-
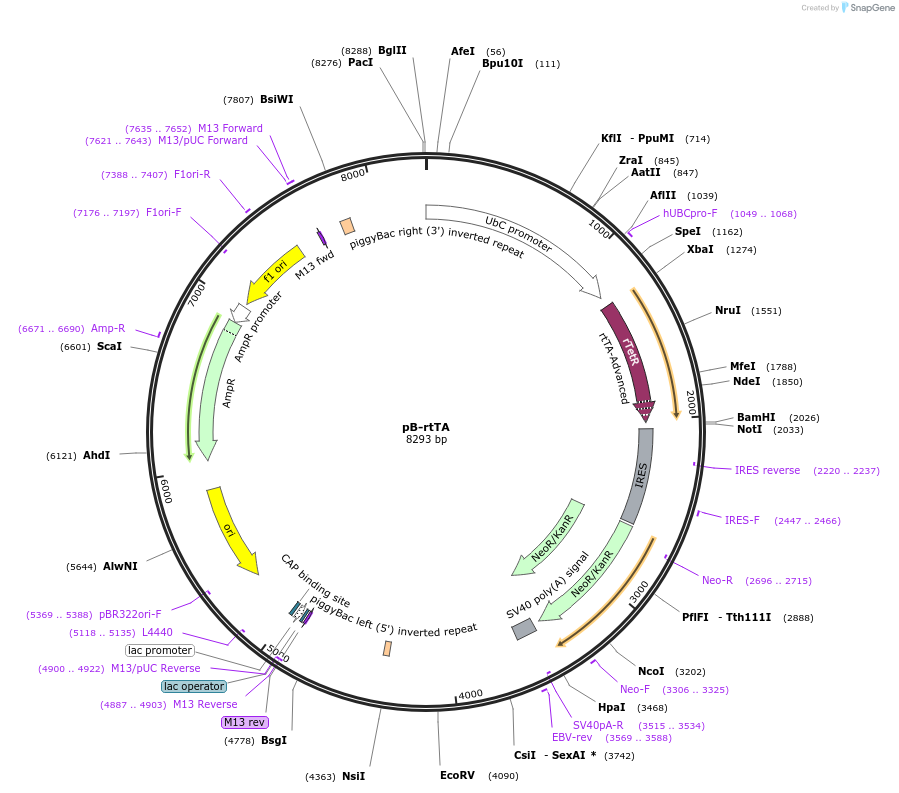
PurposePiggyBac cargo vector expressing rtTA. Also confers resistance to G418.
-
Depositing Lab
-
Sequence Information
Ordering
| Item | Catalog # | Description | Quantity | Price (USD) | |
|---|---|---|---|---|---|
| Plasmid | 126034 | Standard format: Plasmid sent in bacteria as agar stab | 1 | $94 | |
Backbone
-
Vector backbonepiggyBac cargo vector
-
Backbone manufacturerSystem Biosciences
- Total vector size (bp) 8306
-
Vector typeMammalian Expression ; piggyBac
-
Selectable markersNeomycin (select with G418)
Growth in Bacteria
-
Bacterial Resistance(s)Ampicillin, 100 μg/mL
-
Growth Temperature30°C
-
Growth Strain(s)DH5alpha
-
Copy numberUnknown
Gene/Insert
-
Gene/Insert namertTA3-IRES-Neo cassette cloned from Addgene Plasmid #25735
-
SpeciesSynthetic
-
Insert Size (bp)3846
- Promoter hUbiC
Cloning Information
- Cloning method Restriction Enzyme
- 5′ cloning site SfiI (not destroyed)
- 3′ cloning site SalI (not destroyed)
- 5′ sequencing primer NA
- (Common Sequencing Primers)
Resource Information
-
A portion of this plasmid was derived from a plasmid made byThe hUbiC-rtTA3-IRES-Neo cassette was cloned from pSLIK-Neo (Addgene#25735)
-
Articles Citing this Plasmid
Terms and Licenses
-
Academic/Nonprofit Terms
-
Industry Terms
- Not Available to Industry
Trademarks:
- Zeocin® is an InvivoGen trademark.
These plasmids were created by your colleagues. Please acknowledge the Principal Investigator, cite the article in which the plasmids were described, and include Addgene in the Materials and Methods of your future publications.
-
For your Materials & Methods section:
pB-rtTA was a gift from Mauro Calabrese (Addgene plasmid # 126034 ; http://n2t.net/addgene:126034 ; RRID:Addgene_126034) -
For your References section:
Functional classification of long non-coding RNAs by k-mer content. Kirk JM, Kim SO, Inoue K, Smola MJ, Lee DM, Schertzer MD, Wooten JS, Baker AR, Sprague D, Collins DW, Horning CR, Wang S, Chen Q, Weeks KM, Mucha PJ, Calabrese JM. Nat Genet. 2018 Oct;50(10):1474-1482. doi: 10.1038/s41588-018-0207-8. Epub 2018 Sep 17. 10.1038/s41588-018-0207-8 [pii] PubMed 30224646