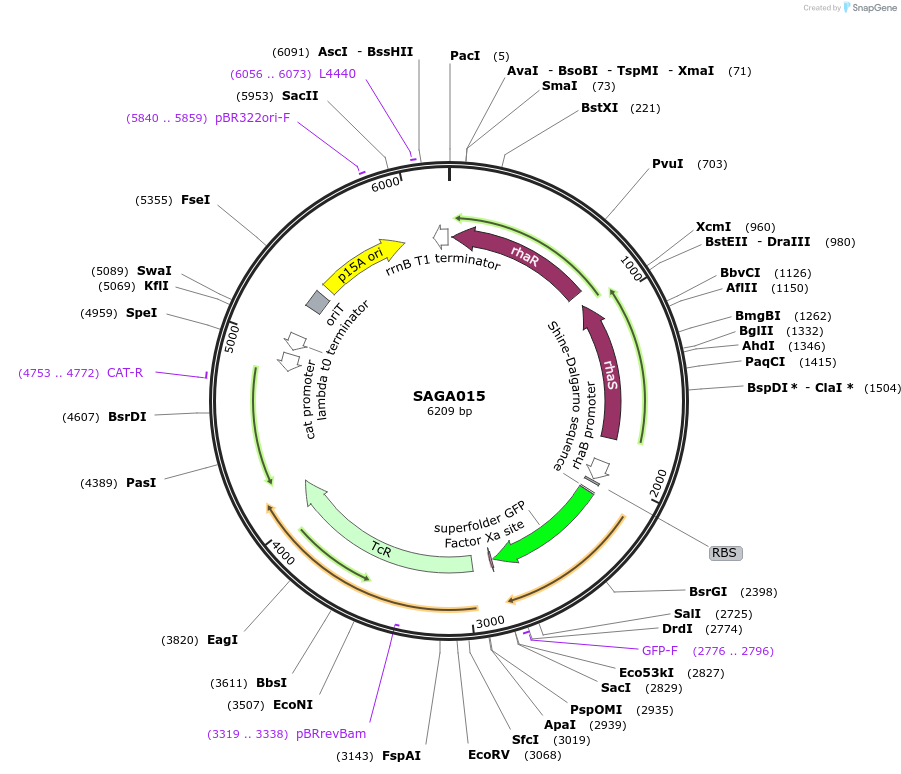
Plasmid#240767PurposePlasmid contains the triple selection cassette gfp - tetApBR322-ROP- CmTrunc that allows for in vivo recombineering in E.coli.DepositorInsertSAGA pBR322-ROP PrhaBAD BCD_low - gfp - tetApBR322-ROP- CmTrunc
UseSynthetic BiologyMutationMutation on translation initiation region of tetAAvailable SinceAug. 20, 2025AvailabilityAcademic Institutions and Nonprofits only -
SAGA015
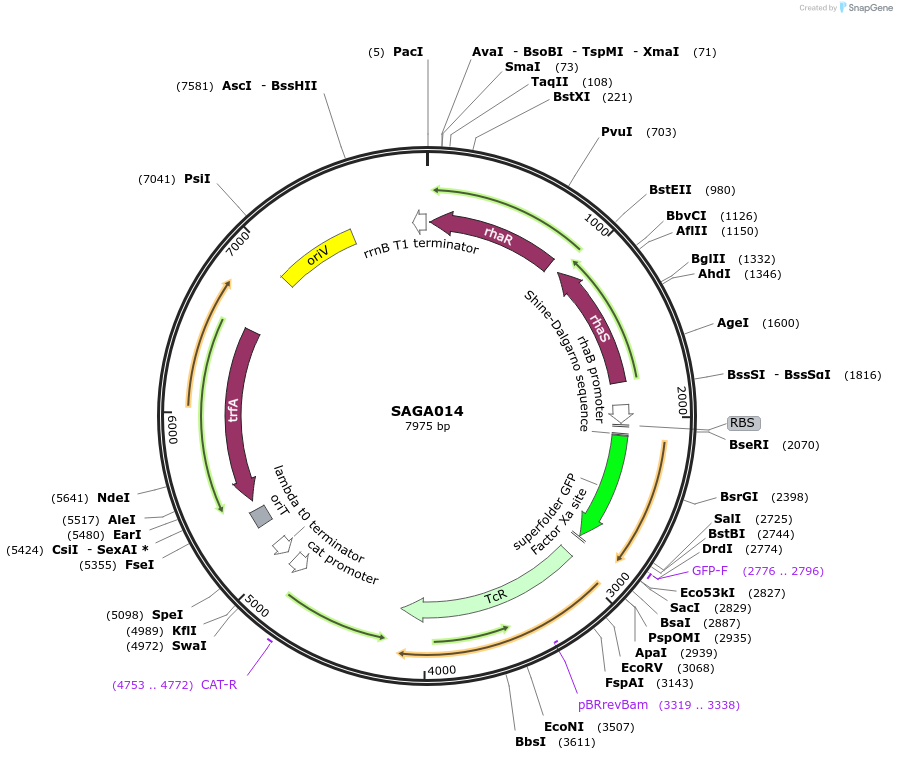
Plasmid#240764PurposePlasmid contains the triple selection cassette gfp - tetAp15A- CmTrunc that allows for in vivo recombineering in E.coli.DepositorInsertSAGA p15A PrhaBAD BCD_low - gfp - tetAp15A- CmTrunc
UseSynthetic BiologyMutationMutation on translation initiation region of tetAAvailable SinceAug. 20, 2025AvailabilityAcademic Institutions and Nonprofits only -
SAGA014
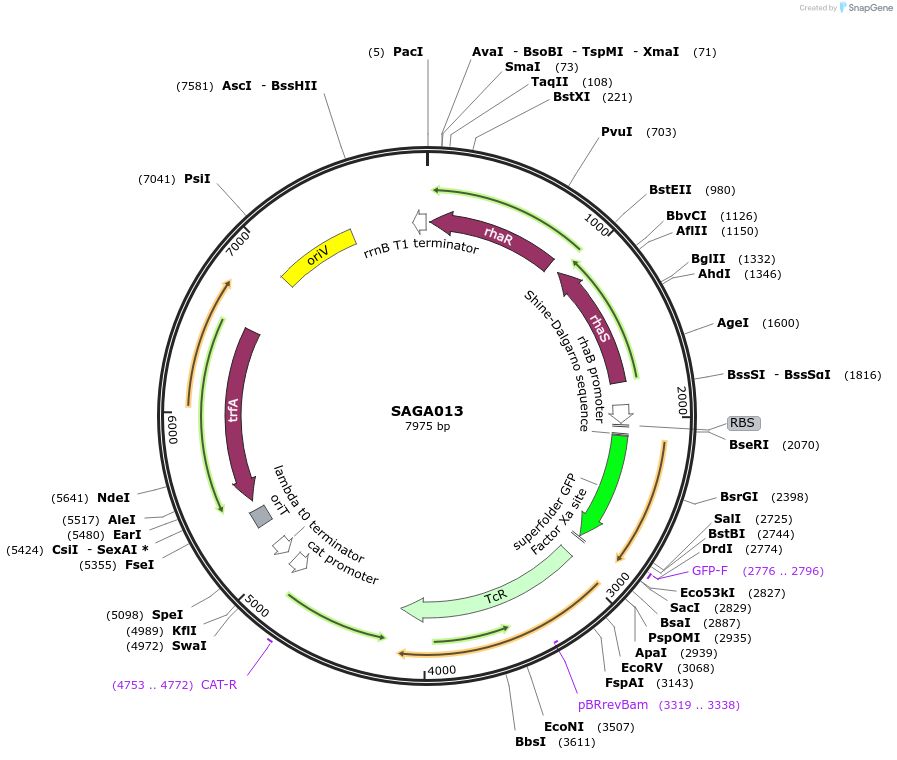
Plasmid#240763PurposePlasmid contains the triple selection cassette gfp - tetARK2- CmTrunc that allows for in vivo recombineering in E.coli.DepositorInsertSAGA RK2 PrhaBAD BCD_high - gfp - tetARK2- CmTrunc
