-
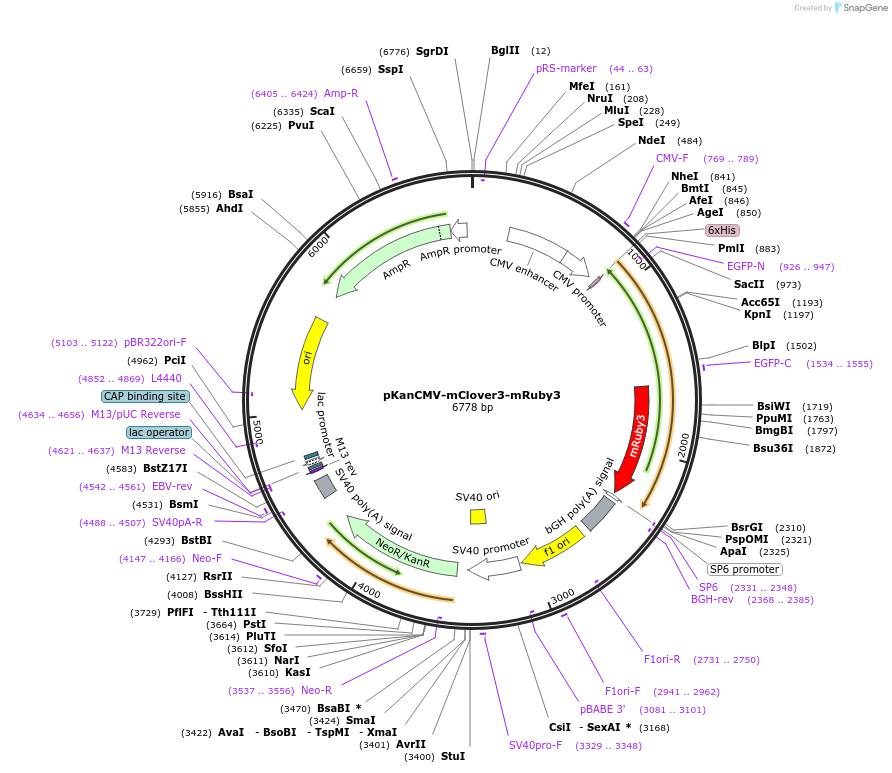
PurposeMammalian expression of tandem fusion of mClover3-mRuby3
-
Depositing Lab
-
Sequence Information
Ordering
| Item | Catalog # | Description | Quantity | Price (USD) | |
|---|---|---|---|---|---|
| Plasmid | 74252 | Standard format: Plasmid sent in bacteria as agar stab | 1 | $94 | |
Backbone
-
Vector backbonepKanCMV
-
Backbone manufacturerMichael Davidson Lab
- Total vector size (bp) 6778
-
Vector typeMammalian Expression
-
Selectable markersNeomycin (select with G418)
Growth in Bacteria
-
Bacterial Resistance(s)Ampicillin, 100 μg/mL
-
Growth Temperature37°C
-
Growth Strain(s)DH5alpha
-
Copy numberHigh Copy
Gene/Insert
-
Gene/Insert namemClover3-mRuby3
-
Insert Size (bp)1473
- Promoter CMV
Cloning Information
- Cloning method Restriction Enzyme
- 5′ cloning site BshTI (unknown if destroyed)
- 3′ cloning site ApaI (unknown if destroyed)
- 5′ sequencing primer pshuttle5
- 3′ sequencing primer BGHrev
- (Common Sequencing Primers)
Resource Information
-
Articles Citing this Plasmid
Terms and Licenses
-
Academic/Nonprofit Terms
-
Industry Terms
- Not Available to Industry
Trademarks:
- Zeocin® is an InvivoGen trademark.
Depositor Comments
The specific substitutions used in the development of mRuby3 relative to its parent mRuby2 are N33R, M36E, T38V, K74A, G75D, M105T, C114E, H118N, Q120K, H159D, M160I, S171H, S173N, I192V, L202I, M209T, F210Y, H216V, F221Y, A222S, G223N.
The specific substitutions used in the development of mClover3 relative to its parent Clover are N149Y, G160C, and A206K.
These plasmids were created by your colleagues. Please acknowledge the Principal Investigator, cite the article in which the plasmids were described, and include Addgene in the Materials and Methods of your future publications.
-
For your Materials & Methods section:
pKanCMV-mClover3-mRuby3 was a gift from Michael Lin (Addgene plasmid # 74252 ; http://n2t.net/addgene:74252 ; RRID:Addgene_74252) -
For your References section:
Improving brightness and photostability of green and red fluorescent proteins for live cell imaging and FRET reporting. Bajar BT, Wang ES, Lam AJ, Kim BB, Jacobs CL, Howe ES, Davidson MW, Lin MZ, Chu J. Sci Rep. 2016 Feb 16;6:20889. doi: 10.1038/srep20889. 10.1038/srep20889 PubMed 26879144