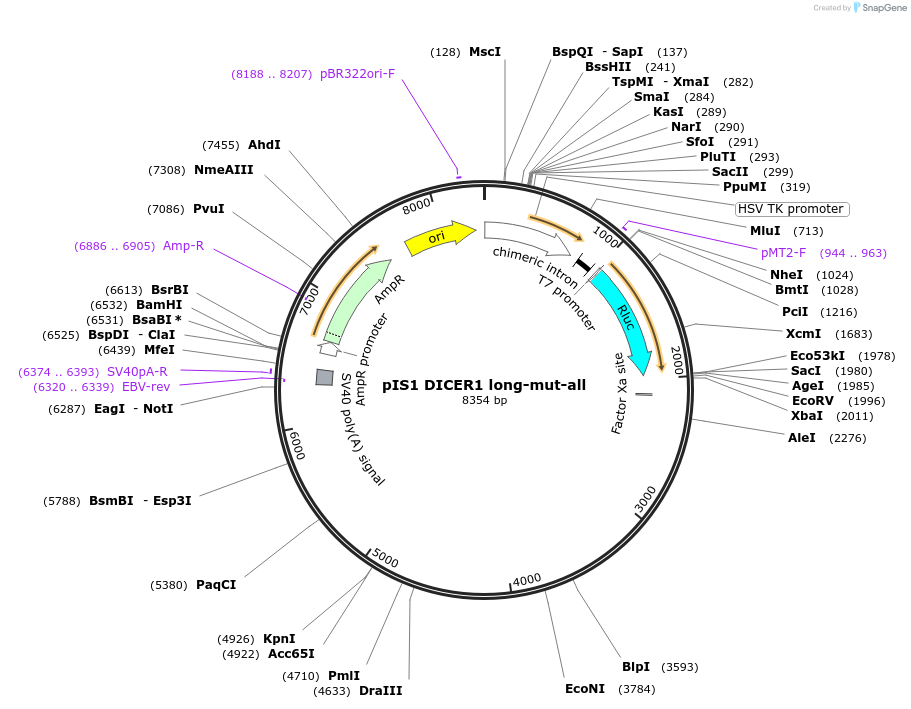
pIS1 DICER1 long-mut-all
(Plasmid
#21653)
-
Depositing Lab
-
Publication
-
Sequence Information
Ordering
| Item | Catalog # | Description | Quantity | Price (USD) | |
|---|---|---|---|---|---|
| Plasmid | 21653 | Standard format: Plasmid sent in bacteria as agar stab | 1 | $94 | |
Backbone
-
Vector backbonepIS1
-
Backbone manufacturerAvailable from Addgene (#12179)
- Backbone size w/o insert (bp) 4085
-
Vector typeMammalian Expression, Luciferase
Growth in Bacteria
-
Bacterial Resistance(s)Ampicillin, 100 μg/mL
-
Growth Temperature37°C
-
Growth Strain(s)DH5alpha
-
Copy numberHigh Copy
Gene/Insert
-
Gene/Insert nameDicer 1
-
Alt nameCM12
-
SpeciesH. sapiens (human)
-
Insert Size (bp)4200
-
MutationMutated let-7 and miR103/107 microRNA binding sites
-
Entrez GeneDICER1 (a.k.a. DCR1, Dicer, Dicer1e, GLOW, HERNA, K12H4.8-LIKE, MNG1, RMSE2, aviD)
-
Tag
/ Fusion Protein
- Luciferase (N terminal on backbone)
Cloning Information
- Cloning method Restriction Enzyme
- 5′ cloning site Xba I (not destroyed)
- 3′ cloning site Not I (not destroyed)
- 5′ sequencing primer n/a
- 3′ sequencing primer EBV Reverse
- (Common Sequencing Primers)
Resource Information
-
Article Citing this Plasmid
Terms and Licenses
-
Academic/Nonprofit Terms
-
Industry Terms
- Not Available to Industry
Trademarks:
- Zeocin® is an InvivoGen trademark.
Depositor Comments
Long means that all functional alternative polyadenylation signals have been deleted.
Mut means that all microRNA binding sites for an indicated microRNA have been mutated.
These plasmids were created by your colleagues. Please acknowledge the Principal Investigator, cite the article in which the plasmids were described, and include Addgene in the Materials and Methods of your future publications.
-
For your Materials & Methods section:
pIS1 DICER1 long-mut-all was a gift from David Bartel (Addgene plasmid # 21653 ; http://n2t.net/addgene:21653 ; RRID:Addgene_21653) -
For your References section:
Widespread shortening of 3'UTRs by alternative cleavage and polyadenylation activates oncogenes in cancer cells. Mayr C, Bartel DP. Cell. 2009 Aug 21. 138(4):673-84. 10.1016/j.cell.2009.06.016 PubMed 19703394