UseSynthetic BiologyMutationMutation on translation initiation region of tetAAvailable SinceAug. 20, 2025AvailabilityAcademic Institutions and Nonprofits only -
SAGA013
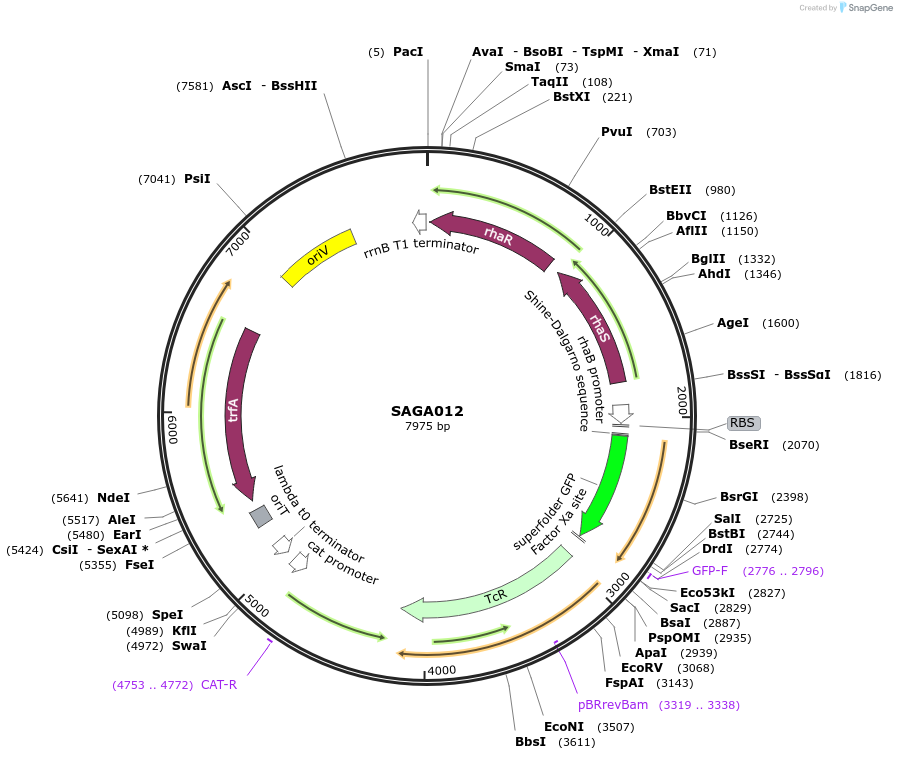
Plasmid#240762PurposePlasmid contains the triple selection cassette gfp - tetARK2- CmTrunc that allows for in vivo recombineering in E.coli.DepositorInsertSAGA RK2 PrhaBAD BCD_medium - gfp - tetARK2- CmTrunc
UseSynthetic BiologyMutationMutation on translation initiation region of tetAAvailable SinceAug. 20, 2025AvailabilityAcademic Institutions and Nonprofits only -
SAGA012
Plasmid#240761PurposePlasmid contains the triple selection cassette gfp - tetARK2- CmTrunc that allows for in vivo recombineering in E.coli.DepositorInsertSAGA RK2 PrhaBAD BCD_low - gfp - tetARK2- CmTrunc
UseSynthetic BiologyMutationMutation on translation initiation region of tetAAvailable SinceAug. 20, 2025AvailabilityAcademic Institutions and Nonprofits only -
