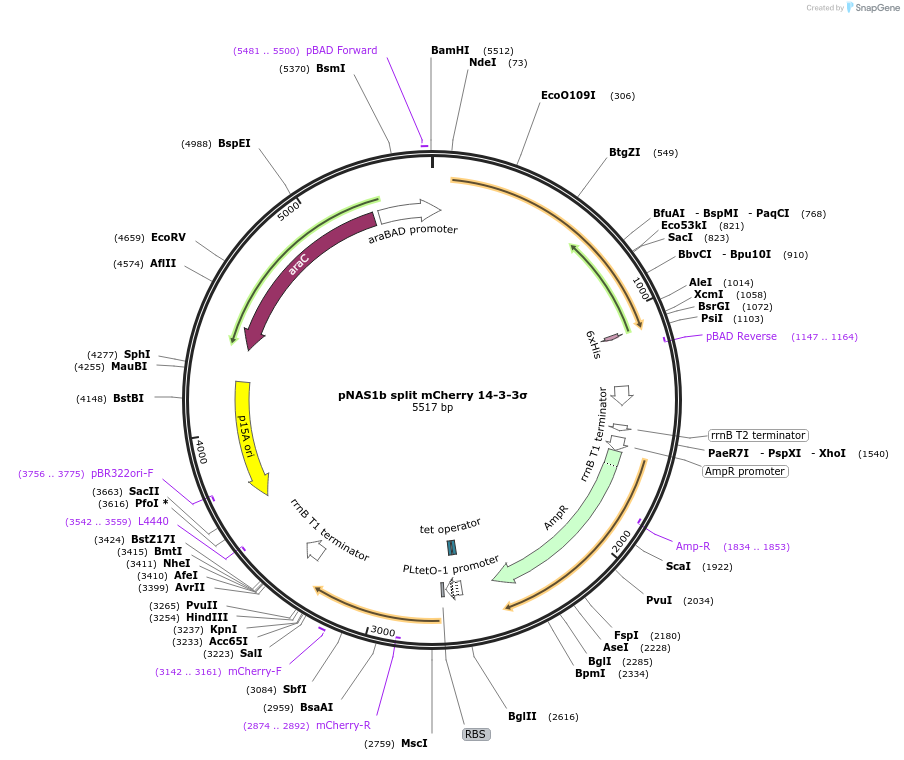
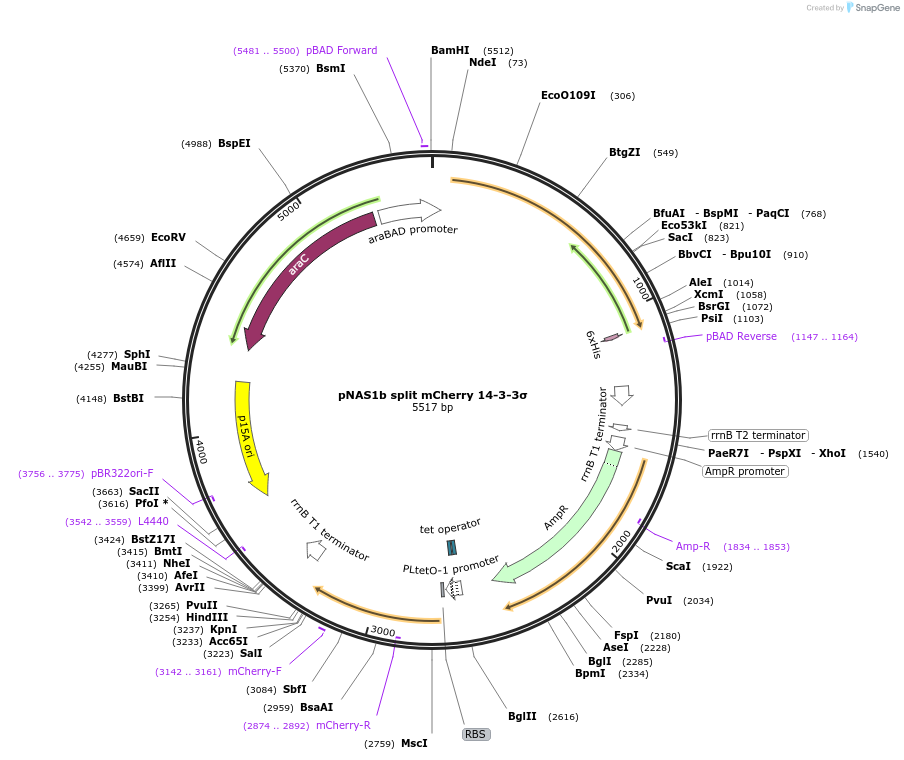
pNAS1b split mCherry 14-3-3σ
(Plasmid
#111881)
-
Purpose(Empty Backbone) Mode #2 phosphosite expression (split mCherry, using 14-3-3σ binding domain)
-
Depositing Lab
-
Sequence Information
Ordering
| Item | Catalog # | Description | Quantity | Price (USD) | |
|---|---|---|---|---|---|
| Plasmid | 111881 | Standard format: Plasmid sent in bacteria as agar stab | 1 | $94 | |
Backbone
-
Vector backbonepNAS1b
- Backbone size (bp) 5500
-
Vector typeBacterial Expression
-
Tag
/ Fusion Protein
- Split mCherry system: 14-3-3σ fused to C-terminal split mCherry; phosphosite empty cassette fused to N-terminal split mCherry. (N terminal on backbone)
Growth in Bacteria
-
Bacterial Resistance(s)Ampicillin, 100 μg/mL
-
Growth Temperature37°C
-
Growth Strain(s)DH5alpha
-
Copy numberUnknown
Cloning Information
- Cloning method Restriction Enzyme
Terms and Licenses
-
Academic/Nonprofit Terms
-
Industry Terms
- Not Available to Industry
Trademarks:
- Zeocin® is an InvivoGen trademark.
Depositor Comments
Backbone originally described in Sawyer, et al, Designed Phosphoprotein Recognition in Escherichia coli, 2014.
These plasmids were created by your colleagues. Please acknowledge the Principal Investigator, cite the article in which the plasmids were described, and include Addgene in the Materials and Methods of your future publications.
-
For your Materials & Methods section:
pNAS1b split mCherry 14-3-3σ was a gift from Jesse Rinehart (Addgene plasmid # 111881 ; http://n2t.net/addgene:111881 ; RRID:Addgene_111881) -
For your References section:
Encoding human serine phosphopeptides in bacteria for proteome-wide identification of phosphorylation-dependent interactions. Barber KW, Muir P, Szeligowski RV, Rogulina S, Gerstein M, Sampson JR, Isaacs FJ, Rinehart J. Nat Biotechnol. 2018 Aug;36(7):638-644. doi: 10.1038/nbt.4150. Epub 2018 Jun 11. 10.1038/nbt.4150 PubMed 29889213