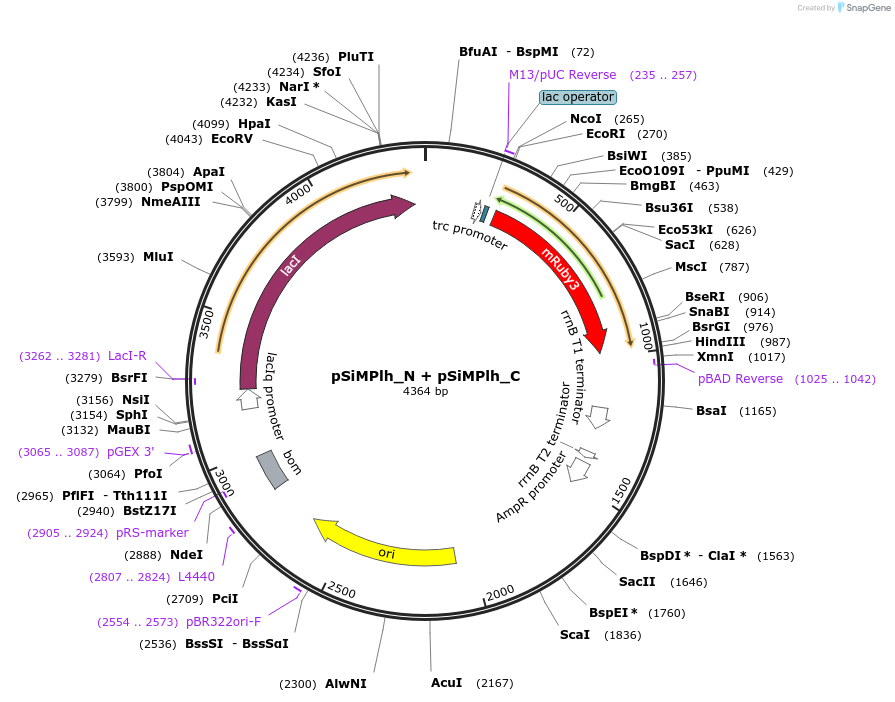
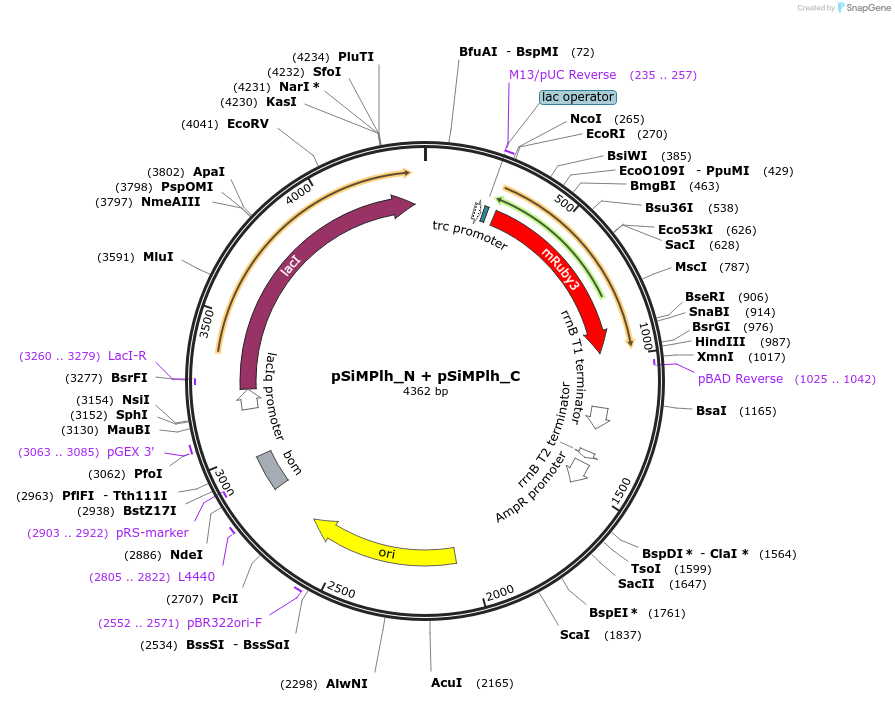
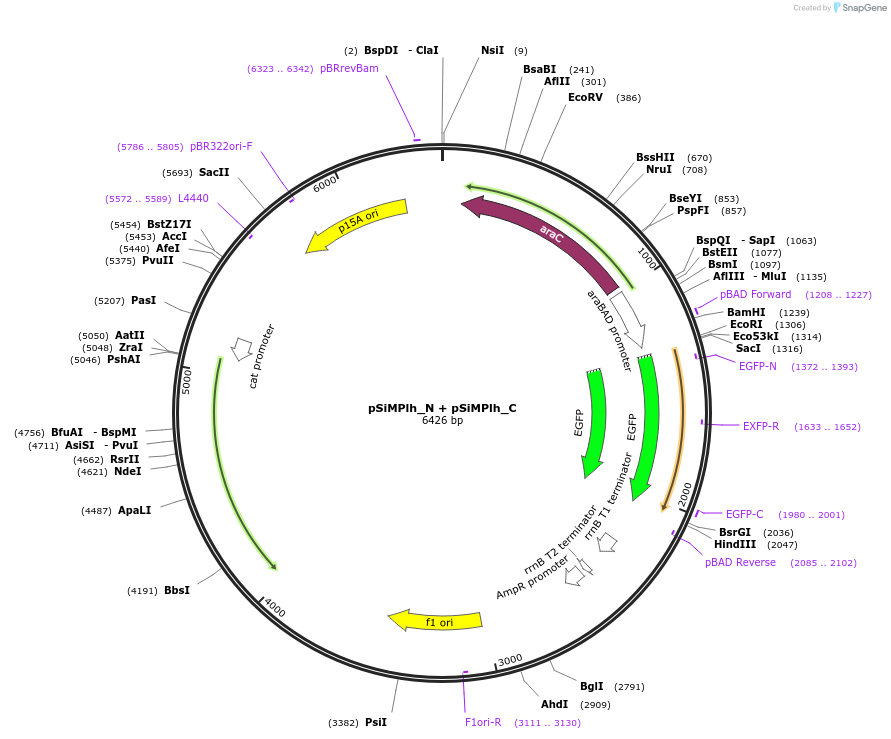
pSiMPlh_N + pSiMPlh_C
(Plasmid
#134316)
-
PurposeMaintenance of 2 plasmids with a single antibiotic in E. coli
-
Depositing Lab
-
Sequence Information
Ordering
| Item | Catalog # | Description | Quantity | Price (USD) | |
|---|---|---|---|---|---|
| Plasmid | 134316 | Standard format: Plasmid sent in bacteria as agar stab | 1 | $94 | |
Backbone
-
Vector backbonepBAD33 + pTrc99a
-
Vector typeBacterial Expression
Growth in Bacteria
-
Bacterial Resistance(s)Hygromycin, 200 μg/mL
-
Growth Temperature37°C
-
Growth Strain(s)DH5alpha
-
Copy numberHigh Copy
Gene/Insert
-
Gene/Insert namehygR_gp41-1 N-intein + hygR_gp41-1 C-intein
-
Alt nameHPT
-
SpeciesSynthetic
-
MutationP254Q in hygR_gp41-1 N-intein
Cloning Information
- Cloning method Gibson Cloning
- 5′ sequencing primer Unknown
- (Common Sequencing Primers)
Terms and Licenses
-
Academic/Nonprofit Terms
-
Industry Terms
- Not Available to Industry
Trademarks:
- Zeocin® is an InvivoGen trademark.
Depositor Comments
pSiMPlh_N should be cotransformed with pSiMPlh_C (or vice versa) in E. Coli to get a reconstituted functional HPT thereby conferring resistance against hygromycin. mRuby3 gene was amplified from a construct bought by us from Addgene (ID: 74252).
Please see the Sequences link for the individual plasmid sequences.
The sample from Addgene will have both the plasmids pSiMPlh_N + pSiMPlh_C in it for the interested
antibiotic resistance. Separating individual plasmid backbones will be possible via PCR. Agarose gel electrophoresis can be used to visualize the presence of both the plasmids. But, separating individual plasmids via gel electrophoresis, in our hands, always resulted in cross-contamination of the two backbones.
These plasmids were created by your colleagues. Please acknowledge the Principal Investigator, cite the article in which the plasmids were described, and include Addgene in the Materials and Methods of your future publications.
-
For your Materials & Methods section:
pSiMPlh_N + pSiMPlh_C was a gift from Barbara Di Ventura (Addgene plasmid # 134316 ; http://n2t.net/addgene:134316 ; RRID:Addgene_134316) -
For your References section:
Split intein-mediated selection of cells containing two plasmids using a single antibiotic. Palanisamy N, Degen A, Morath A, Ballestin Ballestin J, Juraske C, Ozturk MA, Sprenger GA, Youn JW, Schamel WW, Di Ventura B. Nat Commun. 2019 Oct 31;10(1):4967. doi: 10.1038/s41467-019-12911-1. 10.1038/s41467-019-12911-1 PubMed 31672972