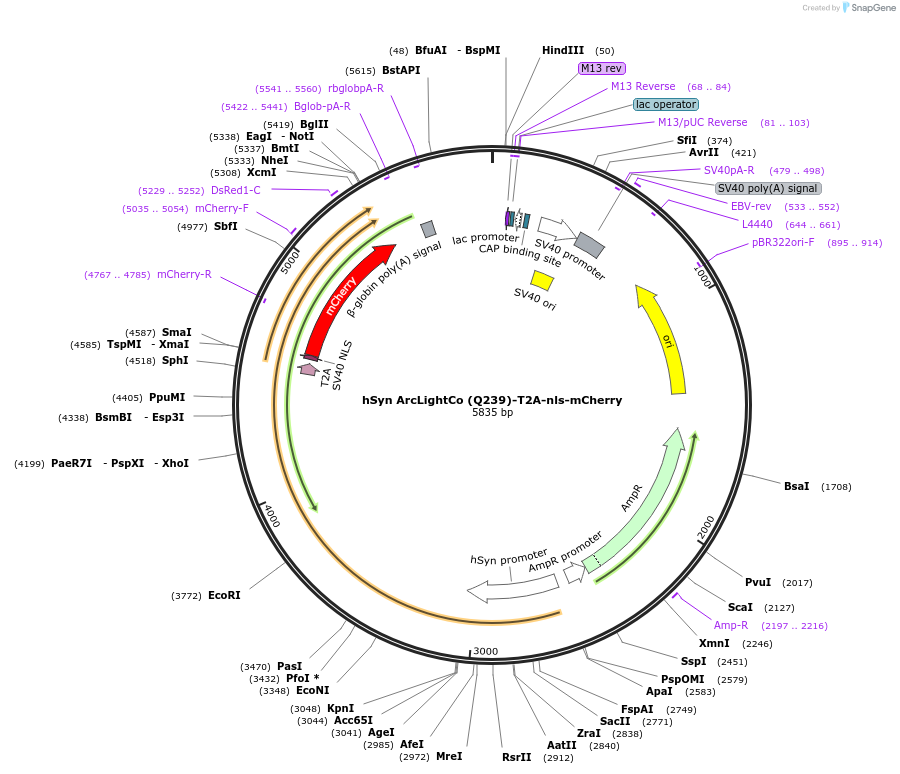
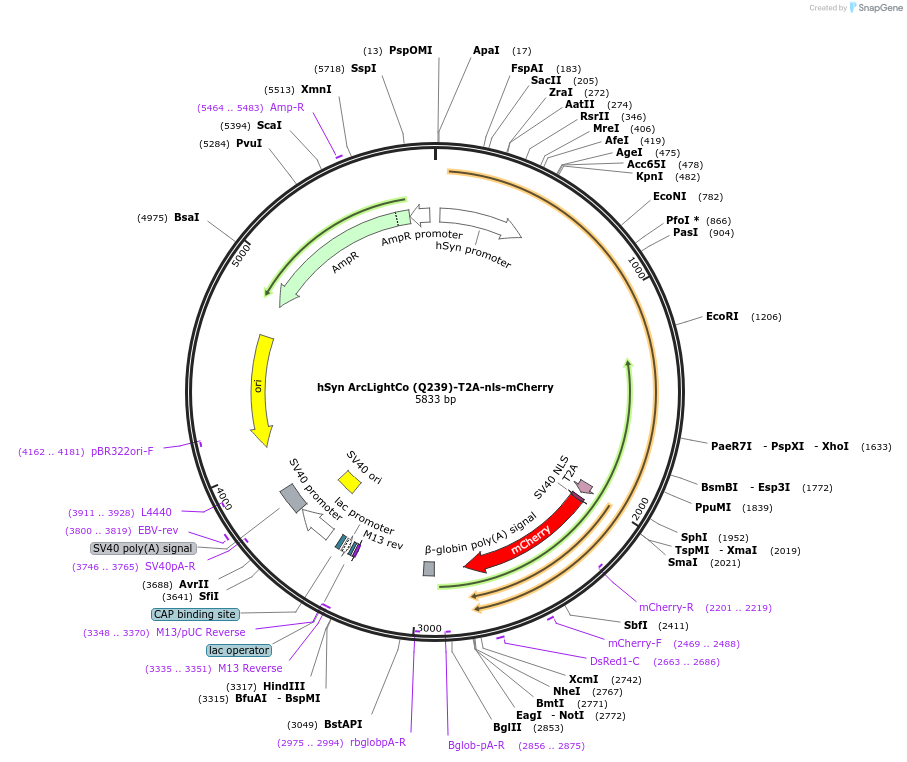
hSyn ArcLightCo (Q239)-T2A-nls-mCherry
(Plasmid
#85844)
-
PurposeGenetically encoded voltage sensor ArcLight- codon optimized
-
Depositing Lab
-
Sequence Information
Ordering
| Item | Catalog # | Description | Quantity | Price (USD) | |
|---|---|---|---|---|---|
| Plasmid | 85844 | Standard format: Plasmid sent in bacteria as agar stab | 1 | $94 | |
Backbone
-
Vector backbonepCS2+ hSyn1
- Backbone size w/o insert (bp) 4372
- Total vector size (bp) 5833
-
Modifications to backbonea hygromysin resistance gene was inserted to facilitate selection for stably transfected cells.
