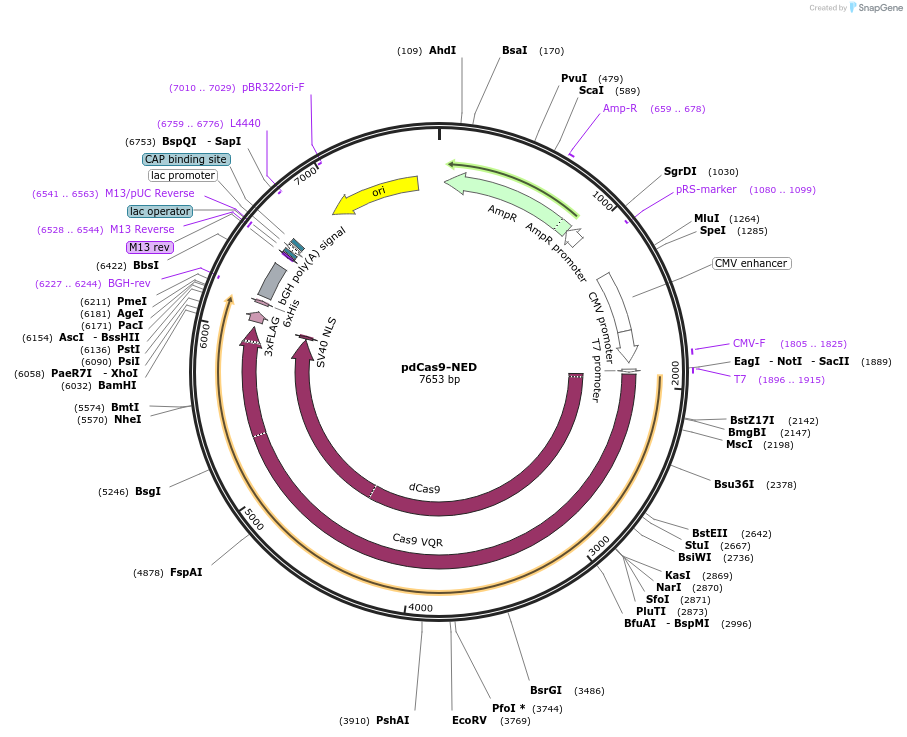
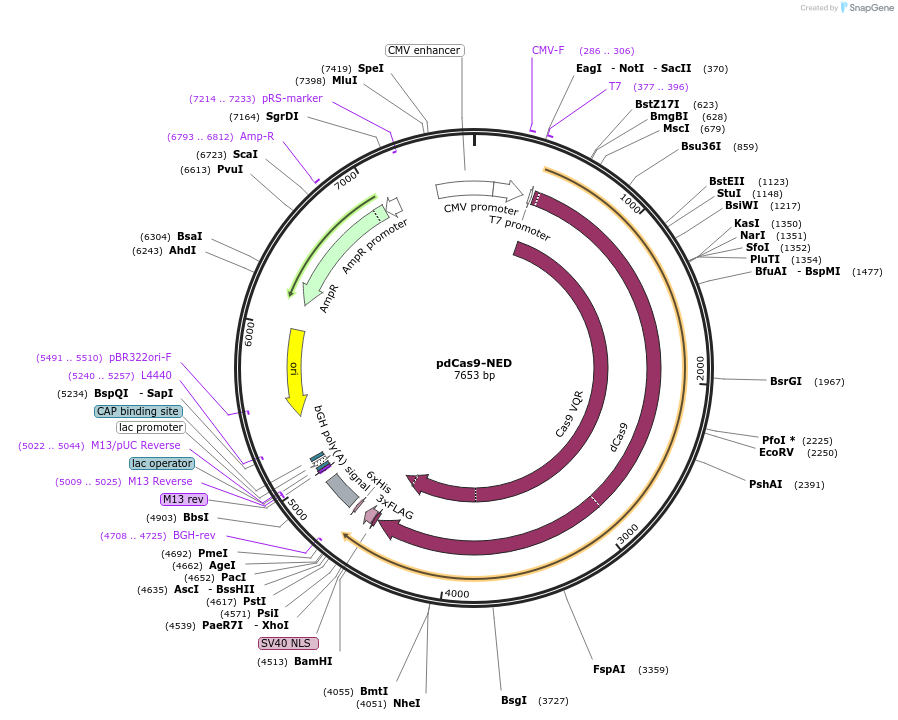
pdCas9-NED
(Plasmid
#109358)
-
Purpose(Empty Backbone) Mammalian vector for expressing effector domains fused to dCas9.
-
Depositing Lab
-
Sequence Information
Ordering
| Item | Catalog # | Description | Quantity | Price (USD) | |
|---|---|---|---|---|---|
| Plasmid | 109358 | Standard format: Plasmid sent in bacteria as agar stab | 1 | $94 | |
Backbone
-
Vector backbonepMLM3705 (Addgene plasmid #47754).
-
Backbone manufacturerJoung lab
- Backbone size (bp) 7636
-
Modifications to backboneThe ORF encoding VP64 was excised from pMLM3705 (Addgene plasmid #47754) by AscI-AgeI double-digestion and replaced with an oligonucleotide containing a PacI site.
-
Vector typeMammalian Expression, CRISPR
- Promoter CMV
-
Tag
/ Fusion Protein
- dCas9, SV40 NLS, 3xFLAG (N terminal on backbone)
Growth in Bacteria
-
Bacterial Resistance(s)Ampicillin, 100 μg/mL
-
Growth Temperature37°C
-
Growth Strain(s)DH5alpha
-
Copy numberHigh Copy
Cloning Information
- Cloning method Restriction Enzyme
- 5′ sequencing primer 5'-TACAAGGATGACGATGACAAG
- 3′ sequencing primer 5'-TAGAAGGCACAGTCGAGG
- (Common Sequencing Primers)
Resource Information
-
A portion of this plasmid was derived from a plasmid made byKeith Joung (Addgene plasmid #47754).
-
Articles Citing this Plasmid
Terms and Licenses
-
Academic/Nonprofit Terms
-
Industry Terms
- Not Available to Industry
Trademarks:
- Zeocin® is an InvivoGen trademark.
Depositor Comments
The DNA fragment encoding the effector domain is synthesized by PCR and cloned between the AscI and PacI sites of pdCas9-NED to create in-frame fusion between dCas9 (D10A, H840A) and the effector domain. For experiments involving cell lines stably expressing the dCas9-effector domain fusion protein, the coding sequence of the effector domain can be excised from the pdCas9-NED-based recombinant plasmid and transferred into pHAGE EF1α dCas9-NED (Addgene plasmid #109369). Cloning the AscI-PacI fragment between the MluI and AsiSI sites of pHAGE EF1α dCas9-NED preserves the reading frame of the fusion protein (Goubert et al., 2018).
These plasmids were created by your colleagues. Please acknowledge the Principal Investigator, cite the article in which the plasmids were described, and include Addgene in the Materials and Methods of your future publications.
-
For your Materials & Methods section:
pdCas9-NED was a gift from Antal Kiss (Addgene plasmid # 109358 ; http://n2t.net/addgene:109358 ; RRID:Addgene_109358) -
For your References section:
Establishment of Cell Lines Stably Expressing dCas9-Fusions to Address Kinetics of Epigenetic Editing. Goubert D, Koncz M, Kiss A, Rots MG. Methods Mol Biol. 2018;1767:395-415. doi: 10.1007/978-1-4939-7774-1_22. 10.1007/978-1-4939-7774-1_22 PubMed 29524148