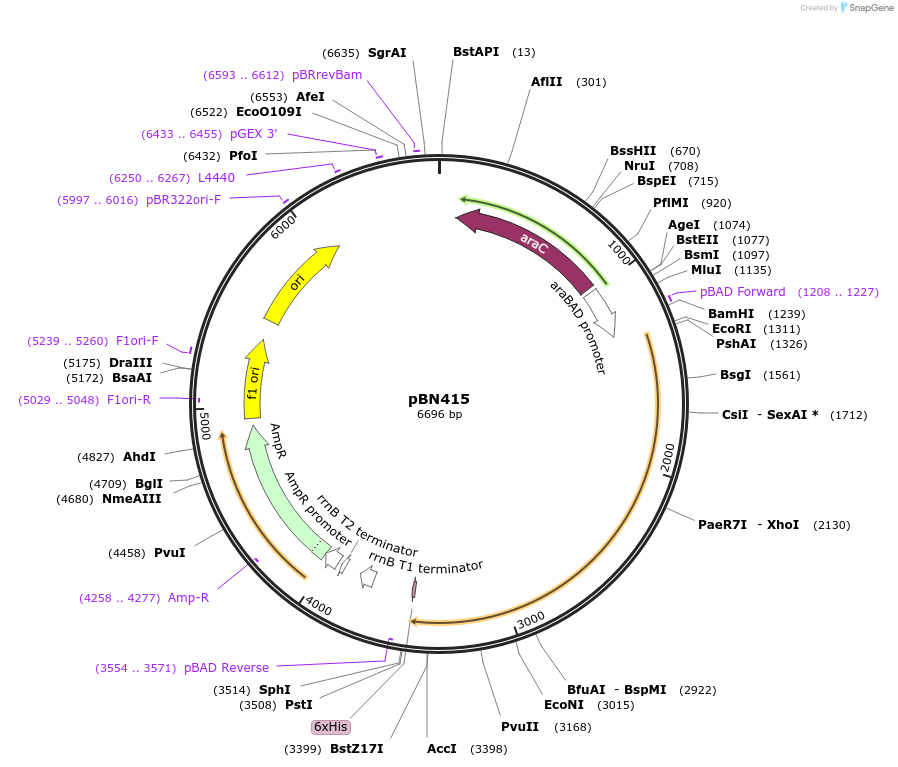
pBN415
Plasmid#217920Purposeexpress E. coli E69 (O9a:K30) Wzc with N711Y mutationDepositorInsertE. coli E69 (O9a:K30) Wzc with N711Y mutation
Tagshexa-histidine tagExpressionBacterialMutationN711YAvailable SinceJuly 10, 2025AvailabilityAcademic Institutions and Nonprofits only -
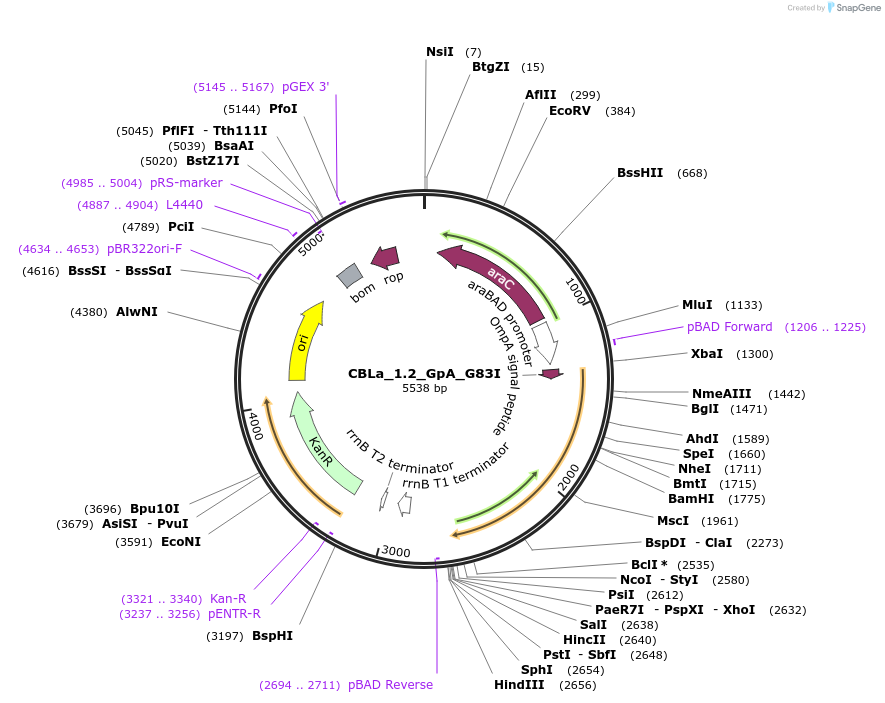
CBLa_1.2_GpA_G83I
Plasmid#207845PurposeBLaTM-System GpA G83I negative control in CBLa 1.2 vectorDepositorInsertN-BLa 1.2 fusion protein
TagsFlagExpressionBacterialAvailable SinceDec. 21, 2023AvailabilityAcademic Institutions and Nonprofits only -
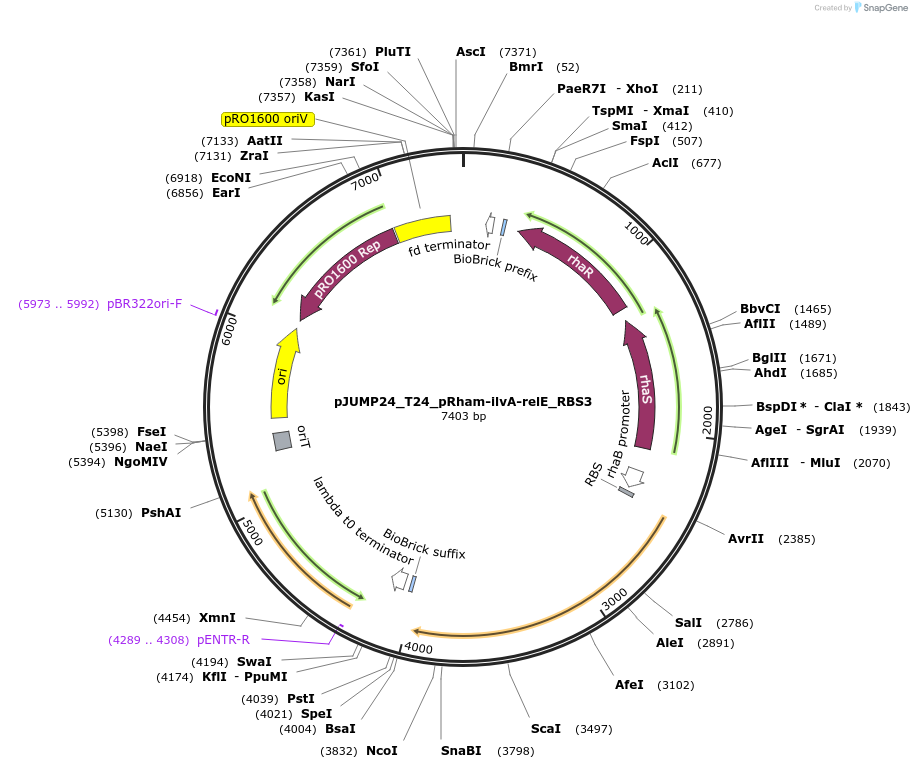
pJUMP24_T24_pRham-ilvA-relE_RBS3
Plasmid#201532PurposeExpressing synthetic overlapping gene, ilvA-relE with internal RBS modification. Origin pRO1600/ColE1 (E.coli - Pseudomonas shuttle)DepositorInsertilvA-relE_RBS3
UseSynthetic BiologyPromoterPrhaBAD (rhamnose)Available SinceJune 23, 2023AvailabilityAcademic Institutions and Nonprofits only -
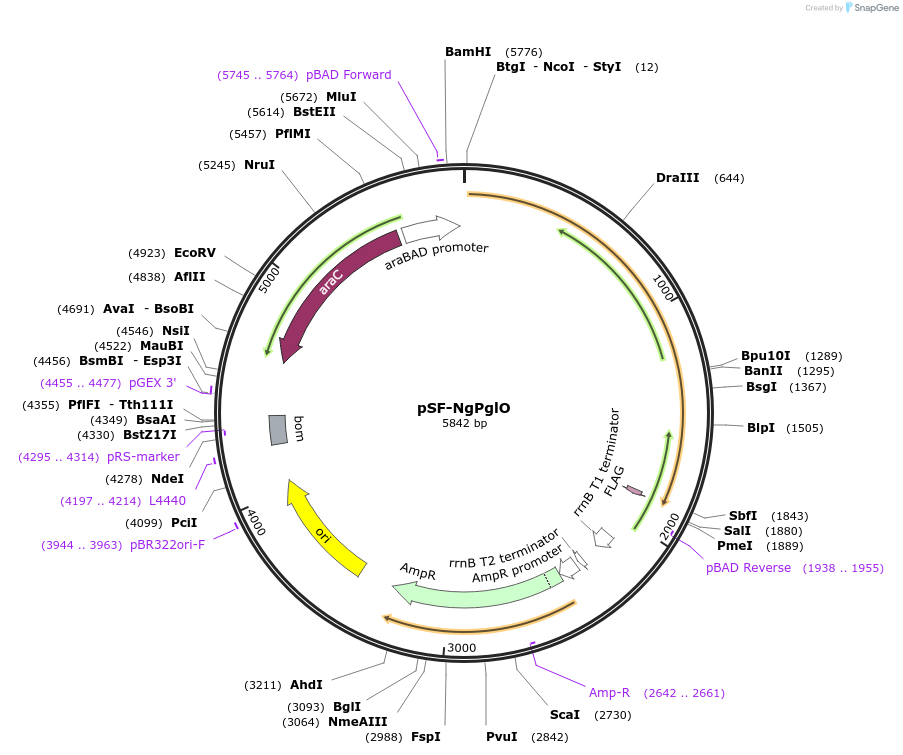
pSF-NgPglO