-
Vector typeMammalian Expression
-
Selectable markersHygromycin
Growth in Bacteria
-
Bacterial Resistance(s)Ampicillin, 100 μg/mL
-
Growth Temperature37°C
-
Growth Strain(s)DH5alpha
-
Copy numberHigh Copy
Gene/Insert
-
Gene/Insert nameArcLightCo-Q239
-
Alt nameArcLight
-
Alt nameCiVSP
-
Alt nameSuper Ecliptic pHluorin
-
SpeciesCiona intestinalis
-
Insert Size (bp)1461
-
MutationCi-VSP contains R217Q mutation; super ecliptic pHluorin contains A227D mutation. The A227D mutation increases the fluorescence response magnitude. We replaced residues 71-91 within CiVSD with an amino acid sequence (KSRITSEGEYIPLDQIDINV) from the Kir2.1 channel Golgi-to-plasma membrane trafficking signal. The Kir 2.1 channel ER export signal sequence (FCYENEV) was added to C-terminus of super ecliptic pHluorin. The sequence is codon optimized using mammalian codon preference.
-
GenBank IDAB183035 AY533296
- Promoter hSyn1
-
Tag
/ Fusion Protein
- mCherry (C terminal on insert)
Cloning Information
- Cloning method Restriction Enzyme
- 5′ cloning site KpnI (not destroyed)
- 3′ cloning site SphI (not destroyed)
- 5′ sequencing primer AGTCGTGTCGTGCCTGAGAG
- 3′ sequencing primer ccttgagcatctgacttctggcta
- (Common Sequencing Primers)
Resource Information
-
A portion of this plasmid was derived from a plasmid made byDr. Atsushi Miyawaki, Laboratory for Cell Function Dynamics, Brain Science Institute, RIKEN, 2-1 Hirosawa, Saitama 351-0198, Japan.
Terms and Licenses
-
Academic/Nonprofit Terms
-
Industry Terms
- Not Available to Industry
Trademarks:
- Zeocin® is an InvivoGen trademark.
Depositor Comments
A portion of this plasmid was derived from a plasmid made by Dr. Atsushi Miyawaki, Laboratory for Cell Function Dynamics, Brain Science Institute, RIKEN, 2-1 Hirosawa, Saitama 351-0198, Japan.
These plasmids were created by your colleagues. Please acknowledge the Principal Investigator, cite the article in which the plasmids were described, and include Addgene in the Materials and Methods of your future publications.
-
For your Materials & Methods section:
hSyn ArcLightCo (Q239)-T2A-nls-mCherry was a gift from Vincent Pieribone (Addgene plasmid # 85844 ; http://n2t.net/addgene:85844 ; RRID:Addgene_85844) -
For your References section:
Directed evolution of key residues in fluorescent protein inverses the polarity of voltage sensitivity in the genetically-encoded indicator ArcLight. Platisa J, Vasan G, Yang A, Pieribone VA. ACS Chem Neurosci. 2017 Jan 3. doi: 10.1021/acschemneuro.6b00234. 10.1021/acschemneuro.6b00234 PubMed 28045247