-
Depositing Lab
-
Publication
-
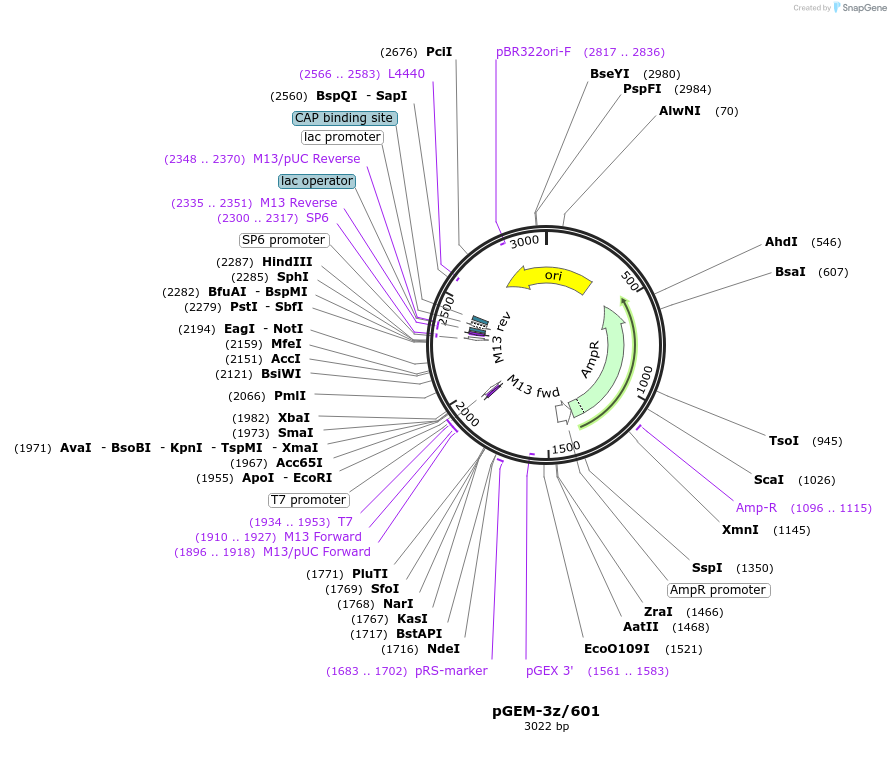
Sequence Information
Ordering
| Item | Catalog # | Description | Quantity | Price (USD) | |
|---|---|---|---|---|---|
| Plasmid | 26656 | Standard format: Plasmid sent in bacteria as agar stab | 1 | $94 | |
Backbone
-
Vector backbonepGEM-3Z
-
Backbone manufacturerPromega
- Backbone size w/o insert (bp) 2743
-
Vector typeBacterial Expression
Growth in Bacteria
-
Bacterial Resistance(s)Ampicillin, 100 μg/mL
-
Growth Temperature37°C
-
Growth Strain(s)DH5alpha
-
Copy numberHigh Copy
Gene/Insert
-
Gene/Insert nameNone
-
Insert Size (bp)282
Cloning Information
- Cloning method Restriction Enzyme
- 5′ cloning site HincII (unknown if destroyed)
- 3′ cloning site HincII (unknown if destroyed)
- 5′ sequencing primer T7
- 3′ sequencing primer SP6
- (Common Sequencing Primers)
Resource Information
-
Articles Citing this Plasmid
Terms and Licenses
-
Academic/Nonprofit Terms
-
Industry Terms
- Not Available to Industry
Trademarks:
- Zeocin® is an InvivoGen trademark.
Depositor Comments
This plasmid contains a 147 bp nucleosome positioning sequence and is referred to as clone"601".
Primers that are used by the Widom lab to amplify the 147 bp fragment:
Forward:
ctggagaatcccggtgccg
Reverse:
acaggatgtatatatctgacacg
Please note that there may be discrepancies between Addgene's sequencing results and the depositor's sequence. Potential discrepancies are not a concern according to the depositing scientist because the clones were selected for their ability to bind to histone octamer, not for their specific sequence.
These plasmids were created by your colleagues. Please acknowledge the Principal Investigator, cite the article in which the plasmids were described, and include Addgene in the Materials and Methods of your future publications.
-
For your Materials & Methods section:
pGEM-3z/601 was a gift from Jonathan Widom (Addgene plasmid # 26656 ; http://n2t.net/addgene:26656 ; RRID:Addgene_26656) -
For your References section:
New DNA sequence rules for high affinity binding to histone octamer and sequence-directed nucleosome positioning. Lowary PT, Widom J. J Mol Biol. 1998 Feb 13. 276(1):19-42. 10.1006/jmbi.1997.1494 PubMed 9514715