-
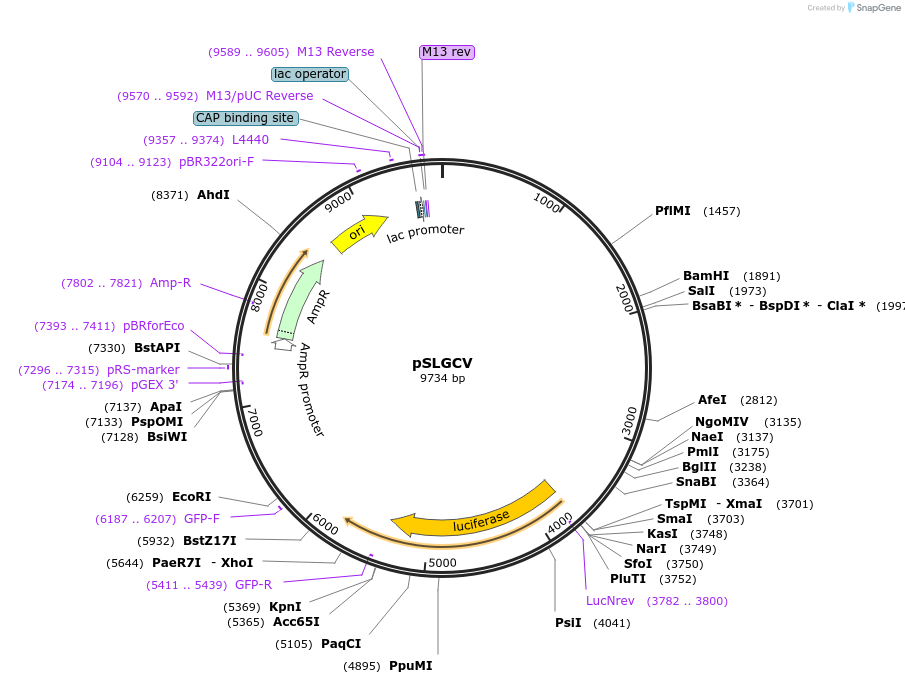
PurposeExpression in Caenorhabditis elegans of the luc+ gene (Promega), fused in frame to GFP (S65C), under the sur-5 promoter (from plasmid pTG96, John Yochem and Min Han). Backbone pPD95.79 (Firelab).
-
Depositing Labs
-
Sequence Information
Ordering
| Item | Catalog # | Description | Quantity | Price (USD) | |
|---|---|---|---|---|---|
| Plasmid | 49862 | Standard format: Plasmid sent in bacteria as agar stab | 1 | $94 | |
Backbone
-
Vector backbonepPD95.79
-
Backbone manufacturerAndrew Fire, Carnegie Institution for Science
-
Modifications to backboneadded sequentially the sur-5 promoter [36], PCR-amplified from plasmid pTG96 and then the luciferase gene PCR-amplified from vector pSP-luc+ (Promega). A 3.7 kb fragment upstream of the sur-5 gene containing the promoter, was amplified using primers 5'- ATAAGCTTGCATGCCTGCATTGC-3' and 5'-AGACACCCCGGGCTTTCTGAAAAC-3' (this introduces a Sma I site and removes the start codon for the sur-5 gene). The sur-5 promoter was inserted in the pPD95.79 backbone using appropriate restriction sites (Sph I and Sma I). The luc+ gene was amplified using primers 5'-TCCCGGGAAGCTTTCCATGGAAGAC-3' and 5'-CTAGGGTACCACGGCGATCTTTCC-3'. Sma I and Kpn I were used to insert luc+ downstream of the sur-5 promoter and upstream of and in frame with gfp. The peroxisome tagging sequence is absent from luc+ and the NLS for sur-5 was excluded from pSLGCV, so that LUC+::GFP was expressed in the cytoplasm
-
Vector typeWorm Expression
Growth in Bacteria
-
Bacterial Resistance(s)Ampicillin, 100 μg/mL
-
Growth Temperature37°C
-
Growth Strain(s)DH5alpha
-
Copy numberHigh Copy
Gene/Insert
-
Gene/Insert namefirefly luciferase luc+ fused to GFP (S65C)
-
SpeciesPhotinus pyralis
- Promoter sur-5
-
Tag
/ Fusion Protein
- GFP (S65C) (C terminal on backbone)
Cloning Information
- Cloning method Restriction Enzyme
- 5′ cloning site SmaI (unknown if destroyed)
- 3′ cloning site KpnI (unknown if destroyed)
- 5′ sequencing primer 5'-TCCCGGGAAGCTTTCCATGGAAGAC-3'
- 3′ sequencing primer 5'-CTAGGGTACCACGGCGATCTTTCC-3'
- (Common Sequencing Primers)
Resource Information
-
A portion of this plasmid was derived from a plasmid made byvector backbone pPD95.79 from Firelab, Carnegie Institution for Science. pTG96 plasmid (from which promoter was PCR- amplified) obtained from John Yochem and Min Han; University of Colorado, Boulder. vector pSP-luc+ (bought from Promega. Luc+ gene was PCR-amplified from here.
-
Articles Citing this Plasmid
Terms and Licenses
-
Academic/Nonprofit Terms
-
Industry Terms
- Not Available to Industry
Trademarks:
- Zeocin® is an InvivoGen trademark.
Depositor Comments
Although no formal MTAs signed with Firelab or University of Colorado Boulder, we accepted that the material was provided for academic research only.
These plasmids were created by your colleagues. Please acknowledge the Principal Investigator, cite the article in which the plasmids were described, and include Addgene in the Materials and Methods of your future publications.
-
For your Materials & Methods section:
pSLGCV was a gift from Anne Glover & Jonathan Pettitt (Addgene plasmid # 49862 ; http://n2t.net/addgene:49862 ; RRID:Addgene_49862) -
For your References section:
Bridging the phenotypic gap: real-time assessment of mitochondrial function and metabolism of the nematode Caenorhabditis elegans. Lagido C, Pettitt J, Flett A, Glover LA. BMC Physiol. 2008 Apr 2;8:7. doi: 10.1186/1472-6793-8-7. 10.1186/1472-6793-8-7 PubMed 18384668