Plasmid#198204PurposepSN18 derivative encoding N. gonorrhoeae PglO with C-terminal FLAG epitope tagDepositorInsertNgPglO
TagsFlag tagExpressionBacterialPromoterpBADAvailable SinceMay 26, 2023AvailabilityAcademic Institutions and Nonprofits only -
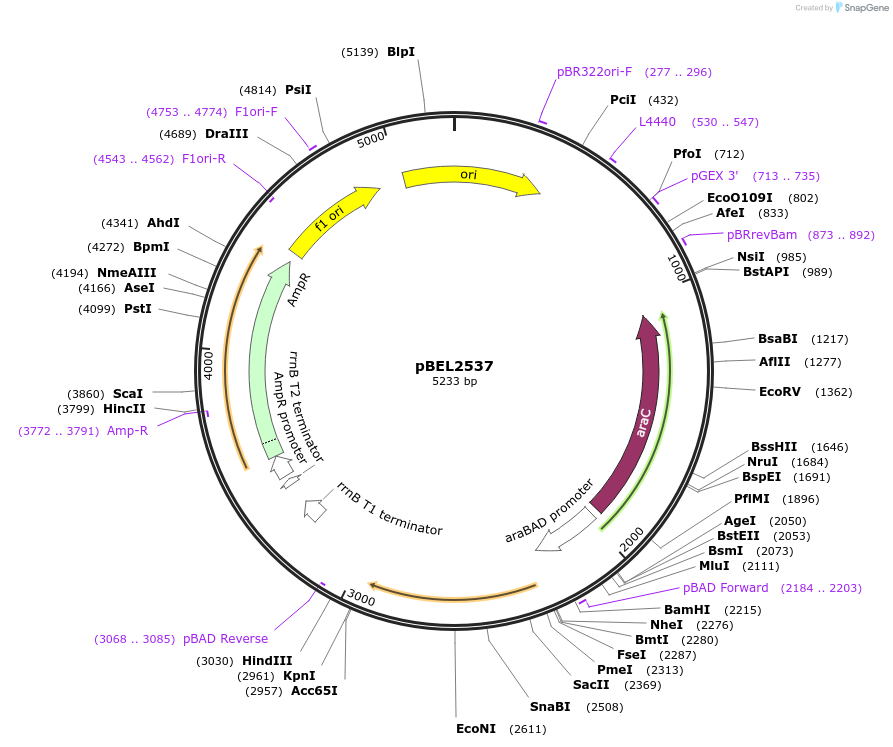
pBEL2537
Plasmid#195658PurposeComplementation of MlaC(R153A)DepositorInsertMlaC
ExpressionBacterialMutationMlaC(R153A)Available SinceMay 2, 2023AvailabilityAcademic Institutions and Nonprofits only -
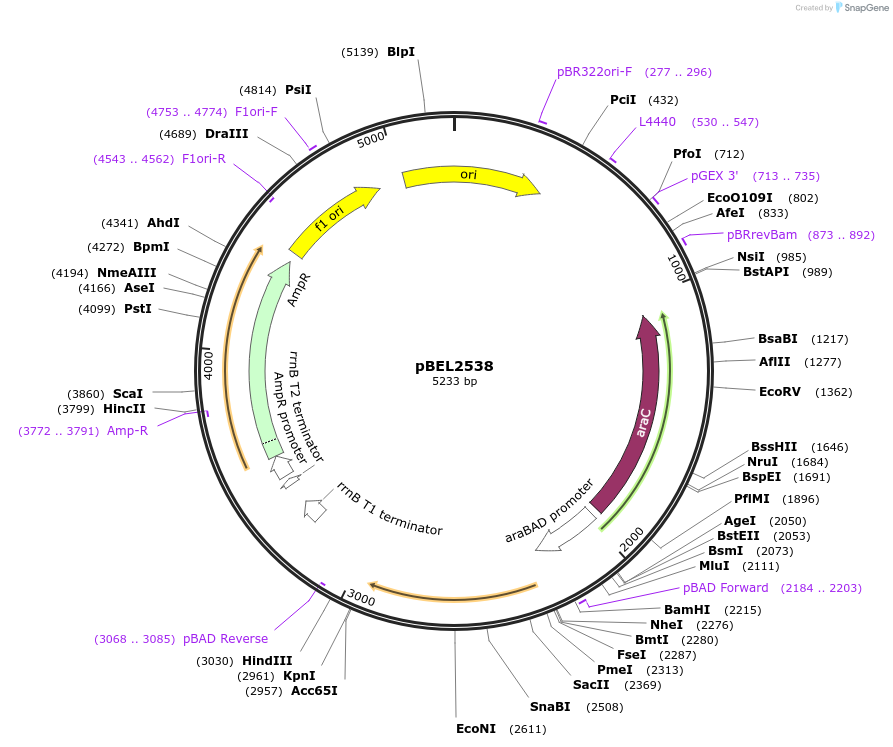
pBEL2538
Plasmid#195659PurposeComplementation of MlaC(R186A)DepositorInsertMlaC
ExpressionBacterialMutationMlaC(R186A)Available SinceMay 2, 2023AvailabilityAcademic Institutions and Nonprofits only -
pBEL2539
Plasmid#195660PurposeComplementation of MlaC(T116D)DepositorInsertMlaC
ExpressionBacterialMutationMlaC(T116D)Available SinceMay 2, 2023AvailabilityAcademic Institutions and Nonprofits only -
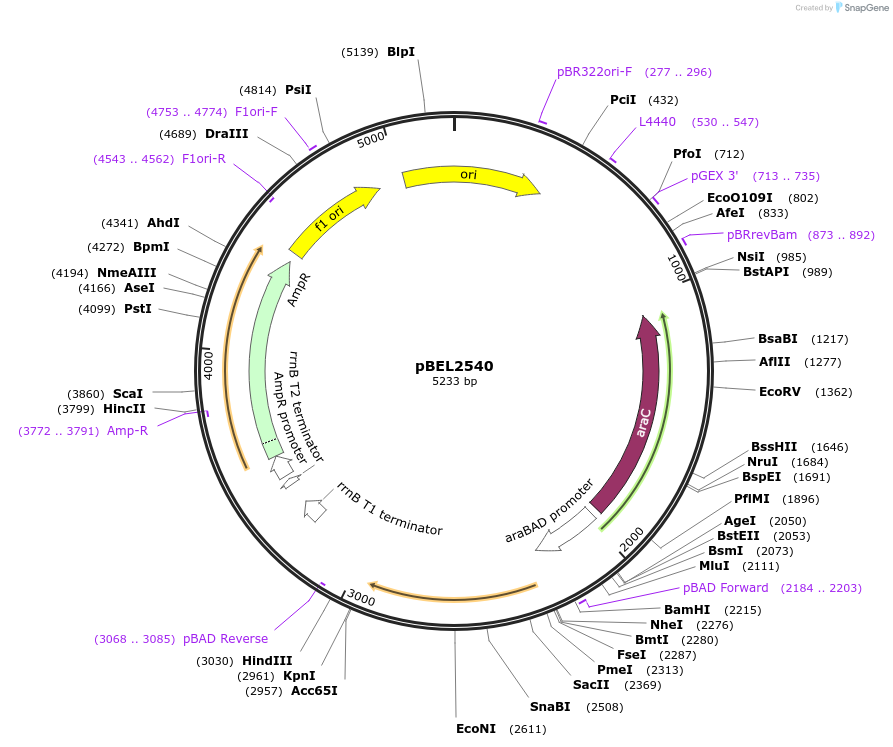
pBEL2540
Plasmid#195661PurposeComplementation of MlaC(T176R)DepositorInsertMlaC
