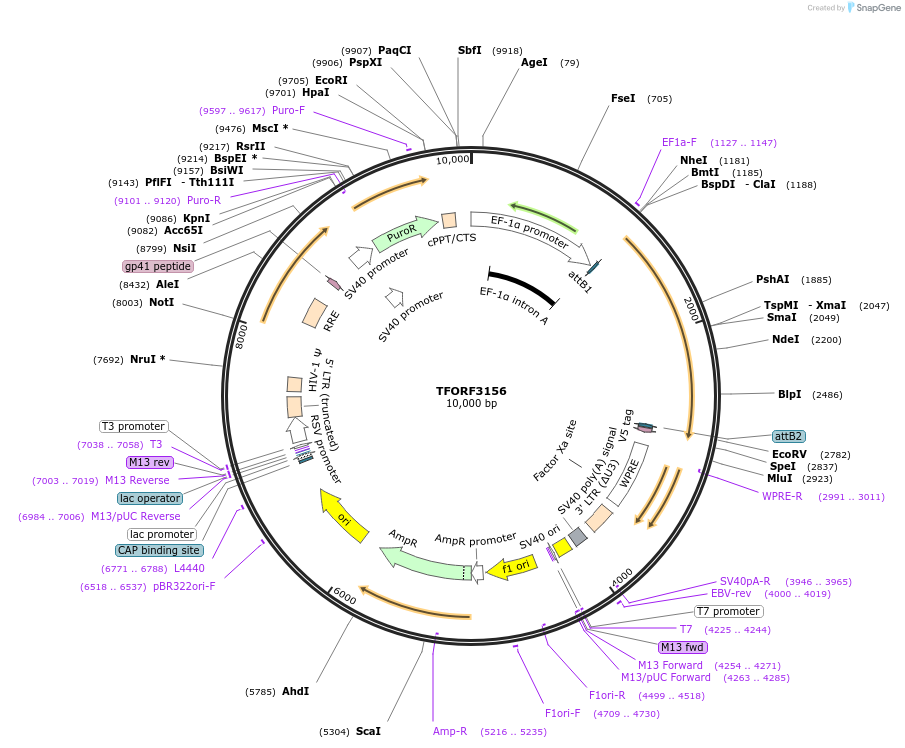
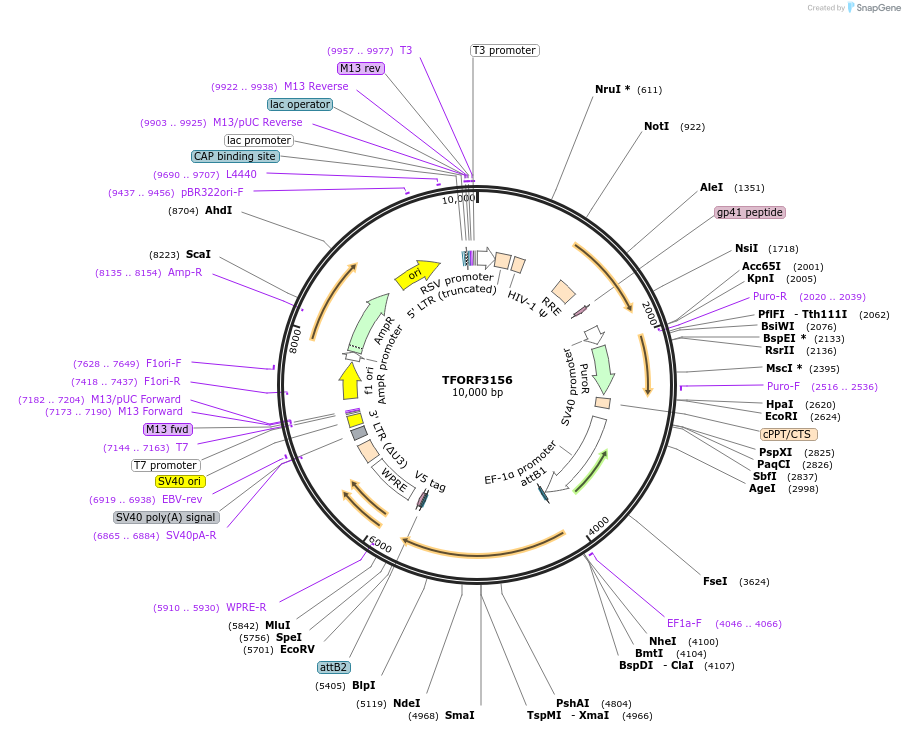
TFORF3156
(Plasmid
#144632)
-
PurposeLentiviral vector for overexpressing transcription factor ORFs with unique 24-bp barcodes. Barcodes facilitate identification of transcription factors in pooled screens.
-
Depositing Lab
-
Sequence Information
Ordering
| Item | Catalog # | Description | Quantity | Price (USD) | |
|---|---|---|---|---|---|
| Plasmid | 144632 | Standard format: Plasmid sent in bacteria as agar stab | 1 | $94 | |
Backbone
-
Vector backbonepLX_TRC317
-
Backbone manufacturerBroad Institute Genetic Perturbation Platform
- Backbone size w/o insert (bp) 8264
-
Vector typeMammalian Expression, Lentiviral
-
Selectable markersPuromycin
Growth in Bacteria
-
Bacterial Resistance(s)Ampicillin, 100 μg/mL
-
Growth Temperature37°C
-
Growth Strain(s)NEB Stable
-
Copy numberHigh Copy
Gene/Insert
-
Gene/Insert nameETV5
-
SpeciesH. sapiens (human)
-
Insert Size (bp)1736
-
Entrez GeneETV5 (a.k.a. ERM)
- Promoter EF1a
-
Tag
/ Fusion Protein
- V5 (C terminal on insert)
Cloning Information
- Cloning method Gateway Cloning
- 5′ sequencing primer GGGTGGAGACTGAAGTTAGGCCAG
- 3′ sequencing primer CACATAGCGTAAAAGGAGCAACATAG
- (Common Sequencing Primers)
Resource Information
-
A portion of this plasmid was derived from a plasmid made byBroad Institute Genetic Perturbation Platform
-
Article Citing this Plasmid
Terms and Licenses
-
Academic/Nonprofit Terms
-
Industry Terms
- Not Available to Industry
Trademarks:
- Zeocin® is an InvivoGen trademark.
Depositor Comments
NM_004454.2 Please test multiple small colonies in case of plasmid recombination. To make this collection available in a timely manner, a portion of this collection was not fully sequenced by Addgene. If an Addgene verified full plasmid sequence is not available, please contact us at [email protected] prior to placing an order to request the full sequence.
These plasmids were created by your colleagues. Please acknowledge the Principal Investigator, cite the article in which the plasmids were described, and include Addgene in the Materials and Methods of your future publications.
-
For your Materials & Methods section:
TFORF3156 was a gift from Feng Zhang (Addgene plasmid # 144632 ; http://n2t.net/addgene:144632 ; RRID:Addgene_144632) -
For your References section:
A transcription factor atlas of directed differentiation. Joung J, Ma S, Tay T, Geiger-Schuller KR, Kirchgatterer PC, Verdine VK, Guo B, Arias-Garcia MA, Allen WE, Singh A, Kuksenko O, Abudayyeh OO, Gootenberg JS, Fu Z, Macrae RK, Buenrostro JD, Regev A, Zhang F. Cell. 2023 Jan 5;186(1):209-229.e26. doi: 10.1016/j.cell.2022.11.026. 10.1016/j.cell.2022.11.026 PubMed 36608654