-
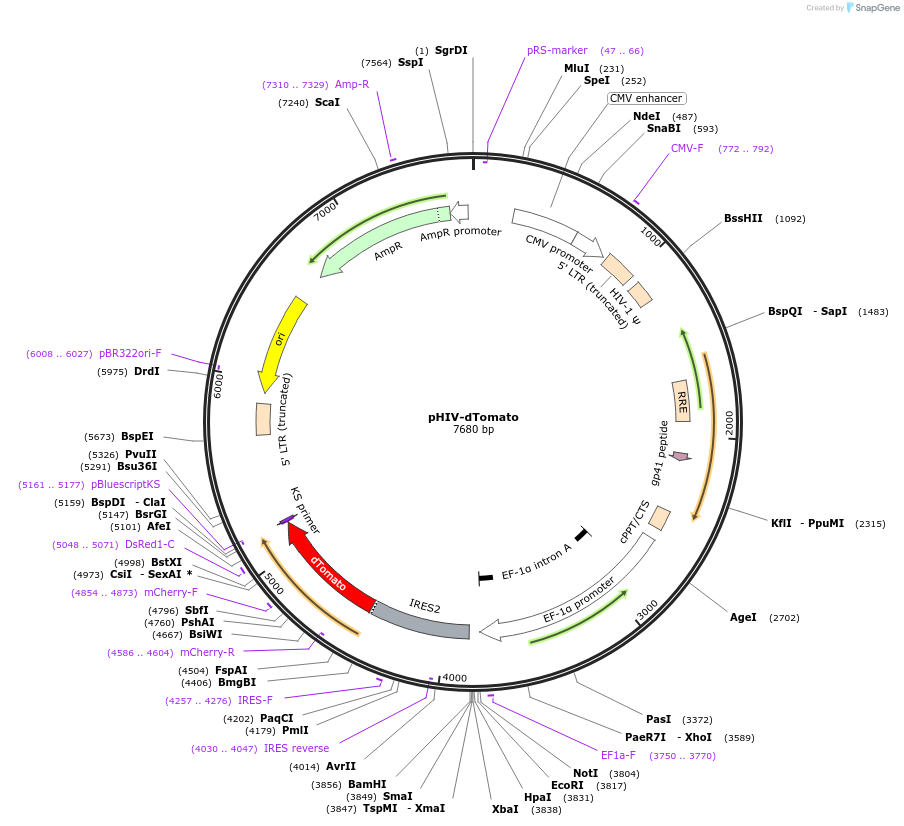
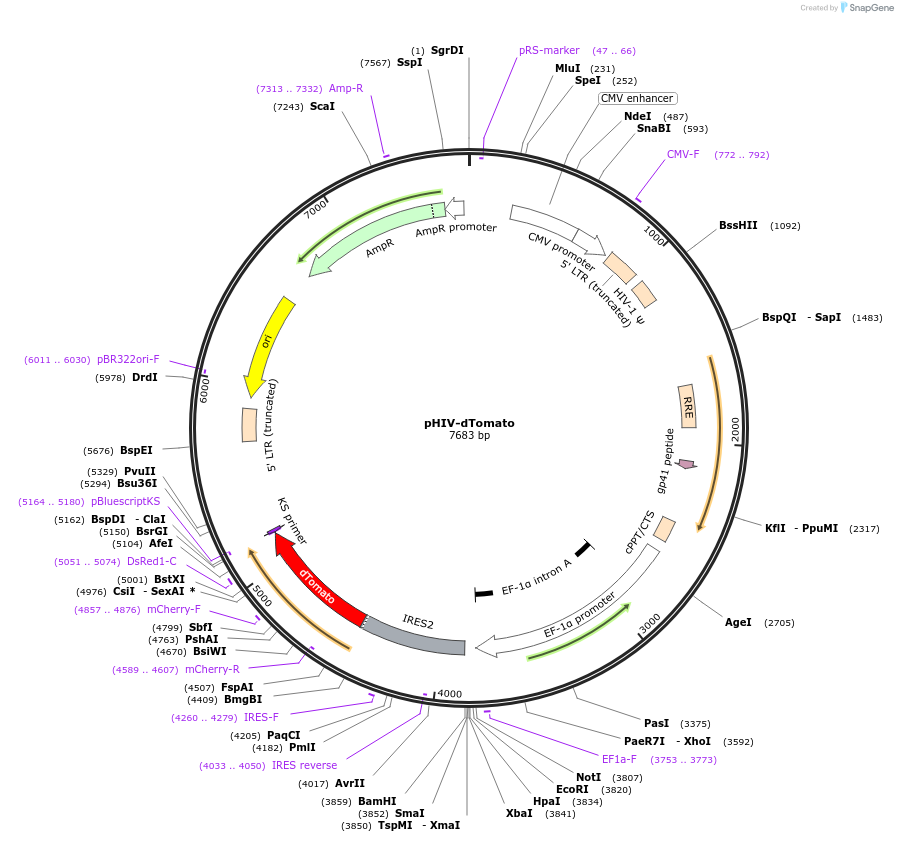
Purpose(Empty Backbone) self-inactivating lentiviral plasmid for co-expression of your gene of interest and dTomato
-
Depositing Lab
-
Publication
-
Sequence Information
Ordering
| Item | Catalog # | Description | Quantity | Price (USD) | |
|---|---|---|---|---|---|
| Plasmid | 21374 | Standard format: Plasmid sent in bacteria as agar stab | 1 | $94 | |
Backbone
-
Vector backbonepSICO
- Backbone size (bp) 6900
-
Vector typeMammalian Expression, Lentiviral
Growth in Bacteria
-
Bacterial Resistance(s)Ampicillin, 100 μg/mL
-
Growth Temperature37°C
-
Growth Strain(s)Stbl3
-
Growth instructionsGrowth in Stbl3 is preferred
-
Copy numberHigh Copy
Gene/Insert
-
Gene/Insert nameNone
Cloning Information
- Cloning method Restriction Enzyme
- 5′ cloning site NA (unknown if destroyed)
- 3′ cloning site NA (unknown if destroyed)
- 5′ sequencing primer 5'-TGGAATTTGCCCTTTTTGAG-3'
- 3′ sequencing primer 5'-AGGAACTGCTTCCTTCACGA-3'
- (Common Sequencing Primers)
Resource Information
-
A portion of this plasmid was derived from a plasmid made byThe red fluorescent protein dTomato was obtained from Roger Tsien's laboratory and was published in: Nature Biotechnology 22, 1567 - 1572 (2004) and pSICO was obtained from Tyler Jacks laboratory.
-
Articles Citing this Plasmid
Terms and Licenses
-
Academic/Nonprofit Terms
-
Industry Terms
- Not Available to Industry
Trademarks:
- Zeocin® is an InvivoGen trademark.
Depositor Comments
The HIV-dTomato plasmid encodes a self-inactivating lentiviral plasmid that was cloned by removing the U6-TATAlox-CMVie-EGFP-TATAlox- WPRE content of pSICO (Ventura et al., 2004), and adding the EF1-alpha promoter, a multiple cloning site (MCS), an internal ribosome entry site (IRES) and EGFP. A cDNA can be cloned into the MCS (NotI, EcoRI, HpaI, XbaI, SmaI, and BamHI sites) to enable bicistronic expression with dTomato. This construct was made in Zena Werb's laboratory and has not been published. Note: This plasmid contains dTomato and not the tandem conjugate TdTomato.
These plasmids were created by your colleagues. Please acknowledge the Principal Investigator, cite the article in which the plasmids were described, and include Addgene in the Materials and Methods of your future publications.
-
For your Materials & Methods section:
pHIV-dTomato was a gift from Bryan Welm (Addgene plasmid # 21374 ; http://n2t.net/addgene:21374 ; RRID:Addgene_21374)