ExpressionBacterialMutationMlaC(T176R)Available SinceMay 2, 2023AvailabilityAcademic Institutions and Nonprofits only -
pBEL2541
Plasmid#195662PurposeComplementation of MlaC(V68R)DepositorInsertMlaC
ExpressionBacterialMutationMlaC(V68R)Available SinceMay 2, 2023AvailabilityAcademic Institutions and Nonprofits only -
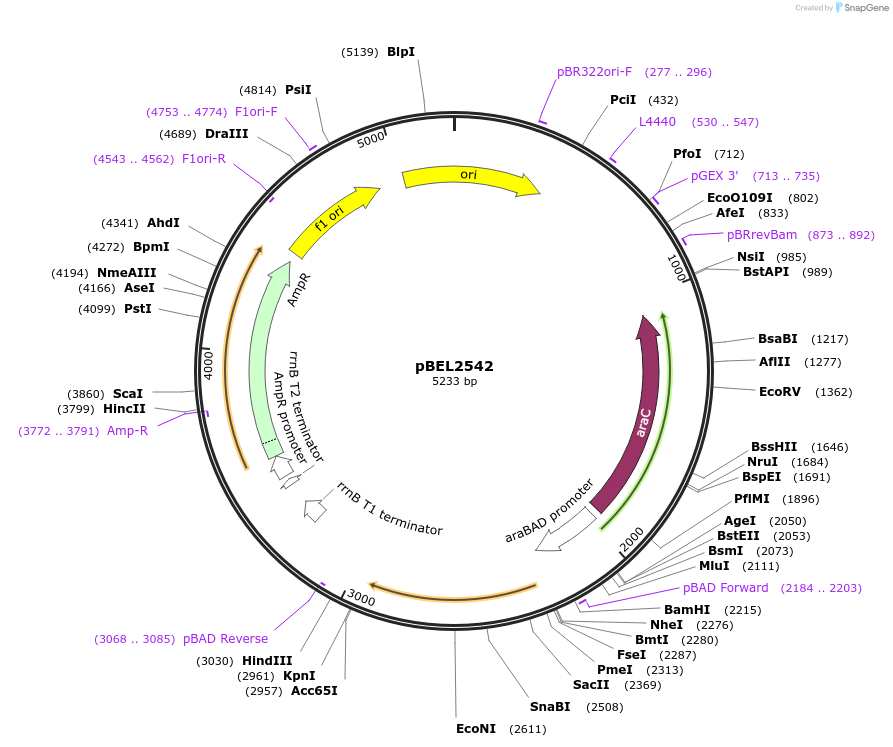
pBEL2542
Plasmid#195663PurposeComplementation of MlaC(V135R)DepositorInsertMlaC
ExpressionBacterialMutationMlaC(V135R)Available SinceMay 2, 2023AvailabilityAcademic Institutions and Nonprofits only -
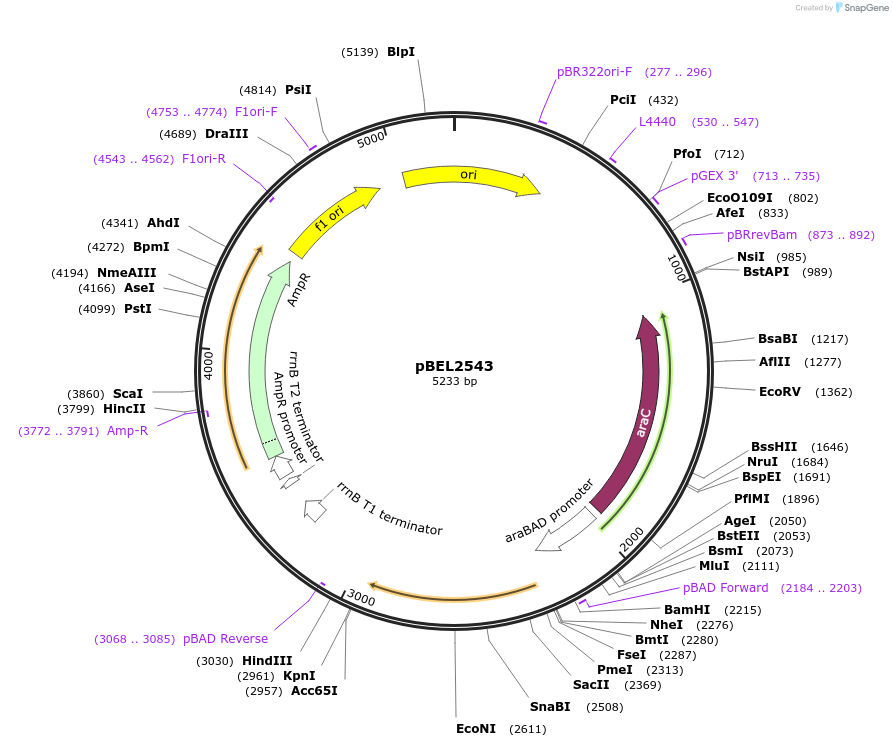
pBEL2543
Plasmid#195664PurposeComplementation of MlaC(V146R)DepositorInsertMlaC
ExpressionBacterialMutationMlaC(V146R)Available SinceMay 2, 2023AvailabilityAcademic Institutions and Nonprofits only -
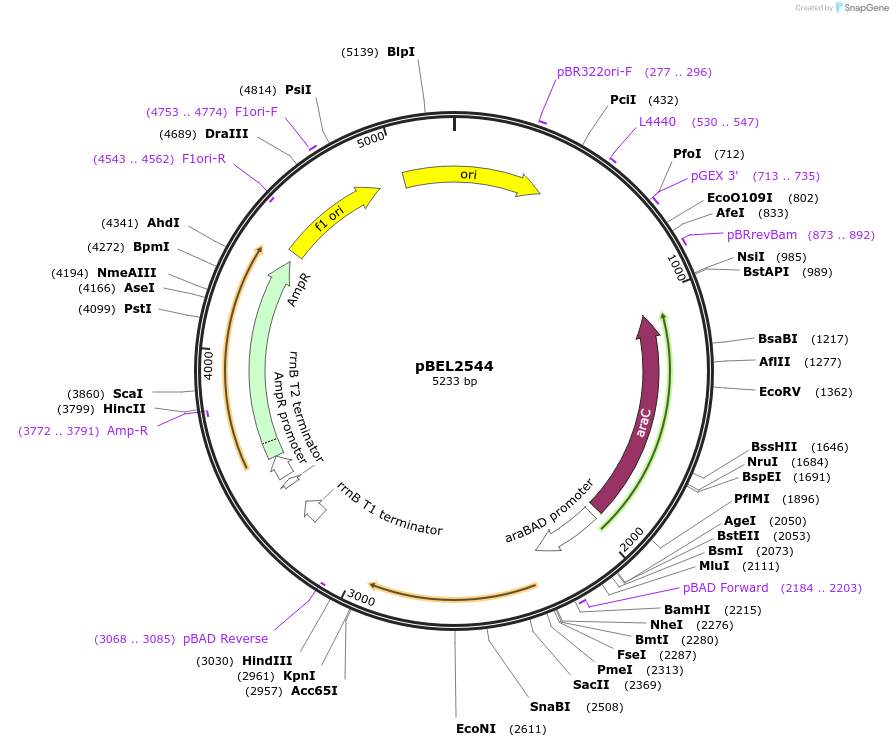
pBEL2544
Plasmid#195665PurposeComplementation of MlaC(V171R)DepositorInsertMlaC
ExpressionBacterialMutationMlaC(V171R)Available SinceMay 2, 2023AvailabilityAcademic Institutions and Nonprofits only -
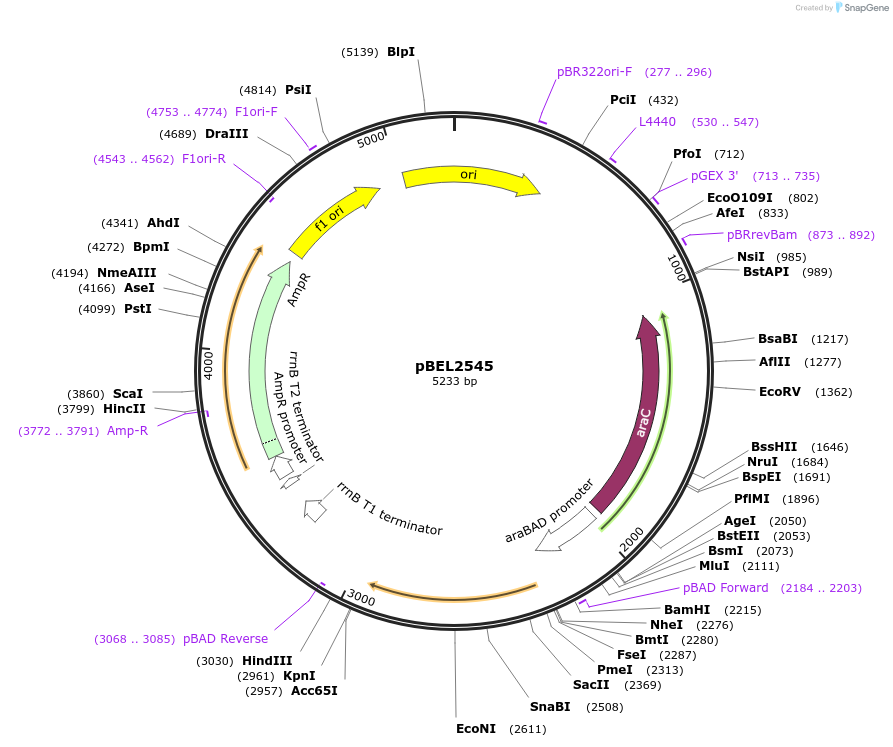
pBEL2545
Plasmid#195666PurposeComplementation of MlaC(Y72A)DepositorInsertMlaC
ExpressionBacterialMutationMlaC(Y72A)Available SinceMay 2, 2023AvailabilityAcademic Institutions and Nonprofits only -
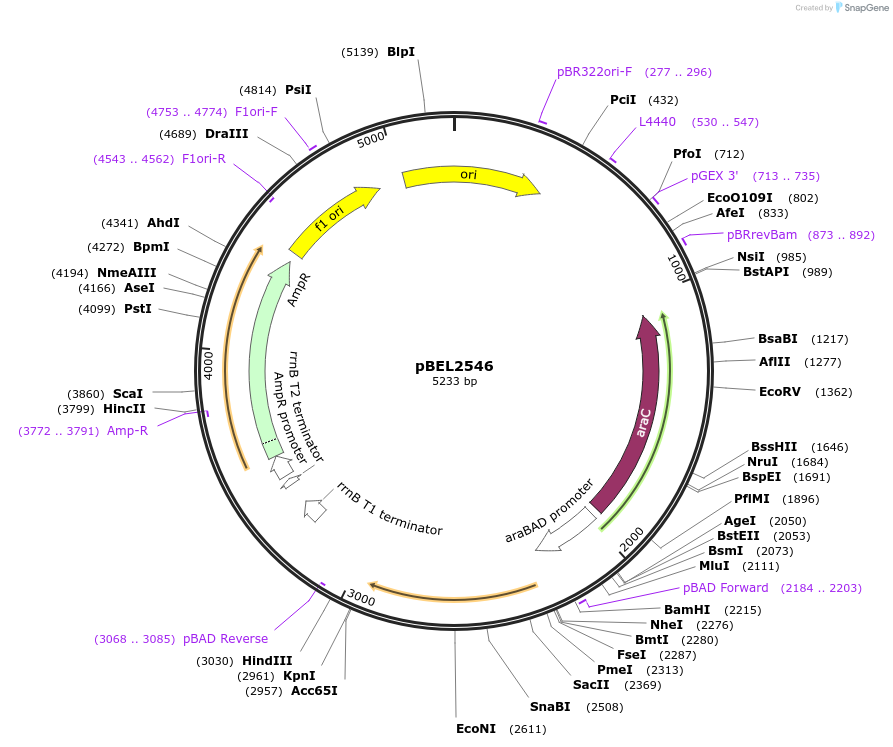
pBEL2546
Plasmid#195667PurposeComplementation of MlaC(Y82K)DepositorInsertMlaC
ExpressionBacterialMutationMlaC(Y82K)Available SinceMay 2, 2023AvailabilityAcademic Institutions and Nonprofits only -
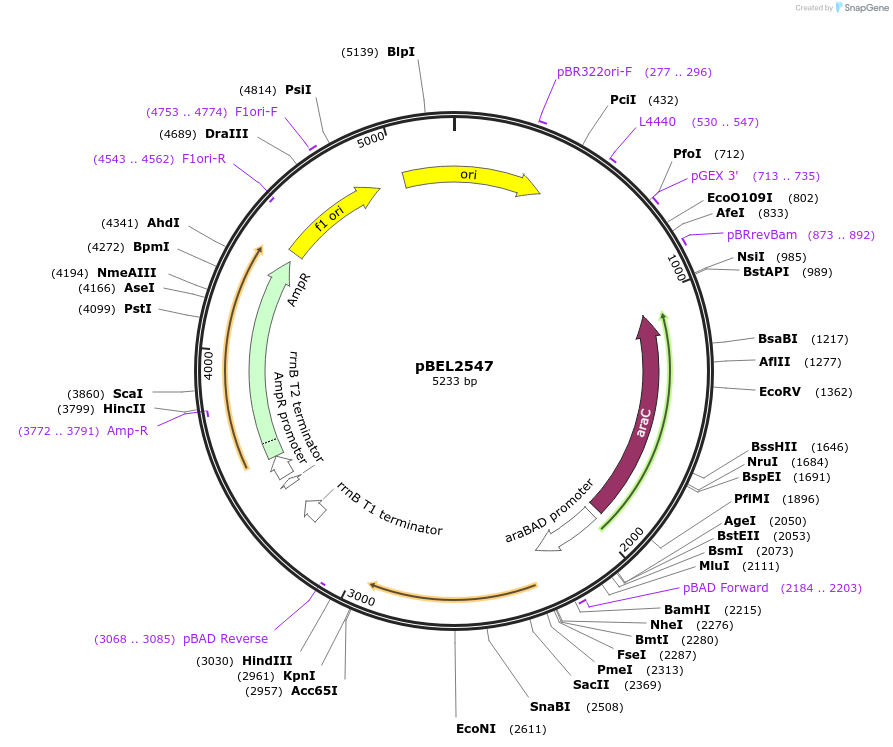
pBEL2547
Plasmid#195668PurposeComplementation of MlaC(Y105A)DepositorInsertMlaC
ExpressionBacterialMutationMlaC(Y105A)Available SinceMay 2, 2023AvailabilityAcademic Institutions and Nonprofits only -
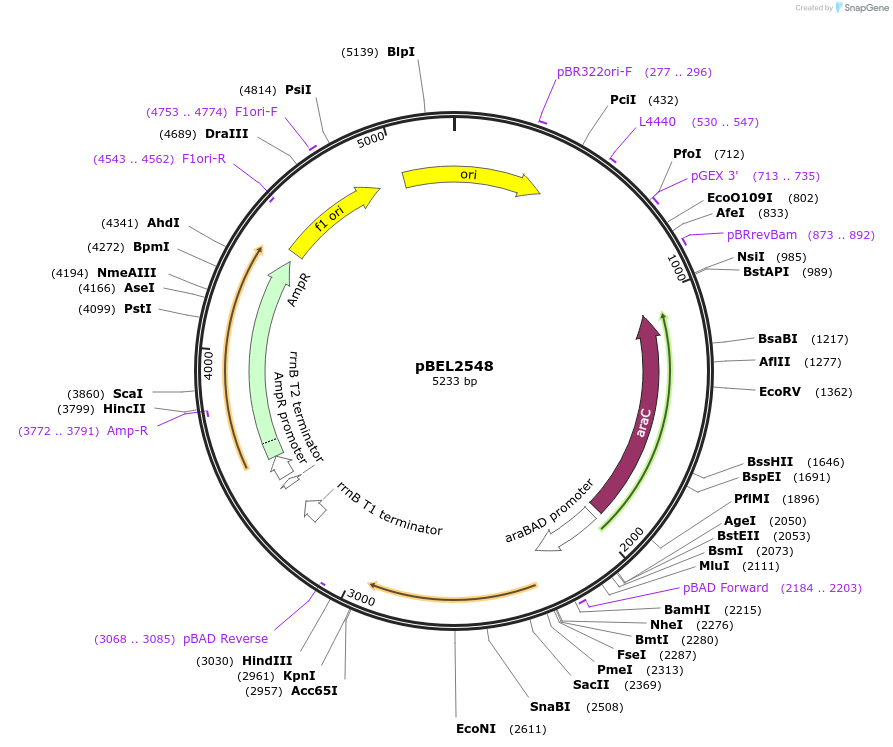
pBEL2548
Plasmid#195669PurposeComplementation of MlaC(Y164R)DepositorInsertMlaC
ExpressionBacterialMutationMlaC(Y164R)Available SinceMay 2, 2023AvailabilityAcademic Institutions and Nonprofits only -
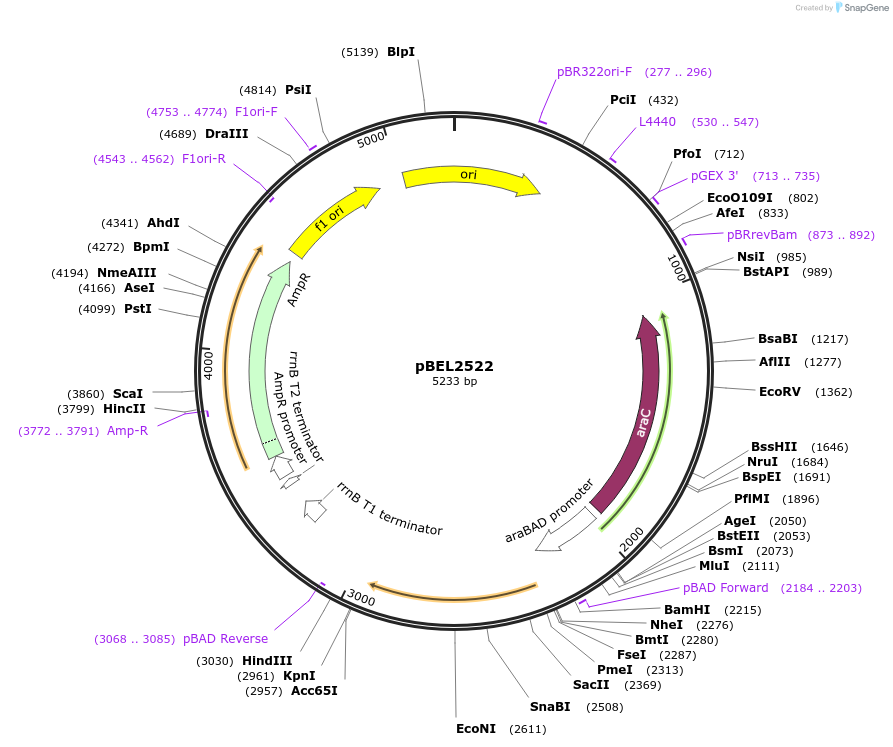
pBEL2522
Plasmid#195644PurposeComplementation of MlaC(A108R)DepositorInsertMlaC
ExpressionBacterialMutationMlaC(A108R)Available SinceMay 2, 2023AvailabilityAcademic Institutions and Nonprofits only -
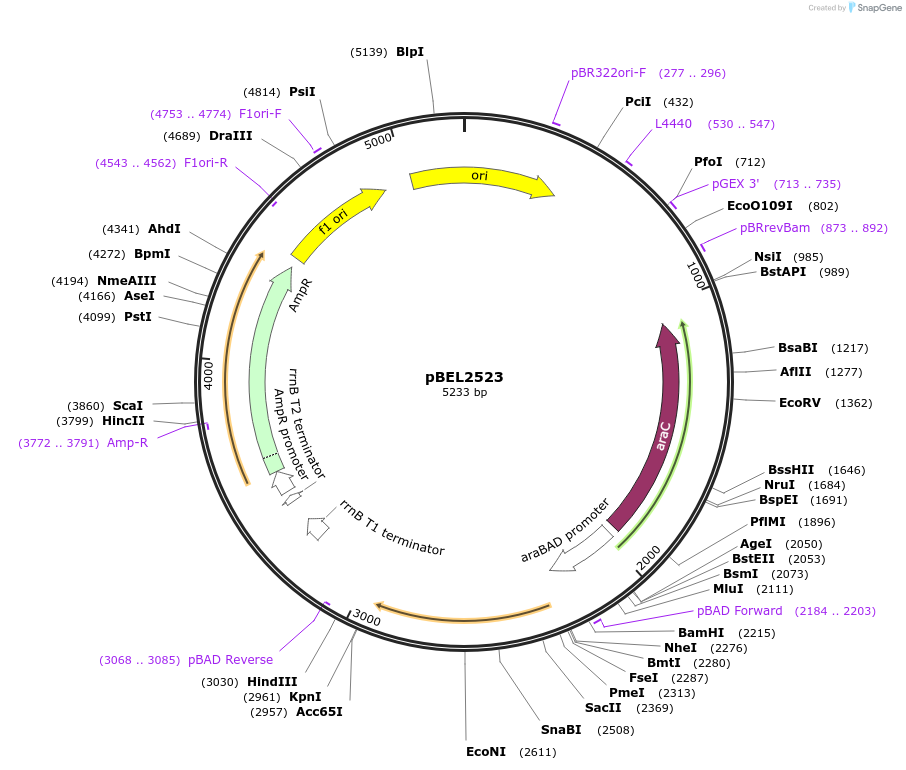
pBEL2523
Plasmid#195645PurposeComplementation of MlaC(A163R)DepositorInsertMlaC
ExpressionBacterialMutationMlaC(A163R)Available SinceMay 2, 2023AvailabilityAcademic Institutions and Nonprofits only -
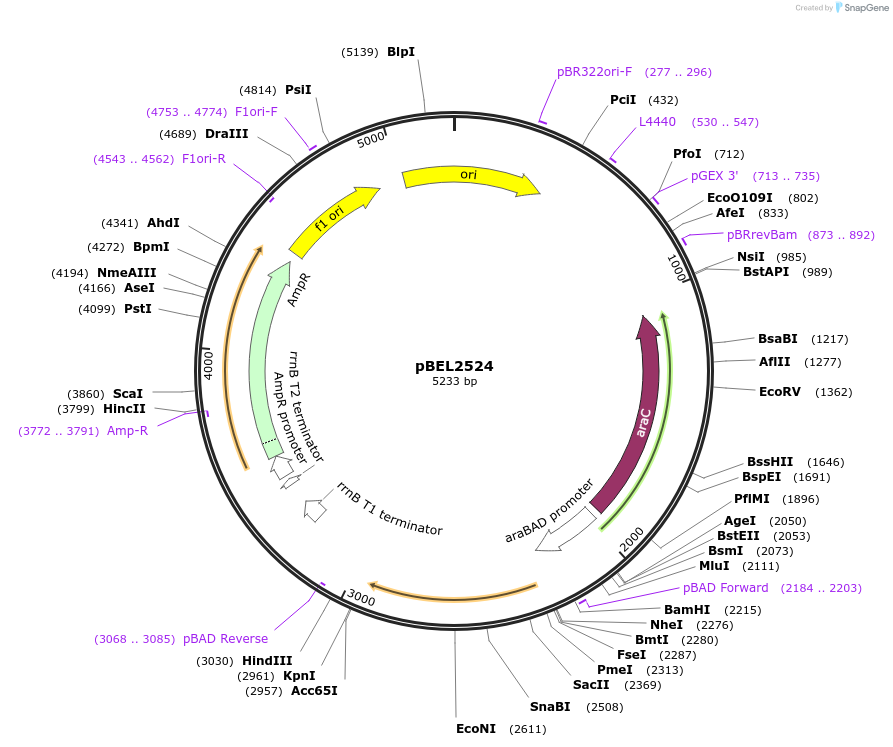
pBEL2524
Plasmid#195646PurposeComplementation of MlaC(A168R)DepositorInsertMlaC
ExpressionBacterialMutationMlaC(A168R)Available SinceMay 2, 2023AvailabilityAcademic Institutions and Nonprofits only -
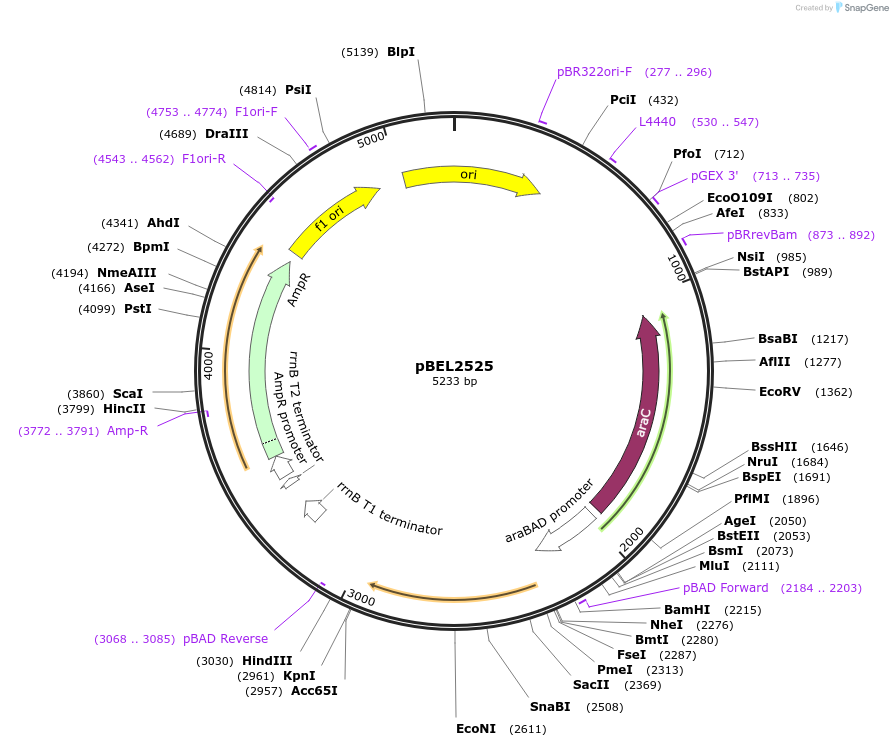
pBEL2525
Plasmid#195647PurposeComplementation of MlaC(D165I)DepositorInsertMlaC
ExpressionBacterialMutationMlaC(D165I)Available SinceMay 2, 2023AvailabilityAcademic Institutions and Nonprofits only -
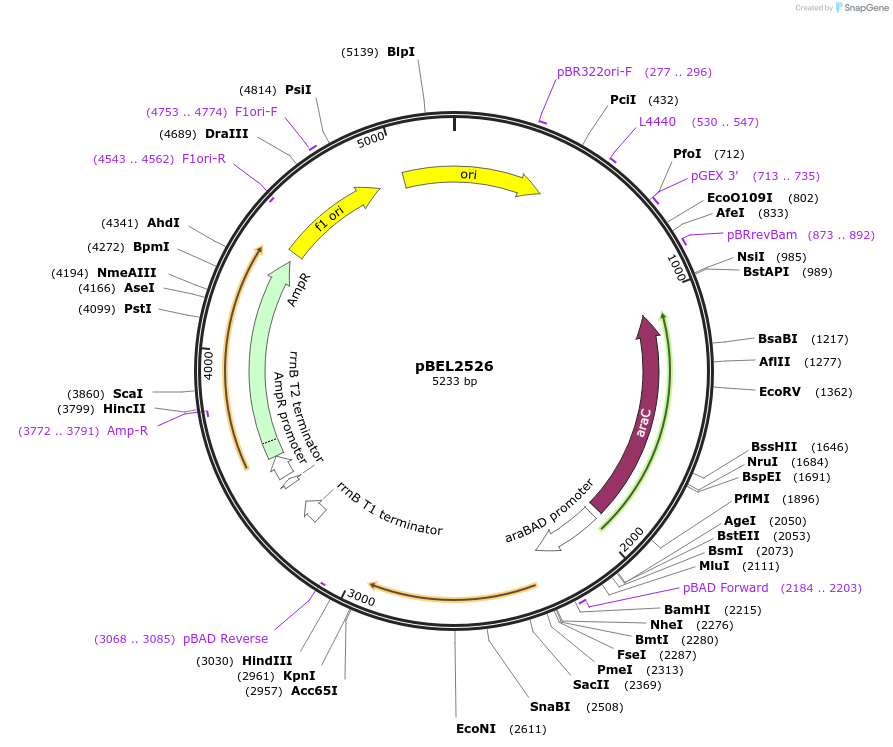
pBEL2526
Plasmid#195648PurposeComplementation of MlaC(E169R)DepositorInsertMlaC
ExpressionBacterialMutationMlaC(E169R)Available SinceMay 2, 2023AvailabilityAcademic Institutions and Nonprofits only -
pBEL2527
Plasmid#195649PurposeComplementation of MlaC(G170K)DepositorInsertMlaC
ExpressionBacterialMutationMlaC(G170K)Available SinceMay 2, 2023AvailabilityAcademic Institutions and Nonprofits only -
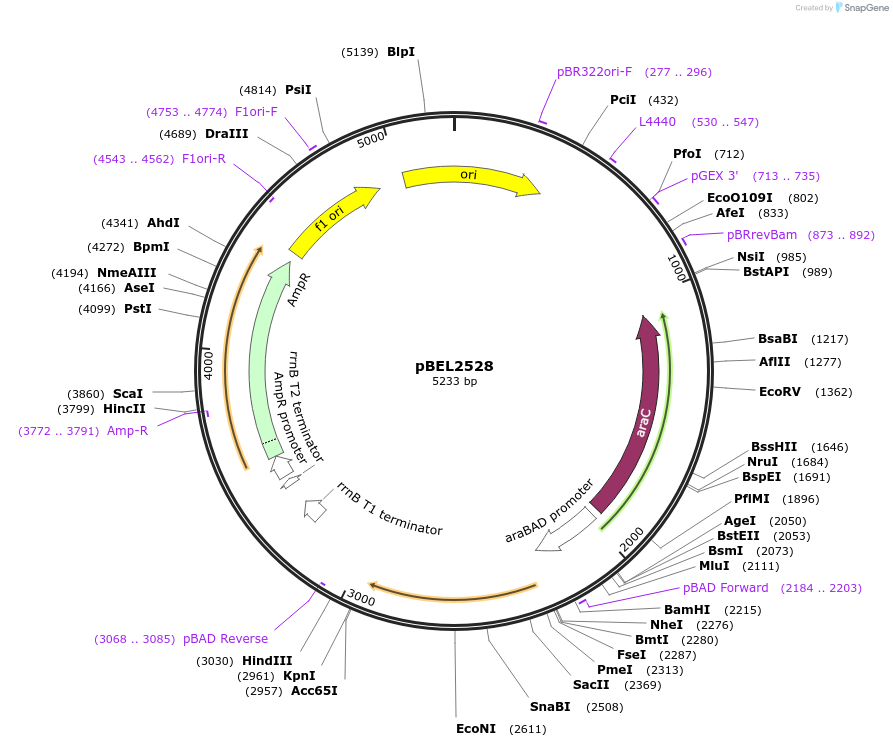
pBEL2528
Plasmid#195650PurposeComplementation of MlaC(I130D)DepositorInsertMlaC
ExpressionBacterialMutationMlaC(I130D)Available SinceMay 2, 2023AvailabilityAcademic Institutions and Nonprofits only -
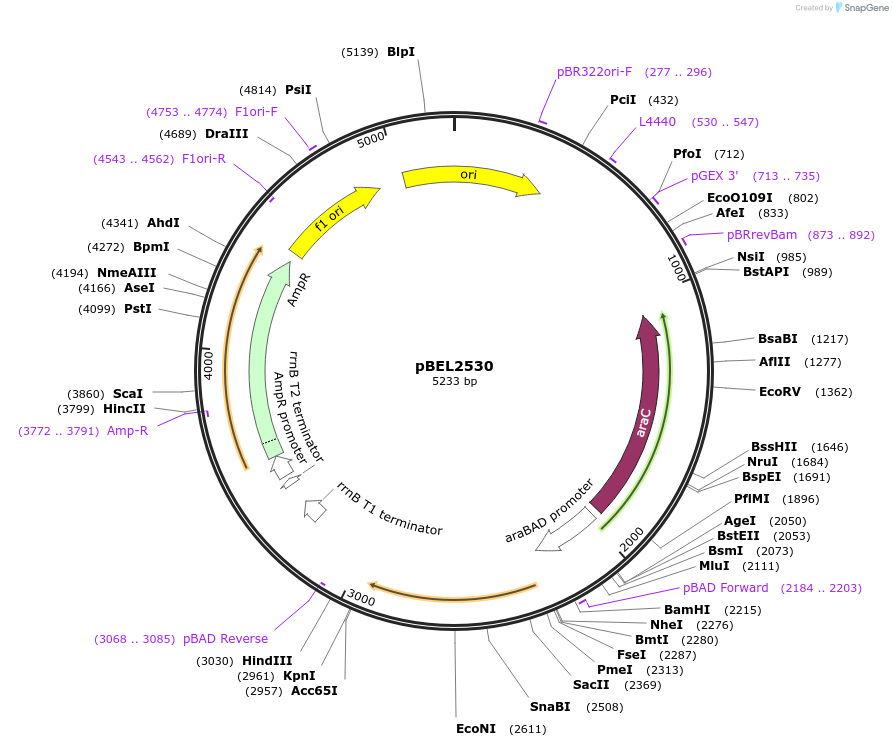
pBEL2530
Plasmid#195651PurposeComplementation of MlaC(I137D)DepositorInsertMlaC
ExpressionBacterialMutationMlaC(I137D)Available SinceMay 2, 2023AvailabilityAcademic Institutions and Nonprofits only -
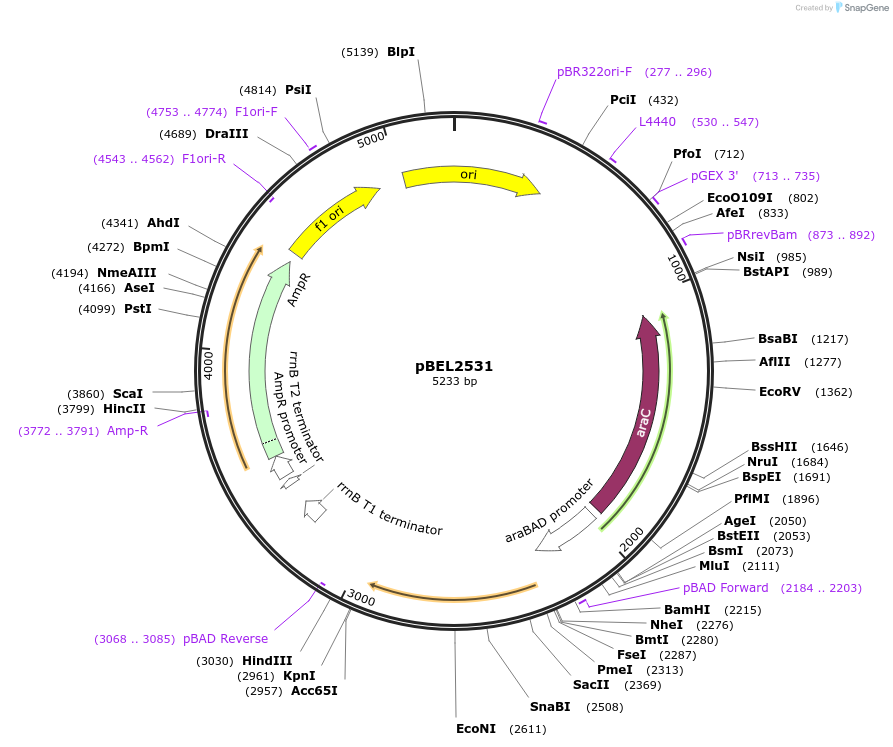
pBEL2531
Plasmid#195652PurposeComplementation of MlaC(L76D)DepositorInsertMlaC
ExpressionBacterialMutationMlaC(L76D)Available SinceMay 2, 2023AvailabilityAcademic Institutions and Nonprofits only